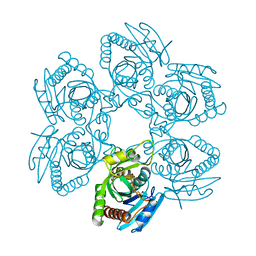
3EMV
 | |
3ENZ
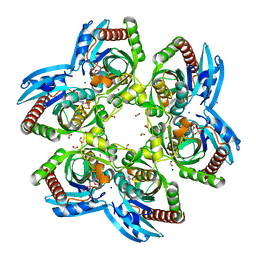 
 | | Arsenolytic structure of Plasmodium falciparum purine nucleoside phosphorylase with hypoxanthine, ribose and arsenate ion | | Descriptor: | 1,4-anhydro-D-ribitol, ARSENATE, FORMIC ACID, ... | | Authors: | Chaikuad, A, Brady, R.L. | | Deposit date: | 2008-09-26 | | Release date: | 2009-08-04 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | Conservation of structure and activity in Plasmodium purine nucleoside phosphorylases.
Bmc Struct.Biol., 9, 2009
|
|
1OC4
 
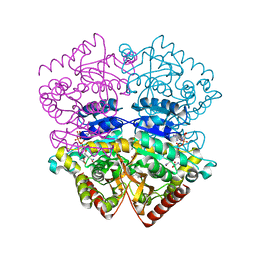 | | Lactate dehydrogenase from Plasmodium berghei | | Descriptor: | GLYCEROL, L-LACTATE DEHYDROGENASE, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, ... | | Authors: | Winter, V.J, Brady, R.L. | | Deposit date: | 2003-02-05 | | Release date: | 2003-09-18 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal Structure of Plasmodium Berghei Lactate Dehydrogenase Indicates the Unique Structural Differences of These Enzymes are Shared Across the Plasmodium Genus
Mol.Biochem.Parasitol., 131, 2003
|
|
1QOU
 
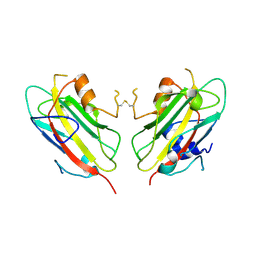 | |
4ZQF
 
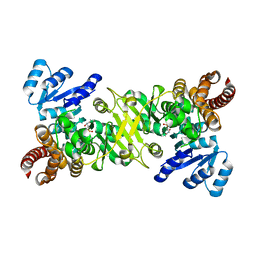 | |
2PET
 
 | | Lutheran glycoprotein, N-terminal domains 1 and 2. | | Descriptor: | Lutheran blood group glycoprotein | | Authors: | Burton, N, Brady, R.L. | | Deposit date: | 2007-04-03 | | Release date: | 2007-12-04 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | The Laminin 511/521-binding site on the Lutheran blood group glycoprotein is located at the flexible junction of Ig domains 2 and 3.
Blood, 110, 2007
|
|
2PF6
 
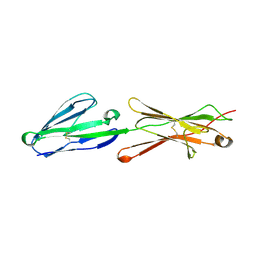 | | Lutheran glycoprotein, N-terminal domains 1 and 2 | | Descriptor: | Lutheran blood group glycoprotein | | Authors: | Burton, N, Brady, R.L. | | Deposit date: | 2007-04-04 | | Release date: | 2007-12-04 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The Laminin 511/521-binding site on the Lutheran blood group glycoprotein is located at the flexible junction of Ig domains 2 and 3.
Blood, 110, 2007
|
|
4ZQG
 
 | |
4ZQE
 
 | |
4ZQH
 
 | |
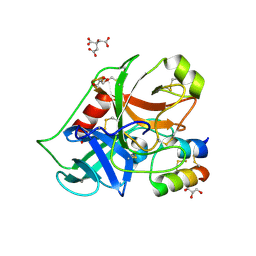
2PLX
 | |
2QYQ
 
 | |
2QIH
 | |
2X8L
 
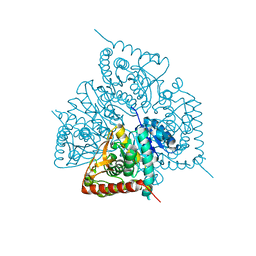 | |
1CET
 
 | | CHLOROQUINE BINDS IN THE COFACTOR BINDING SITE OF PLASMODIUM FALCIPARUM LACTATE DEHYDROGENASE. | | Descriptor: | N4-(7-CHLORO-QUINOLIN-4-YL)-N1,N1-DIETHYL-PENTANE-1,4-DIAMINE, PROTEIN (L-LACTATE DEHYDROGENASE) | | Authors: | Read, J.A, Wilkinson, K.W, Tranter, R, Sessions, R.B, Brady, R.L. | | Deposit date: | 1999-03-10 | | Release date: | 1999-03-19 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Chloroquine binds in the cofactor binding site of Plasmodium falciparum lactate dehydrogenase.
J.Biol.Chem., 274, 1999
|
|
1CEQ
 
 | | CHLOROQUINE BINDS IN THE COFACTOR BINDING SITE OF PLASMODIUM FALCIPARUM LACTATE DEHYDROGENASE. | | Descriptor: | PROTEIN (L-LACTATE DEHYDROGENASE) | | Authors: | Read, J.A, Wilkinson, K.W, Tranter, R, Sessions, R.B, Brady, R.L. | | Deposit date: | 1999-03-10 | | Release date: | 1999-03-19 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Chloroquine binds in the cofactor binding site of Plasmodium falciparum lactate dehydrogenase.
J.Biol.Chem., 274, 1999
|
|
1CLO
 
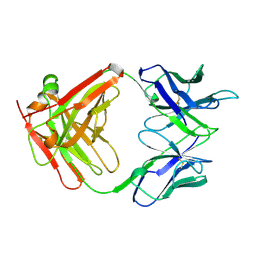 | |
6ZT1
 | |
7A1T
 
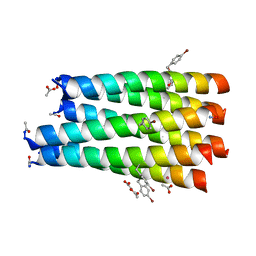 | | A collapsed hexameric state of a de novo coiled-coil assembly: CC-Type2-(GgLaId)4-W19BrPhe. | | Descriptor: | ACETATE ION, CADMIUM ION, CC-Type2-(GgLaId)4-W19BrPhe, ... | | Authors: | Rhys, G.G, Brady, R.L, Woolfson, D.N. | | Deposit date: | 2020-08-14 | | Release date: | 2021-05-19 | | Last modified: | 2021-11-17 | | Method: | X-RAY DIFFRACTION (1.42 Å) | | Cite: | Coiled coils 9-to-5: rational de novo design of alpha-helical barrels with tunable oligomeric states.
Chem Sci, 12, 2021
|
|
1WKO
 
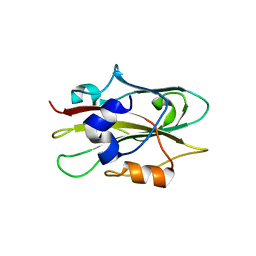 | |
1WKP
 
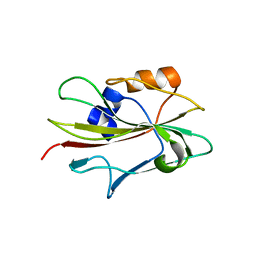 | | Flowering locus t (ft) from arabidopsis thaliana | | Descriptor: | FLOWERING LOCUS T protein, SULFATE ION | | Authors: | Miller, D, Banfield, M.J, Winter, V.J, Brady, R.L. | | Deposit date: | 2004-06-01 | | Release date: | 2005-06-28 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | A divergent external loop confers antagonistic activity on floral regulators FT and TFL1.
Embo J., 25, 2006
|
|
1XIV
 
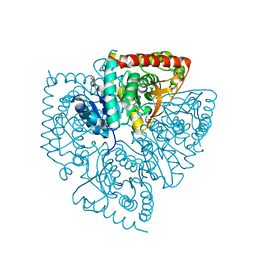 | | Plasmodium falciparum lactate dehydrogenase complexed with 2-({4-chloro-[hydroxy(methoxy)methyl]cyclohexyl}amino)ethane-1,1,2-triol | | Descriptor: | 2-({4-CHLORO-2-[HYDROXY(METHOXY)METHYL]CYCLOHEXYL}AMINO)ETHANE-1,1,2-TRIOL, GLYCEROL, L-lactate dehydrogenase | | Authors: | Conners, R, Cameron, A, Read, J.A, Schambach, F, Sessions, R.B, Brady, R.L. | | Deposit date: | 2004-09-22 | | Release date: | 2005-06-21 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Mapping the binding site for gossypol-like inhibitors of Plasmodium
falciparum lactate dehydrogenase
MOL.BIOCHEM.PARASITOL., 142, 2005
|
|
7BIM
 | | A de novo designed nonameric coiled coil, CC-Type2-(GgLaId)4 | | Descriptor: | GLYCEROL, ISOPROPYL ALCOHOL, Nonameric de novo coiled coil CC-Type2-(GgLaId)4 | | Authors: | Shelley, K.L, Rhys, G.G, Dawson, W.M, Brady, R.L, Woolfson, D.N. | | Deposit date: | 2021-01-12 | | Release date: | 2021-04-21 | | Last modified: | 2021-11-17 | | Method: | X-RAY DIFFRACTION (1.64 Å) | | Cite: | Coiled coils 9-to-5: rational de novo design of alpha-helical barrels with tunable oligomeric states.
Chem Sci, 12, 2021
|
|
7BAT
 
 | | A hexameric barrel state of a de novo coiled-coil assembly: CC-Type2-(GgIaId)4 | | Descriptor: | CC-Type2-(GgIaId)4, N-PROPANOL | | Authors: | Martin, F.J.O, Dawson, W.M, Shelley, K, Brady, R.L, Woolfson, D.N. | | Deposit date: | 2020-12-16 | | Release date: | 2021-04-28 | | Last modified: | 2021-11-17 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | Coiled coils 9-to-5: rational de novo design of alpha-helical barrels with tunable oligomeric states.
Chem Sci, 12, 2021
|
|
7BAS
 | | A de novo pentameric coiled-coil assembly: CC-Type2-(TgLaId)4-W19BrPhe. | | Descriptor: | CC-Type2-(TgLaId)4-W19BrPhe. | | Authors: | Martin, F.J.O, Dawson, W.M, Shelley, K, Brady, R.L, Woolfson, D.N. | | Deposit date: | 2020-12-16 | | Release date: | 2021-04-28 | | Last modified: | 2021-11-17 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | Coiled coils 9-to-5: rational de novo design of alpha-helical barrels with tunable oligomeric states.
Chem Sci, 12, 2021
|
|
