3TX8
 
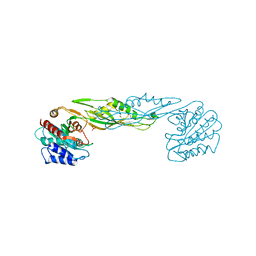 | | Crystal structure of a succinyl-diaminopimelate desuccinylase (ArgE) from Corynebacterium glutamicum ATCC 13032 at 2.97 A resolution | | Descriptor: | CHLORIDE ION, PHOSPHATE ION, Succinyl-diaminopimelate desuccinylase | | Authors: | Joint Center for Structural Genomics (JCSG), Brunger, A.T, Terwilliger, T.C, Read, R.J, Adams, P.D, Levitt, M, Schroder, G.F. | | Deposit date: | 2011-09-22 | | Release date: | 2011-10-26 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (2.972 Å) | | Cite: | Application of DEN refinement and automated model building to a difficult case of molecular-replacement phasing: the structure of a putative succinyl-diaminopimelate desuccinylase from Corynebacterium glutamicum.
Acta Crystallogr.,Sect.D, 68, 2012
|
|
3H0N
 
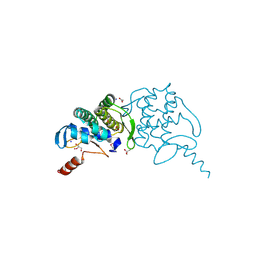 | |
3H41
 
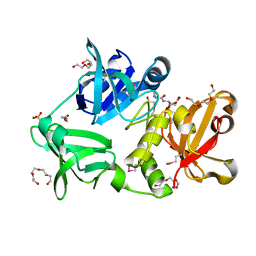 | |
3ETN
 
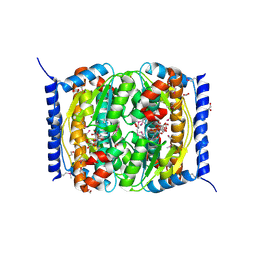 | |
3HBZ
 
 | |
3G23
 | |
3G0T
 
 | |
3GF8
 
 | |
3GO5
 
 | |
3EQX
 
 | |
3F1Z
 
 | |
3H50
 
 | |
2QTP
 
 | |
2IIZ
 
 | |
2QTD
 | |
2RA9
 
 | |
2ICH
 
 | |
2INB
 
 | |
2HUJ
 
 | |
1Z9F
 
 | |
2B8N
 
 | |
2A6A
 
 | |
1ZKG
 
 | |
2IAY
 
 | |
1ZX8
 
 | |
