1SQR
 
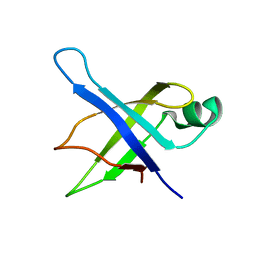 | | Solution Structure of the 50S Ribosomal Protein L35AE from Pyrococcus furiosus. Northeast Structural Genomics Consortium Target PfR48. | | 分子名称: | 50S ribosomal protein L35Ae | | 著者 | Snyder, D.A, Aramini, J.M, Huang, Y.J, Xiao, R, Cort, J.R, Shastry, R, Ma, L.C, Liu, J, Rost, B, Acton, T.B, Kennedy, M.A, Montelione, G.T, Northeast Structural Genomics Consortium (NESG) | | 登録日 | 2004-03-19 | | 公開日 | 2004-11-16 | | 最終更新日 | 2024-05-22 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Solution Structure of the 50S Ribosomal Protein L35AE from Pyrococcus furiosus: Northeast Structural Genomics Consortium Target PfR48
To be Published
|
|
1SOU
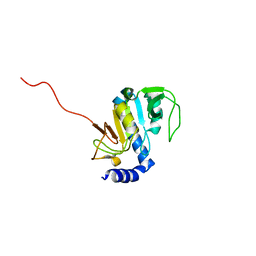 
 | | NMR structure of Aquifex aeolicus 5,10-methenyltetrahydrofolate synthetase: Northeast Structural Genomics Consortium Target QR46 | | 分子名称: | 5,10-methenyltetrahydrofolate synthetase | | 著者 | Cort, J.R, Chiang, Y, Acton, T, Wu, M, Montelione, G.T, Kennedy, M.A, Northeast Structural Genomics Consortium (NESG) | | 登録日 | 2004-03-15 | | 公開日 | 2004-06-22 | | 最終更新日 | 2024-05-22 | | 実験手法 | SOLUTION NMR | | 主引用文献 | NMR structure of Aquifex aeolicus 5,10-methenyltetrahydrofolate synthetase: Northeast Structural Genomics Consortium Target QR46
To be Published
|
|
1SSV
 
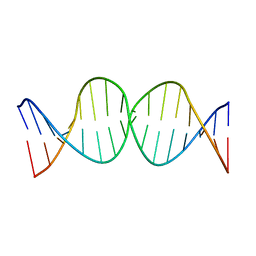 | | Compensating bends in a 16 base-pair DNA oligomer containing a T3A3 segment | | 分子名称: | 5'-D(*CP*GP*AP*GP*GP*TP*TP*TP*AP*AP*AP*CP*CP*TP*CP*G)-3' | | 著者 | McAteer, K, Aceves-Gaona, A, Michalczyk, R, Buchko, G.W, Isern, N.G, Silks, L.A, Miller, J.H, Kennedy, M.A. | | 登録日 | 2004-03-24 | | 公開日 | 2004-12-07 | | 最終更新日 | 2024-05-22 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Compensating bends in a 16-base-pair DNA oligomer containing a T(3)A(3) segment: A NMR study of global DNA curvature
Biopolymers, 75, 2004
|
|
1NEI
 
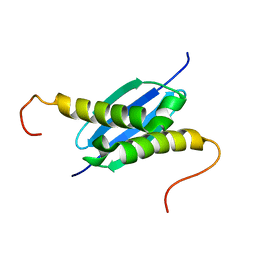 | | Solution NMR Structure of Protein yoaG from Escherichia coli. Ontario Centre for Structural Proteomics Target EC0264_1_60; Northeast Structural Genomics Consortium Target ET94. | | 分子名称: | hypothetical protein yoaG | | 著者 | Wu, B, Pineda-Lucena, A, Yee, A, Cort, J, Kennedy, M.A, Edwards, A.M, Arrowsmith, C.H, Northeast Structural Genomics Consortium (NESG) | | 登録日 | 2002-12-11 | | 公開日 | 2004-04-13 | | 最終更新日 | 2024-05-22 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Solution structure of hypothetical protein dimer encoded by the Yoag gene from Escherichia coli
To be published
|
|
1T0A
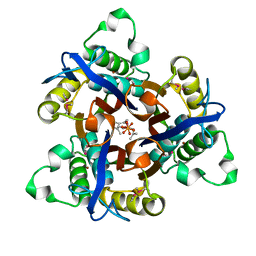 
 | | Crystal Structure of 2C-Methyl-D-Erythritol-2,4-cyclodiphosphate Synthase from Shewanella Oneidensis | | 分子名称: | 2C-methyl-D-erythritol 2,4-cyclodiphosphate synthase, COBALT (II) ION, FARNESYL DIPHOSPHATE, ... | | 著者 | Ni, S, Robinson, H, Marsing, G.C, Bussiere, D.E, Kennedy, M.A. | | 登録日 | 2004-04-08 | | 公開日 | 2004-10-26 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.6 Å) | | 主引用文献 | Structure of 2C-methyl-D-erythritol-2,4-cyclodiphosphate synthase from Shewanella oneidensis at 1.6 A: identification of farnesyl pyrophosphate trapped in a hydrophobic cavity.
Acta Crystallogr.,Sect.D, 60, 2004
|
|
1SS7
 
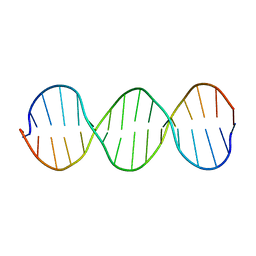 | | Compensating bends in a 16 base-pair DNA oligomer containing a T3A3 segment | | 分子名称: | 5'-D(*CP*GP*AP*GP*GP*TP*TP*TP*AP*AP*AP*CP*CP*TP*CP*G)-3' | | 著者 | McAteer, K, Aceves-Gaona, A, Michalczyk, R, Buchko, G.W, Isern, N.G, Silks, L.A, Miller, J.H, Kennedy, M.A. | | 登録日 | 2004-03-23 | | 公開日 | 2004-12-07 | | 最終更新日 | 2024-05-22 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Compensating bends in a 16-base-pair DNA oligomer containing a T(3)A(3) segment: A NMR study of global DNA curvature
Biopolymers, 75, 2004
|
|
1NY4
 
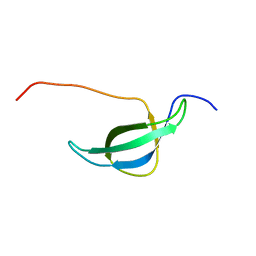 | | Solution structure of the 30S ribosomal protein S28E from Pyrococcus horikoshii. Northeast Structural Genomics Consortium target JR19. | | 分子名称: | 30S ribosomal protein S28E | | 著者 | Aramini, J.M, Cort, J.R, Huang, Y.J, Xiao, R, Acton, T.B, Ho, C.K, Shih, L.-Y, Kennedy, M.A, Montelione, G.T, Northeast Structural Genomics Consortium (NESG) | | 登録日 | 2003-02-11 | | 公開日 | 2003-09-02 | | 最終更新日 | 2024-05-22 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Solution NMR structure of the 30S ribosomal protein S28E from Pyrococcus horikoshii.
Protein Sci., 12, 2003
|
|
1NXI
 
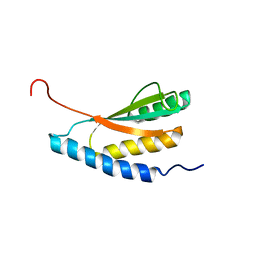 | | Solution structure of Vibrio cholerae protein VC0424 | | 分子名称: | conserved hypothetical protein VC0424 | | 著者 | Ramelot, T.A, Ni, S, Goldsmith-Fischman, S, Cort, J.R, Honig, B, Kennedy, M.A, Northeast Structural Genomics Consortium (NESG) | | 登録日 | 2003-02-10 | | 公開日 | 2003-07-01 | | 最終更新日 | 2024-05-08 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Solution structure of Vibrio cholerae protein VC0424: a variation of the ferredoxin-like fold.
Protein Sci., 12, 2003
|
|
1TTD
 
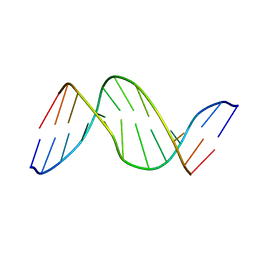 | | SOLUTION-STATE STRUCTURE OF A DNA DODECAMER DUPLEX CONTAINING A CIS-SYN THYMINE CYCLOBUTANE DIMER | | 分子名称: | DNA (5'-D(*CP*TP*TP*AP*AP*TP*TP*CP*GP*TP*GP*C)-3'), DNA (5'-D(*GP*CP*AP*CP*GP*AP*AP*(TTD)P*AP*AP*G)-3') | | 著者 | Mcateer, K, Jing, Y, Kao, J, Taylor, J.-S, Kennedy, M.A. | | 登録日 | 1999-01-20 | | 公開日 | 1999-02-02 | | 最終更新日 | 2023-12-27 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Solution-state structure of a DNA dodecamer duplex containing a Cis-syn thymine cyclobutane dimer, the major UV photoproduct of DNA.
J.Mol.Biol., 282, 1998
|
|
1Q48
 
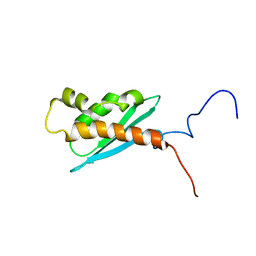 | | Solution NMR Structure of The Haemophilus Influenzae Iron-Sulfur Cluster Assembly Protein U (IscU) with Zinc Bound at the Active Site. Northeast Structural Genomics Consortium Target IR24. This protein is not apo, it is a model without zinc binding constraints. | | 分子名称: | NifU-like protein | | 著者 | Ramelot, T.A, Cort, J.R, Xiao, R, Shastry, R, Acton, T.B, Montelione, G.T, Kennedy, M.A, Northeast Structural Genomics Consortium (NESG) | | 登録日 | 2003-08-01 | | 公開日 | 2003-11-18 | | 最終更新日 | 2024-05-01 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Solution NMR structure of the iron-sulfur cluster assembly protein U (IscU) with zinc bound at the active site.
J.Mol.Biol., 344, 2004
|
|
8E0M
 
 | |
8E12
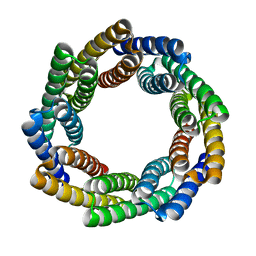 
 | |
8E0N
 
 | |
8E0O
 
 | |
7KHU
 
 | | Crystal structure of the carbohydrate-binding domain VP8* of human P[4] rotavirus strain BM5265 in complex with LNDFH I | | 分子名称: | Outer capsid protein VP4, alpha-L-fucopyranose-(1-2)-beta-D-galactopyranose-(1-3)-[alpha-L-fucopyranose-(1-4)]2-acetamido-2-deoxy-beta-D-glucopyranose-(1-3)-beta-D-galactopyranose-(1-4)-beta-D-glucopyranose | | 著者 | Xu, S, Stuckert, M, Burnside, R, McGinnis, K, Jiang, X, Kennedy, M.A. | | 登録日 | 2020-10-22 | | 公開日 | 2021-08-18 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (2.54 Å) | | 主引用文献 | Structural basis of P[II] rotavirus evolution and host ranges under selection of histo-blood group antigens.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
7KI5
 
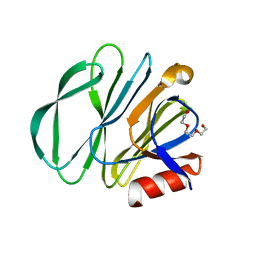 | |
5VKI
 
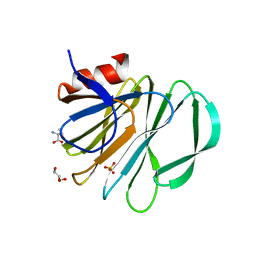 | | Crystal structure of P[19] rotavirus VP8* complexed with mucin core 2 | | 分子名称: | GLYCEROL, Outer capsid protein VP4, SULFATE ION, ... | | 著者 | Xu, S, Liu, Y, Woodruff, A, Zhong, W, Jiang, X, Kennedy, M.A. | | 登録日 | 2017-04-21 | | 公開日 | 2017-11-08 | | 最終更新日 | 2023-10-04 | | 実験手法 | X-RAY DIFFRACTION (1.9 Å) | | 主引用文献 | Structural basis of glycan specificity of P[19] VP8*: Implications for rotavirus zoonosis and evolution.
PLoS Pathog., 13, 2017
|
|
5VKS
 
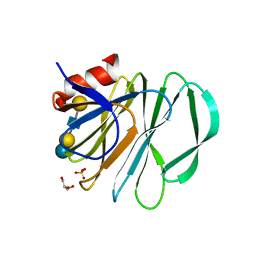 | | Crystal structure of P[19] rotavirus VP8* complexed with LNFPI | | 分子名称: | GLYCEROL, Outer capsid protein VP4, SULFATE ION, ... | | 著者 | Xu, S, Liu, Y, Woodruff, A, Zhong, W, Jiang, X, Kennedy, M.A. | | 登録日 | 2017-04-22 | | 公開日 | 2017-11-08 | | 最終更新日 | 2023-10-04 | | 実験手法 | X-RAY DIFFRACTION (1.94 Å) | | 主引用文献 | Structural basis of glycan specificity of P[19] VP8*: Implications for rotavirus zoonosis and evolution.
PLoS Pathog., 13, 2017
|
|
5VPY
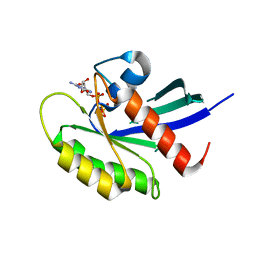 
 | | Crystal structure of human KRAS G12A mutant in complex with GppNHp | | 分子名称: | 2-amino-9-{5-O-[(S)-hydroxy{[(R)-hydroxy(phosphonoamino)phosphoryl]oxy}phosphoryl]-alpha-L-xylofuranosyl}-1,9-dihydro-6H-purin-6-one, GTPase KRas, MAGNESIUM ION, ... | | 著者 | Xu, S, Long, B, Boris, G, Ni, S, Kennedy, M.A. | | 登録日 | 2017-05-06 | | 公開日 | 2017-12-06 | | 最終更新日 | 2024-03-13 | | 実験手法 | X-RAY DIFFRACTION (2 Å) | | 主引用文献 | Structural insight into the rearrangement of the switch I region in GTP-bound G12A K-Ras.
Acta Crystallogr D Struct Biol, 73, 2017
|
|
5VQ2
 
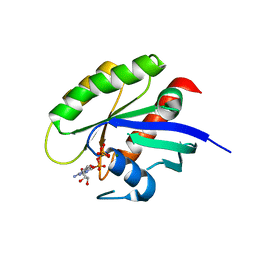 | | Crystal structure of human WT-KRAS in complex with GTP | | 分子名称: | GTPase KRas, GUANOSINE-5'-TRIPHOSPHATE, MAGNESIUM ION | | 著者 | Xu, S, Long, B, Boris, G, Ni, S, Kennedy, M.A. | | 登録日 | 2017-05-07 | | 公開日 | 2017-12-06 | | 最終更新日 | 2017-12-20 | | 実験手法 | X-RAY DIFFRACTION (1.96 Å) | | 主引用文献 | Structural insight into the rearrangement of the switch I region in GTP-bound G12A K-Ras.
Acta Crystallogr D Struct Biol, 73, 2017
|
|
5VQ8
 
 | | Crystal structure of human WT-KRAS in complex with GDP (EDTA soaked) | | 分子名称: | GTPase KRas, GUANOSINE-5'-DIPHOSPHATE, MAGNESIUM ION | | 著者 | Xu, S, Long, B, Boris, G, Ni, S, Kennedy, M.A. | | 登録日 | 2017-05-08 | | 公開日 | 2017-12-06 | | 最終更新日 | 2024-03-13 | | 実験手法 | X-RAY DIFFRACTION (2.3 Å) | | 主引用文献 | Structural insight into the rearrangement of the switch I region in GTP-bound G12A K-Ras.
Acta Crystallogr D Struct Biol, 73, 2017
|
|
5VPI
 
 | | Crystal structure of human KRAS G12A mutant in complex with GTP | | 分子名称: | GTPase KRas, GUANOSINE-5'-TRIPHOSPHATE, MAGNESIUM ION | | 著者 | Xu, S, Long, B, Boris, G, Ni, S, Kennedy, M.A. | | 登録日 | 2017-05-05 | | 公開日 | 2017-12-06 | | 最終更新日 | 2024-03-13 | | 実験手法 | X-RAY DIFFRACTION (1.62 Å) | | 主引用文献 | Structural insight into the rearrangement of the switch I region in GTP-bound G12A K-Ras.
Acta Crystallogr D Struct Biol, 73, 2017
|
|
5W22
 
 | | Crystal structure of human WT-KRAS in complex with GDP | | 分子名称: | GTPase KRas, GUANOSINE-5'-DIPHOSPHATE, MAGNESIUM ION | | 著者 | Xu, S, Long, B, Boris, G, Ni, S, Kennedy, M.A. | | 登録日 | 2017-06-05 | | 公開日 | 2017-12-06 | | 最終更新日 | 2024-03-13 | | 実験手法 | X-RAY DIFFRACTION (1.762 Å) | | 主引用文献 | Structural insight into the rearrangement of the switch I region in GTP-bound G12A K-Ras.
Acta Crystallogr D Struct Biol, 73, 2017
|
|
5VQ1
 
 | | Crystal structure of human KRAS Q61A mutant in complex with GDP | | 分子名称: | GTPase KRas, GUANOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, ... | | 著者 | Xu, S, Long, B, Boris, G, Ni, S, Kennedy, M.A. | | 登録日 | 2017-05-07 | | 公開日 | 2017-12-06 | | 最終更新日 | 2024-03-13 | | 実験手法 | X-RAY DIFFRACTION (1.78 Å) | | 主引用文献 | Structural insight into the rearrangement of the switch I region in GTP-bound G12A K-Ras.
Acta Crystallogr D Struct Biol, 73, 2017
|
|
5VQ0
 
 | | Crystal structure of human KRAS G12A mutant in complex with GDP (EDTA soaked) | | 分子名称: | GTPase KRas, GUANOSINE-5'-DIPHOSPHATE, MAGNESIUM ION | | 著者 | Xu, S, Long, B, Boris, G, Ni, S, Kennedy, M.A. | | 登録日 | 2017-05-06 | | 公開日 | 2017-12-06 | | 最終更新日 | 2024-03-13 | | 実験手法 | X-RAY DIFFRACTION (2.3 Å) | | 主引用文献 | Structural insight into the rearrangement of the switch I region in GTP-bound G12A K-Ras.
Acta Crystallogr D Struct Biol, 73, 2017
|
|
