1LWR
 
 | | Solution structure of the NCAM fibronectin type III module 2 | | Descriptor: | Neural Cell Adhesion Molecule 1, 140 kDa isoform | | Authors: | Kiselyov, V.V, Skladchikova, G, Hinsby, A.M, Jensen, P.H, Kulahin, N, Pedersen, N, Tsetlin, V, Poulsen, F.M, Berezin, V, Bock, E. | | Deposit date: | 2002-06-03 | | Release date: | 2003-06-03 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structural basis for a direct interaction between FGFR1 and NCAM and evidence for a regulatory role of ATP
Structure, 11, 2003
|
|
3NCM
 
 | | NEURAL CELL ADHESION MOLECULE, MODULE 2, NMR, 20 STRUCTURES | | Descriptor: | PROTEIN (NEURAL CELL ADHESION MOLECULE, LARGE ISOFORM) | | Authors: | Jensen, P.H, Soroka, V, Thomsen, N.K, Berezin, V, Bock, E, Poulsen, F.M. | | Deposit date: | 1998-09-21 | | Release date: | 1999-07-23 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Structure and interactions of NCAM modules 1 and 2, basic elements in neural cell adhesion
Nat.Struct.Biol., 6, 1999
|
|
2LCP
 
 | | NMR structure of calcium loaded, un-myristoylated human NCS-1 | | Descriptor: | CALCIUM ION, Neuronal calcium sensor 1 | | Authors: | Heidarsson, P.O, Bjerrum-Bohr, I.J, Bellucci, L, Gitte, J, Corni, S, Poulsen, F.M, Finn, B.E, Di Felice, R, Kragelund, B. | | Deposit date: | 2011-05-05 | | Release date: | 2012-02-01 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | The C-terminal tail of human neuronal calcium sensor 1 regulates the conformational stability of the ca(2+)-activated state.
J.Mol.Biol., 417, 2012
|
|
2LUI
 
 | | Structure of the PICK PDZ domain in complex with the DAT C-terminal | | Descriptor: | PICK1 PDZ DOMAIN FUSED TO THE C10 DAT LIGAND | | Authors: | Erlendsson, S, Rathje, M, Heidarsson, P.O, Poulsen, F.M, Madsen, K.L, Teilum, K, Gether, U. | | Deposit date: | 2012-06-14 | | Release date: | 2013-06-19 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Protein interacting with C-kinase 1 (PICK1) binding promiscuity relies on unconventional PSD-95/discs-large/ZO-1 homology (PDZ) binding modes for nonclass II PDZ ligands.
J.Biol.Chem., 289, 2014
|
|
2LVS
 
 | |
2NCM
 
 | |
2DEZ
 
 | | Structure of human PYY | | Descriptor: | Peptide YY | | Authors: | Nygaard, R. | | Deposit date: | 2006-02-20 | | Release date: | 2006-07-18 | | Last modified: | 2022-03-09 | | Method: | SOLUTION NMR | | Cite: | The PP-Fold Solution Structure of Human Polypeptide YY and Human PYY3-36 As Determined by NMR(,)
Biochemistry, 45, 2006
|
|
2DF0
 
 | | Solution structure of human PYY3-36 | | Descriptor: | Peptide YY | | Authors: | Nygaard, R. | | Deposit date: | 2006-02-20 | | Release date: | 2006-07-18 | | Last modified: | 2022-03-09 | | Method: | SOLUTION NMR | | Cite: | The PP-Fold Solution Structure of Human Polypeptide YY and Human PYY3-36 As Determined by NMR(,)
Biochemistry, 45, 2006
|
|
2QIM
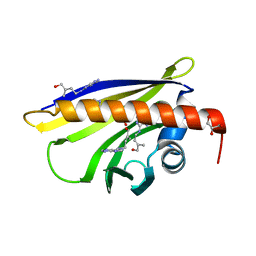 
 | | Crystal Structure of Pathogenesis-related Protein LlPR-10.2B from yellow lupine in complex with Cytokinin | | Descriptor: | (2E)-2-methyl-4-(9H-purin-6-ylamino)but-2-en-1-ol, CALCIUM ION, GLYCEROL, ... | | Authors: | Fernandes, H.C, Pasternak, O, Bujacz, G, Bujacz, A, Sikorski, M.M, Jaskolski, M. | | Deposit date: | 2007-07-05 | | Release date: | 2008-04-29 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Lupinus luteus pathogenesis-related protein as a reservoir for cytokinin.
J.Mol.Biol., 378, 2008
|
|
1XDF
 
 | | Crystal structure of pathogenesis-related protein LlPR-10.2A from yellow lupine | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, PR10.2A, SODIUM ION | | Authors: | Pasternak, O, Biesiadka, J, Dolot, R, Handschuh, L, Bujacz, G, Sikorski, M.M, Jaskolski, M. | | Deposit date: | 2004-09-06 | | Release date: | 2005-02-15 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure of a yellow lupin pathogenesis-related PR-10 protein belonging to a novel subclass.
Acta Crystallogr.,Sect.D, 61, 2005
|
|
1BNR
 
 | | BARNASE | | Descriptor: | BARNASE (G SPECIFIC ENDONUCLEASE) | | Authors: | Bycroft, M. | | Deposit date: | 1995-03-31 | | Release date: | 1995-07-31 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Determination of the three-dimensional solution structure of barnase using nuclear magnetic resonance spectroscopy.
Biochemistry, 30, 1991
|
|
2CI2
 
 | |
1LIP
 
 | |
2FLH
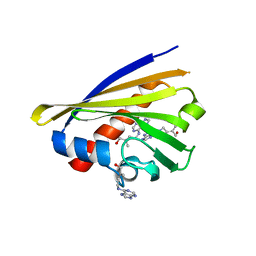 
 | | Crystal structure of cytokinin-specific binding protein from mung bean in complex with cytokinin | | Descriptor: | (2E)-2-methyl-4-(9H-purin-6-ylamino)but-2-en-1-ol, SODIUM ION, cytokinin-specific binding protein | | Authors: | Pasternak, O, Bujacz, G.D, Sikorski, M.M, Jaskolski, M. | | Deposit date: | 2006-01-06 | | Release date: | 2006-11-21 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Crystal Structure of Vigna radiata Cytokinin-Specific Binding Protein in Complex with Zeatin.
Plant Cell, 18, 2006
|
|
3E85
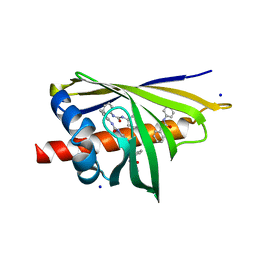 
 | | Crystal Structure of Pathogenesis-related Protein LlPR-10.2B from yellow lupine in complex with Diphenylurea | | Descriptor: | 1,3-DIPHENYLUREA, PR10.2B, SODIUM ION | | Authors: | Fernandes, H.C, Bujacz, G, Bujacz, A, Sikorski, M.M, Jaskolski, M. | | Deposit date: | 2008-08-19 | | Release date: | 2009-03-03 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Cytokinin-induced structural adaptability of a Lupinus luteus PR-10 protein.
Febs J., 276, 2009
|
|
1CQ4
 
 | | CI2 MUTANT WITH TETRAGLUTAMINE (MGQQQQGM) REPLACING MET59 | | Descriptor: | PROTEIN (SERINE PROTEINASE INHIBITOR 2), SULFATE ION | | Authors: | Chen, Y.W, Stott, K.R. | | Deposit date: | 1998-11-17 | | Release date: | 1998-11-25 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of a dimeric chymotrypsin inhibitor 2 mutant containing an inserted glutamine repeat.
Proc.Natl.Acad.Sci.USA, 96, 1999
|
|
1IFV
 
 | |
3IE5
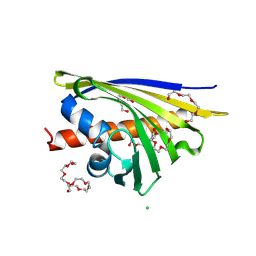 
 | | Crystal structure of Hyp-1 protein from Hypericum perforatum (St John's wort) involved in hypericin biosynthesis | | Descriptor: | 3,6,9,12,15,18,21-HEPTAOXATRICOSANE-1,23-DIOL, CHLORIDE ION, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Michalska, K, Fernandes, H, Sikorski, M.M, Jaskolski, M. | | Deposit date: | 2009-07-22 | | Release date: | 2009-11-10 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.688 Å) | | Cite: | Crystal structure of Hyp-1, a St. John's wort protein implicated in the biosynthesis of hypericin
J.Struct.Biol., 169, 2010
|
|
1ICX
 
 | |
1QMR
 
 | | BIRCH POLLEN ALLERGEN BET V 1 MUTANT N28T, K32Q, E45S, P108G | | Descriptor: | MAJOR POLLEN ALLERGEN BET V 1-A | | Authors: | Henriksen, A, Holm, J.O, Spangfort, M.D, Gajhede, M. | | Deposit date: | 1999-10-06 | | Release date: | 2000-10-12 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Allergy vaccine engineering: epitope modulation of recombinant Bet v 1 reduces IgE binding but retains protein folding pattern for induction of protective blocking-antibody responses.
J Immunol., 173, 2004
|
|
