3X17
 
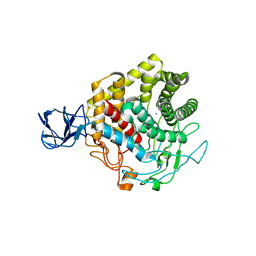 | | Crystal structure of metagenome-derived glycoside hydrolase family 9 endoglucanase | | Descriptor: | CALCIUM ION, Endoglucanase, ZINC ION | | Authors: | Okano, H, Angkawidjaja, C, Kanaya, S. | | Deposit date: | 2014-10-30 | | Release date: | 2015-01-21 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Structure, activity, and stability of metagenome-derived glycoside hydrolase family 9 endoglucanase with an N-terminal Ig-like domain.
Protein Sci., 24, 2015
|
|
3U3G
 
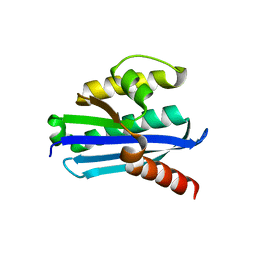 | | Structure of LC11-RNase H1 Isolated from Compost by Metagenomic Approach: Insight into the Structural Bases for Unusual Enzymatic Properties of Sto-RNase H1 | | Descriptor: | CHLORIDE ION, Ribonuclease H, UNKNOWN LIGAND | | Authors: | Nguyen, T.N, Angkawidjaja, C, Kanaya, E, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2011-10-05 | | Release date: | 2012-03-07 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Activity, stability, and structure of metagenome-derived LC11-RNase H1, a homolog of Sulfolobus tokodaii RNase H1
Protein Sci., 21, 2012
|
|
3VN5
 
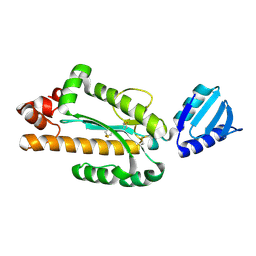 | | Crystal structure of Aquifex aeolicus RNase H3 | | Descriptor: | Ribonuclease HIII | | Authors: | Jongruja, N, You, D.J, Eiko, K, Angkawidjaja, C, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2011-12-22 | | Release date: | 2012-12-26 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | Structure and characterization of RNase H3 from Aquifex aeolicus
Febs J., 279, 2012
|
|
3WX5
 
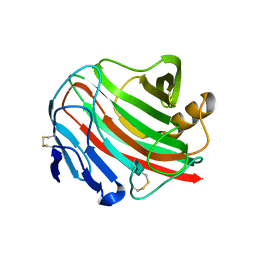 | |
3WIU
 
 | |
3WIV
 
 | |
3WX9
 
 | | Crystal structure of Pyrococcus horikoshii kynurenine aminotransferase in complex with PMP, GLA, 4AD, 2OG, GLU and KYA | | Descriptor: | (2E)-pent-2-enedioic acid, 2-OXOGLUTARIC ACID, 4'-DEOXY-4'-AMINOPYRIDOXAL-5'-PHOSPHATE, ... | | Authors: | Okada, K, Angkawidjaja, C, Koga, Y, Kanaya, S. | | Deposit date: | 2014-07-28 | | Release date: | 2014-09-24 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.58 Å) | | Cite: | Crystal structure of Pyrococcus horikoshii kynurenine aminotransferase in complex with PMP, GLA, 4AD, 2OG, GLU and KYA
To be Published
|
|
3B09
 
 | | Crystal structure of the N-domain of FKBP22 from Shewanella sp. SIB1 | | Descriptor: | Peptidyl-prolyl cis-trans isomerase | | Authors: | Budiman, C, Angkawidjaja, C, Motoike, H, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2011-06-07 | | Release date: | 2012-04-18 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of N-domain of FKBP22 from Shewanella sp. SIB1: dimer dissociation by disruption of Val-Leu knot
Protein Sci., 20, 2011
|
|
3AFG
 
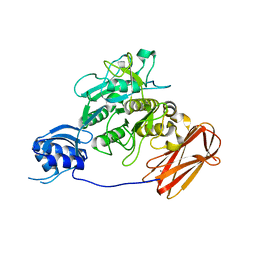 | | Crystal structure of ProN-Tk-SP from Thermococcus kodakaraensis | | Descriptor: | CALCIUM ION, Subtilisin-like serine protease | | Authors: | Foophow, T, Tanaka, S, Angkawidjaja, C, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2010-03-01 | | Release date: | 2010-08-04 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of a subtilisin homologue, Tk-SP, from Thermococcus kodakaraensis: requirement of a C-terminal beta-jelly roll domain for hyperstability.
J.Mol.Biol., 400, 2010
|
|
3AHP
 
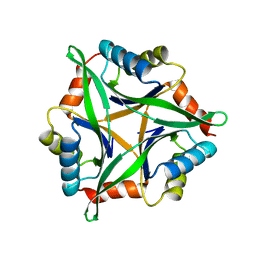 | | Crystal structure of stable protein, CutA1, from a psychrotrophic bacterium Shewanella sp. SIB1 | | Descriptor: | CutA1 | | Authors: | Tanaka, S, Angkawidjaja, C, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2010-04-26 | | Release date: | 2011-01-05 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal structure of stable protein CutA1 from psychrotrophic bacterium Shewanella sp. SIB1
J.SYNCHROTRON RADIAT., 18, 2011
|
|
3AOW
 
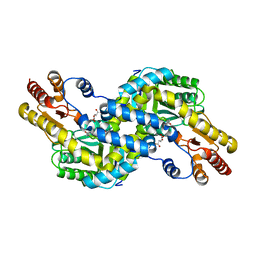 | | Crystal structure of Pyrococcus horikoshii kynurenine aminotransferase in complex with AKG | | Descriptor: | 2-OXOGLUTARIC ACID, PYRIDOXAL-5'-PHOSPHATE, Putative uncharacterized protein PH0207 | | Authors: | Okada, K, Angkawidjaja, C, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2010-10-07 | | Release date: | 2011-10-12 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.56 Å) | | Cite: | Characterization of kynurenine aminotransferase from hyperthermophilic archaeon: enzymatic activity for conversion to kynurenic acid is allosterically regulated by alpha-ketoglutaric acid cooperatively with kynurenine
To be Published
|
|
3AOV
 
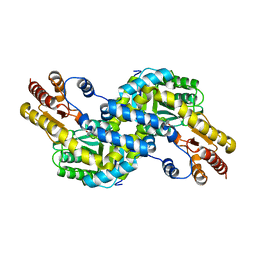 | | Crystal structure of Pyrococcus horikoshii kynurenine aminotransferase in complex with PLP | | Descriptor: | PYRIDOXAL-5'-PHOSPHATE, Putative uncharacterized protein PH0207 | | Authors: | Okada, K, Angkawidjaja, C, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2010-10-07 | | Release date: | 2011-10-12 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.72 Å) | | Cite: | Characterization of kynurenine aminotransferase from hyperthermophilic archaeon: enzymatic activity for conversion to kynurenic acid is allosterically regulated by alpha-ketoglutaric acid cooperatively with kynurenine
To be Published
|
|
3ATH
 
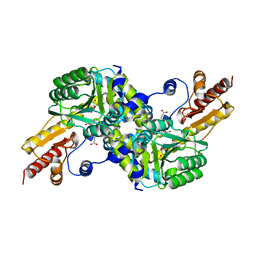 | | Crystal structure of Pyrococcus horikoshii kynurenine aminotransferase in complex with four AKGs as substrates and allosteric effectors | | Descriptor: | 2-OXOGLUTARIC ACID, PYRIDOXAL-5'-PHOSPHATE, Putative uncharacterized protein PH0207 | | Authors: | Okada, K, Angkawidjaja, C, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2011-01-05 | | Release date: | 2012-01-18 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.69 Å) | | Cite: | Characterization of kynurenine aminotransferase from hyperthermophilic archaeon: enzymatic activity for conversion to kynurenic acid is allosterically regulated by alpha-ketoglutaric acid cooperatively with kynurenine
To be Published
|
|
3AV7
 
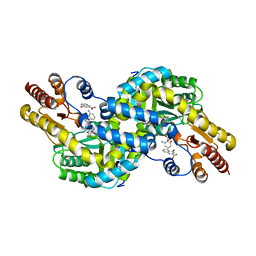 | | Crystal structure of Pyrococcus horikoshii kynurenine aminotransferase in complex with PMP, KYN as substrates and KYA as products | | Descriptor: | (2S)-2-amino-4-(2-aminophenyl)-4-oxobutanoic acid, 4'-DEOXY-4'-AMINOPYRIDOXAL-5'-PHOSPHATE, 4-hydroxyquinoline-2-carboxylic acid, ... | | Authors: | Okada, K, Angkawidjaja, C, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2011-02-23 | | Release date: | 2012-03-07 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.84 Å) | | Cite: | Structural insight into kynurenic acid excretion mechanisms of kynurenine aminotransferase in the hyperthermophilic archaeon Pyrococcus horikoshii
To be Published
|
|
4MAD
 
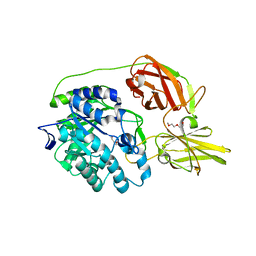 | | Crystal structure of beta-galactosidase C (BgaC) from Bacillus circulans ATCC 31382 | | Descriptor: | 2-(2-METHOXYETHOXY)ETHANOL, Beta-galactosidase | | Authors: | Kamerke, C, You, D.J, Kanaya, S, Elling, L. | | Deposit date: | 2013-08-16 | | Release date: | 2014-08-27 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Rational design of a glycosynthase by the crystal structure of beta-galactosidase from Bacillus circulans (BgaC) and its use for the synthesis of N-acetyllactosamine type 1 glycan structures.
J.Biotechnol., 191, 2014
|
|
5NEN
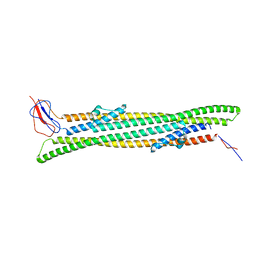 
 | |
3AA4
 
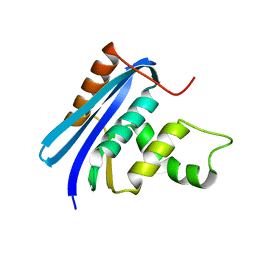 | | A52V E.coli RNase HI | | Descriptor: | Ribonuclease HI | | Authors: | Takano, K. | | Deposit date: | 2009-11-11 | | Release date: | 2010-10-06 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | Protein core adaptability: crystal structures of the cavity-filling variants of Escherichia coli RNase HI
Protein Pept.Lett., 17, 2010
|
|
3AA3
 
 | | A52L E. coli RNase HI | | Descriptor: | Ribonuclease HI | | Authors: | Takano, K. | | Deposit date: | 2009-11-11 | | Release date: | 2010-10-06 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Protein core adaptability: crystal structures of the cavity-filling variants of Escherichia coli RNase HI
Protein Pept.Lett., 17, 2010
|
|
3AA2
 
 | | A52I E. coli RNase HI | | Descriptor: | Ribonuclease HI | | Authors: | Takano, K. | | Deposit date: | 2009-11-11 | | Release date: | 2010-10-06 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Protein core adaptability: crystal structures of the cavity-filling variants of Escherichia coli RNase HI
Protein Pept.Lett., 17, 2010
|
|
3AA5
 
 | | A52F E.coli RNase HI | | Descriptor: | Ribonuclease HI | | Authors: | Takano, K. | | Deposit date: | 2009-11-11 | | Release date: | 2010-10-06 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Protein core adaptability: crystal structures of the cavity-filling variants of Escherichia coli RNase HI
Protein Pept.Lett., 17, 2010
|
|
