6XV0
 
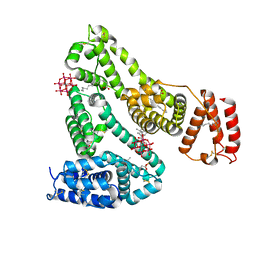 | | lauric acid functionalized hexamolybdoaluminate bound to human serum albumin | | Descriptor: | MYRISTIC ACID, Serum albumin, lauric acid functionalized hexamolybdoaluminate | | Authors: | Bijelic, A, Dobrov, A, Roller, A, Rompel, A. | | Deposit date: | 2020-01-21 | | Release date: | 2020-03-18 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Binding of a Fatty Acid-Functionalized Anderson-Type Polyoxometalate to Human Serum Albumin.
Inorg.Chem., 59, 2020
|
|
5IFO
 
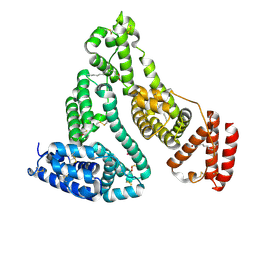 | | X-ray structure of HSA-Myr-KP1019 | | Descriptor: | MYRISTIC ACID, RUTHENIUM ION, Serum albumin | | Authors: | Bijelic, A, Theiner, S, Keppler, B.K, Rompel, A. | | Deposit date: | 2016-02-26 | | Release date: | 2016-06-01 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | X-ray Structure Analysis of Indazolium trans-[Tetrachlorobis(1H-indazole)ruthenate(III)] (KP1019) Bound to Human Serum Albumin Reveals Two Ruthenium Binding Sites and Provides Insights into the Drug Binding Mechanism.
J.Med.Chem., 59, 2016
|
|
5CE9
 
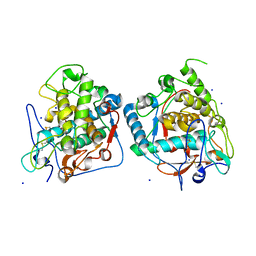 | | structure of tyrosinase from walnut (Juglans regia) | | Descriptor: | COPPER (II) ION, OXYGEN ATOM, Polyphenol oxidase, ... | | Authors: | Bijelic, A, Pretzler, M, Zekiri, F, Rompel, A. | | Deposit date: | 2015-07-06 | | Release date: | 2015-10-28 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The Structure of a Plant Tyrosinase from Walnut Leaves Reveals the Importance of """"Substrate-Guiding Residues"""" for Enzymatic Specificity.
Angew.Chem.Int.Ed.Engl., 54, 2015
|
|
4PHI
 
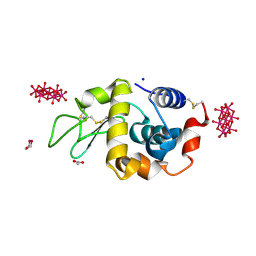 | | Crystal structure of HEWL with hexatungstotellurate(VI) | | Descriptor: | 6-tungstotellurate(VI), ACETATE ION, GLYCEROL, ... | | Authors: | Bijelic, A, Molitor, C, Mauracher, S.G, Al-Oweini, R, Kortz, U, Rompel, A. | | Deposit date: | 2014-05-06 | | Release date: | 2015-01-14 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.811 Å) | | Cite: | Hen Egg-White Lysozyme Crystallisation: Protein Stacking and Structure Stability Enhanced by a Tellurium(VI)-Centred Polyoxotungstate.
Chembiochem, 16, 2015
|
|
9EM2
 
 | |
9EM0
 
 | |
9EM4
 
 | | OPR3 variant Y364P in its dimeric form obtained with ammonium sulfate | | Descriptor: | 12-oxophytodienoate reductase 3, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, FLAVIN MONONUCLEOTIDE, ... | | Authors: | Bijelic, A, Macheroux, P, Kerschbaumer, B. | | Deposit date: | 2024-03-07 | | Release date: | 2024-08-14 | | Last modified: | 2024-08-21 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | Analysis of homodimer formation in 12-oxophytodienoate reductase 3 in solutio and crystallo challenges the physiological role of the dimer.
Sci Rep, 14, 2024
|
|
9FCP
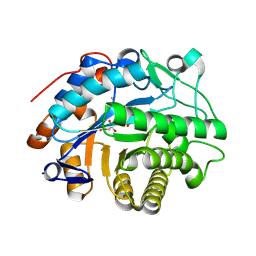 
 | | OPR3 loop swap variant L6(AchrOYE4) in complex with NADH4 | | Descriptor: | 1,4,5,6-Tetrahydronicotinamide adenine dinucleotide, 12-oxophytodienoate reductase 3,Old yellow enzyme OYE4, FLAVIN MONONUCLEOTIDE | | Authors: | Bijelic, A, Kerschbaumer, B, Macheroux, P. | | Deposit date: | 2024-05-15 | | Release date: | 2025-05-28 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Insights into the complexity of coenzyme specificity in ene-reductases
To Be Published
|
|
9FCN
 
 | | OPR3 loop swap variant L6(AchrOYE4) in complex with NADPH4 | | Descriptor: | 1,4,5,6-TETRAHYDRONICOTINAMIDE ADENINE DINUCLEOTIDE PHOSPHATE, 12-oxophytodienoate reductase 3,Old yellow enzyme OYE4, FLAVIN MONONUCLEOTIDE | | Authors: | Bijelic, A, Kerschbaumer, B, Macheroux, P. | | Deposit date: | 2024-05-15 | | Release date: | 2025-05-28 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Insights into the complexity of coenzyme specificity in ene-reductases
To Be Published
|
|
9EM3
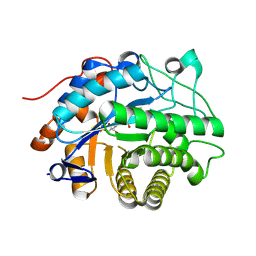 
 | | OPR3 wild type in its monomeric form | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 12-oxophytodienoate reductase 3, FLAVIN MONONUCLEOTIDE | | Authors: | Bijelic, A, Macheroux, P, Kerschbaumer, B. | | Deposit date: | 2024-03-07 | | Release date: | 2024-08-14 | | Last modified: | 2024-08-21 | | Method: | X-RAY DIFFRACTION (2.27 Å) | | Cite: | Analysis of homodimer formation in 12-oxophytodienoate reductase 3 in solutio and crystallo challenges the physiological role of the dimer.
Sci Rep, 14, 2024
|
|
9EM6
 
 | |
9EM5
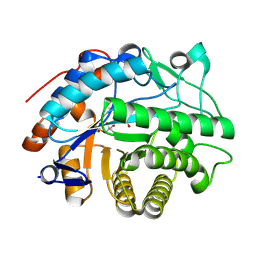 
 | | OPR3 variant Y364P in its monomeric form | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 12-oxophytodienoate reductase 3, CHLORIDE ION, ... | | Authors: | Bijelic, A, Macheroux, P, Kerschbaumer, B. | | Deposit date: | 2024-03-07 | | Release date: | 2024-08-14 | | Last modified: | 2024-08-21 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Analysis of homodimer formation in 12-oxophytodienoate reductase 3 in solutio and crystallo challenges the physiological role of the dimer.
Sci Rep, 14, 2024
|
|
8QN9
 
 | | OPR3 variant - R283E | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 12-oxophytodienoate reductase 3, FLAVIN MONONUCLEOTIDE, ... | | Authors: | Bijelic, A, Macheroux, P, Kerschbaumer, B. | | Deposit date: | 2023-09-26 | | Release date: | 2024-08-14 | | Last modified: | 2024-08-21 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Analysis of homodimer formation in 12-oxophytodienoate reductase 3 in solutio and crystallo challenges the physiological role of the dimer.
Sci Rep, 14, 2024
|
|
8QN1
 
 | | OPR3 variant - R283D | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 12-oxophytodienoate reductase 3, FLAVIN MONONUCLEOTIDE | | Authors: | Bijelic, A, Macheroux, P, Kerschbaumer, B. | | Deposit date: | 2023-09-25 | | Release date: | 2024-08-14 | | Last modified: | 2024-08-21 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Analysis of homodimer formation in 12-oxophytodienoate reductase 3 in solutio and crystallo challenges the physiological role of the dimer.
Sci Rep, 14, 2024
|
|
8QNE
 
 | |
8QNX
 
 | | OPR3 variant with redesigned loop 6 (8aa) in complex with NADH4 | | Descriptor: | 1,4,5,6-Tetrahydronicotinamide adenine dinucleotide, 12-oxophytodienoate reductase 3, FLAVIN MONONUCLEOTIDE, ... | | Authors: | Bijelic, A, Macheroux, P, Kerschbaumer, B. | | Deposit date: | 2023-09-27 | | Release date: | 2025-04-09 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Crystal structures of OPR3 wildtype and variants in complex with NAD(P)H
To Be Published
|
|
8QNK
 
 | |
8QNW
 
 | |
8QO7
 
 | |
8QO8
 
 | | OPR3 variant R283E in complex with NADH4 | | Descriptor: | 1,4,5,6-Tetrahydronicotinamide adenine dinucleotide, 12-oxophytodienoate reductase 3, FLAVIN MONONUCLEOTIDE | | Authors: | Bijelic, A, Macheroux, P, Kerschbaumer, B. | | Deposit date: | 2023-09-28 | | Release date: | 2025-04-09 | | Method: | X-RAY DIFFRACTION (1.47 Å) | | Cite: | Crystal structures of OPR3 wildtype and variants in complex with NAD(P)H
To Be Published
|
|
8QNP
 
 | | OPR3 with redesigned loop 6 (9aa) in complex with NADH4 | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 1,4,5,6-Tetrahydronicotinamide adenine dinucleotide, 12-oxophytodienoate reductase 3, ... | | Authors: | Bijelic, A, Macheroux, P, Kerschbaumer, B. | | Deposit date: | 2023-09-27 | | Release date: | 2025-04-09 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structures of OPR3 wildtype and variants with NAD(P)H
To Be Published
|
|
8QO6
 
 | | OPR3 variant R283D in complex with NADH4 | | Descriptor: | 1,4,5,6-Tetrahydronicotinamide adenine dinucleotide, 12-oxophytodienoate reductase 3, FLAVIN MONONUCLEOTIDE | | Authors: | Bijelic, A, Macheroux, P, Kerschbaumer, B. | | Deposit date: | 2023-09-28 | | Release date: | 2025-04-09 | | Method: | X-RAY DIFFRACTION (2.37 Å) | | Cite: | Crystal structures of OPR3 wildtype and variants in complex with NAD(P)H
To Be Published
|
|
8QNY
 
 | | OPR3 variant with redesigned loop 6 (8aa) in complex with NADPH4 | | Descriptor: | 12-oxophytodienoate reductase 3, FLAVIN MONONUCLEOTIDE, [[(2R,3S,4R,5R)-5-(5-aminocarbonyl-3,4-dihydro-2H-pyridin-1-yl)-3,4-bis(oxidanyl)oxolan-2-yl]methoxy-oxidanyl-phosphoryl] [(2R,3R,4R,5R)-5-(6-aminopurin-9-yl)-3-oxidanyl-4-phosphonooxy-oxolan-2-yl]methyl hydrogen phosphate | | Authors: | Bijelic, A, Macheroux, P, Kerschbaumer, B. | | Deposit date: | 2023-09-27 | | Release date: | 2025-04-09 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Crystal structures of OPR3 wildtype and variants in complex with NAD(P)H
To Be Published
|
|
8QNM
 
 | | OPR3 with redesigned Loop 6 (9aa) in complex with NADPH4 | | Descriptor: | 12-oxophytodienoate reductase 3, FLAVIN MONONUCLEOTIDE, [[(2R,3S,4R,5R)-5-(5-aminocarbonyl-3,4-dihydro-2H-pyridin-1-yl)-3,4-bis(oxidanyl)oxolan-2-yl]methoxy-oxidanyl-phosphoryl] [(2R,3R,4R,5R)-5-(6-aminopurin-9-yl)-3-oxidanyl-4-phosphonooxy-oxolan-2-yl]methyl hydrogen phosphate | | Authors: | Bijelic, A, Macheroux, P, Kerschbaumer, B. | | Deposit date: | 2023-09-27 | | Release date: | 2025-04-09 | | Method: | X-RAY DIFFRACTION (2.08 Å) | | Cite: | Crystal structures of OPR3 wildtype and variants in complex with NAD(P)H
To Be Published
|
|
8QNA
 
 | | OPR3 variant - R366A | | Descriptor: | 12-oxophytodienoate reductase 3, FLAVIN MONONUCLEOTIDE, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | Bijelic, A, Macheroux, P, Kerschbaumer, B. | | Deposit date: | 2023-09-26 | | Release date: | 2025-04-09 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Crystal structures of OPR3 wildtype and variants in complex with NAD(P)H
To Be Published
|
|
