3H2I
 
 | |
3H2G
 
 | |
3CW3
 
 | | Crystal structure of AIM1g1 | | Descriptor: | Absent in melanoma 1 protein, GLYCEROL | | Authors: | Aravind, P, Sankaranarayanan, R, Sharma, Y. | | Deposit date: | 2008-04-21 | | Release date: | 2008-06-17 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Exploring the limits of sequence and structure in a variant betagamma-crystallin domain of the protein absent in melanoma-1 (AIM1).
J.Mol.Biol., 381, 2008
|
|
3G2R
 
 | |
3HZ2
 
 | |
3HZB
 
 | |
3H2J
 
 | |
2ZLU
 
 | | Horse methemoglobin high salt, pH 7.0 (88% relative humidity) | | Descriptor: | Hemoglobin subunit alpha, Hemoglobin subunit beta, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Kaushal, P.S, Sankaranarayanan, R, Vijayan, M. | | Deposit date: | 2008-04-10 | | Release date: | 2008-06-24 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Water-mediated variability in the structure of relaxed-state haemoglobin
Acta Crystallogr.,Sect.F, 64, 2008
|
|
2ZLX
 
 | | Horse methemoglobin high salt, pH 7.0 (66% relative humidity) | | Descriptor: | Hemoglobin subunit alpha, Hemoglobin subunit beta, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Kaushal, P.S, Sankaranarayanan, R, Vijayan, M. | | Deposit date: | 2008-04-10 | | Release date: | 2008-06-24 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Water-mediated variability in the structure of relaxed-state haemoglobin
Acta Crystallogr.,Sect.F, 64, 2008
|
|
3H2K
 
 | | Crystal structure of a ligand-bound form of the rice cell wall degrading esterase LipA from Xanthomonas oryzae | | Descriptor: | esterase, octyl beta-D-glucopyranoside | | Authors: | Aparna, G, Chatterjee, A, Sonti, R.V, Sankaranarayanan, R. | | Deposit date: | 2009-04-14 | | Release date: | 2009-08-18 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | A Cell Wall-Degrading Esterase of Xanthomonas oryzae Requires a Unique Substrate Recognition Module for Pathogenesis on Rice
Plant Cell, 21, 2009
|
|
3H2H
 
 | |
3HTH
 
 | |
3HTJ
 
 | |
3I9H
 
 | |
3HTA
 
 | |
3HTI
 
 | |
3E1H
 
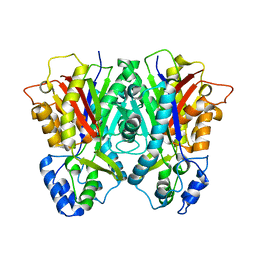 | |
3IAJ
 
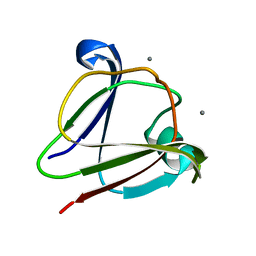 | |
2ZLW
 
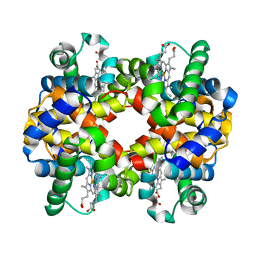 | | Horse methemoglobin high salt, pH 7.0 (75% relative humidity) | | Descriptor: | Hemoglobin subunit alpha, Hemoglobin subunit beta, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Kaushal, P.S, Sankaranarayanan, R, Vijayan, M. | | Deposit date: | 2008-04-10 | | Release date: | 2008-06-24 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Water-mediated variability in the structure of relaxed-state haemoglobin
Acta Crystallogr.,Sect.F, 64, 2008
|
|
2ZLT
 
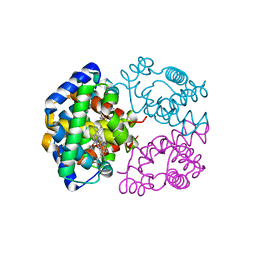 | | Horse methemoglobin high salt, pH 7.0 | | Descriptor: | Hemoglobin subunit alpha, Hemoglobin subunit beta, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Kaushal, P.S, Sankaranarayanan, R, Vijayan, M. | | Deposit date: | 2008-04-10 | | Release date: | 2008-06-24 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Water-mediated variability in the structure of relaxed-state haemoglobin
Acta Crystallogr.,Sect.F, 64, 2008
|
|
2ZLV
 
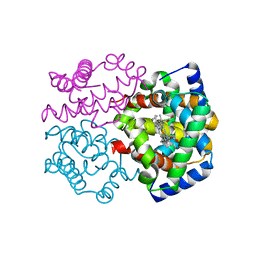 | | Horse methemoglobin high salt, pH 7.0 (79% relative humidity) | | Descriptor: | Hemoglobin subunit alpha, Hemoglobin subunit beta, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Kaushal, P.S, Sankaranarayanan, R, Vijayan, M. | | Deposit date: | 2008-04-10 | | Release date: | 2008-06-24 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Water-mediated variability in the structure of relaxed-state haemoglobin
Acta Crystallogr.,Sect.F, 64, 2008
|
|
3ENT
 
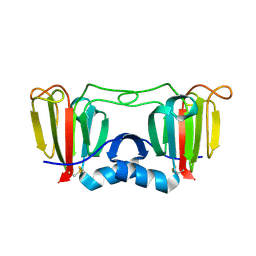 | |
4NSS
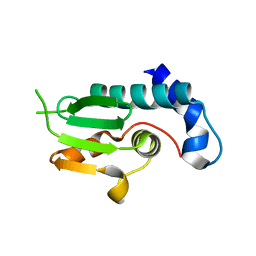 
 | | A structural and functional investigation of a novel protein from Mycobacterium smegmatis implicated in mycobacterial macrophage survivability | | Descriptor: | GLYCEROL, Mycobacterial protein | | Authors: | Shahine, A.E, Littler, D, Brammananath, R, Chan, P.Y, Crellin, P.K, Coppel, R.L, Rossjohn, J, Beddoe, T. | | Deposit date: | 2013-11-28 | | Release date: | 2014-09-10 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | A structural and functional investigation of a novel protein from Mycobacterium smegmatis implicated in mycobacterial macrophage survivability.
Acta Crystallogr.,Sect.D, 70, 2014
|
|
4NBI
 
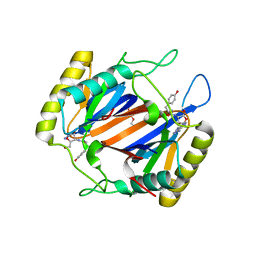 | | D-aminoacyl-tRNA deacylase (DTD) from Plasmodium falciparum in complex with D-tyrosyl-3'-aminoadenosine at 1.86 Angstrom resolution | | Descriptor: | 3'-deoxy-3'-(D-tyrosylamino)adenosine, D-tyrosyl-tRNA(Tyr) deacylase, TRIETHYLENE GLYCOL | | Authors: | Ahmad, S, Routh, S.B, Kamarthapu, V, Sankaranarayanan, R. | | Deposit date: | 2013-10-23 | | Release date: | 2013-12-18 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Mechanism of chiral proofreading during translation of the genetic code.
Elife, 2, 2013
|
|
4NBJ
 
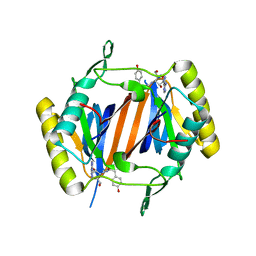 | | D-aminoacyl-tRNA deacylase (DTD) from Plasmodium falciparum in complex with D-tyrosyl-3'-aminoadenosine at 2.20 Angstrom resolution | | Descriptor: | 3'-deoxy-3'-(D-tyrosylamino)adenosine, D-tyrosyl-tRNA(Tyr) deacylase | | Authors: | Ahmad, S, Routh, S.B, Kamarthapu, V, Sankaranarayanan, R. | | Deposit date: | 2013-10-23 | | Release date: | 2013-12-18 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Mechanism of chiral proofreading during translation of the genetic code.
Elife, 2, 2013
|
|
