7Z48
 
 | | Bottom part (C5) of bacteriophage SU10 capsid | | Descriptor: | Major head protein | | Authors: | Siborova, M, Fuzik, T, Prochazkova, M, Novacek, J, Plevka, P. | | Deposit date: | 2022-03-03 | | Release date: | 2022-08-10 | | Last modified: | 2022-10-12 | | Method: | ELECTRON MICROSCOPY (4 Å) | | Cite: | Tail proteins of phage SU10 reorganize into the nozzle for genome delivery.
Nat Commun, 13, 2022
|
|
7Z44
 
 | | Portal of bacteriophage SU10 | | Descriptor: | Portal protein | | Authors: | Siborova, M, Fuzik, T, Prochazkova, M, Novacek, J, Plevka, P. | | Deposit date: | 2022-03-03 | | Release date: | 2022-08-10 | | Last modified: | 2022-10-12 | | Method: | ELECTRON MICROSCOPY (3.6 Å) | | Cite: | Tail proteins of phage SU10 reorganize into the nozzle for genome delivery.
Nat Commun, 13, 2022
|
|
7Z47
 
 | | Tail of bacteriophage SU10 | | Descriptor: | Adaptor protein, Putative structural protein, Putative tail fiber, ... | | Authors: | Siborova, M, Fuzik, T, Prochazkova, M, Novacek, J, Plevka, P. | | Deposit date: | 2022-03-03 | | Release date: | 2022-08-10 | | Last modified: | 2022-10-12 | | Method: | ELECTRON MICROSCOPY (3.8 Å) | | Cite: | Tail proteins of phage SU10 reorganize into the nozzle for genome delivery.
Nat Commun, 13, 2022
|
|
7Z4F
 
 | | Tail of phage SU10 genome release intermediate | | Descriptor: | Adaptor protein, Portal protein, Putative structural protein, ... | | Authors: | Siborova, M, Fuzik, T, Prochazkova, M, Novacek, J, Plevka, P. | | Deposit date: | 2022-03-03 | | Release date: | 2022-08-17 | | Last modified: | 2022-10-12 | | Method: | ELECTRON MICROSCOPY (4.2 Å) | | Cite: | Tail proteins of phage SU10 reorganize into the nozzle for genome delivery.
Nat Commun, 13, 2022
|
|
7Z4A
 
 | | Bacteriophage SU10 tail and bottom part of the capsid (C1) | | Descriptor: | Adaptor protein, Major head protein, Portal protein, ... | | Authors: | Siborova, M, Fuzik, T, Prochazkova, M, Novacek, J, Plevka, P. | | Deposit date: | 2022-03-03 | | Release date: | 2022-08-17 | | Last modified: | 2022-10-12 | | Method: | ELECTRON MICROSCOPY (4.6 Å) | | Cite: | Tail proteins of phage SU10 reorganize into the nozzle for genome delivery.
Nat Commun, 13, 2022
|
|
3J35
 
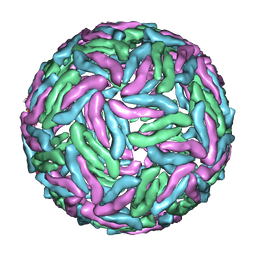 | | Cryo-EM reconstruction of Dengue virus at 37 C | | Descriptor: | envelope protein | | Authors: | Zhang, X.Z, Sheng, J, Plevka, P, Kuhn, R.J, Diamond, M.S, Rossmann, M.G. | | Deposit date: | 2013-02-24 | | Release date: | 2013-04-10 | | Last modified: | 2024-02-21 | | Method: | ELECTRON MICROSCOPY (35 Å) | | Cite: | Dengue structure differs at the temperatures of its human and mosquito hosts.
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
5G52
 
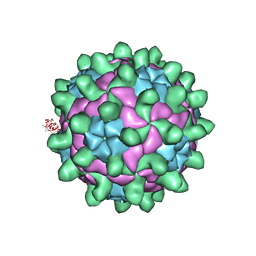 | | Crystallographic structure of full particle of Deformed Wing Virus | | Descriptor: | URIDINE-5'-MONOPHOSPHATE, VP1, VP2, ... | | Authors: | Skubnik, K, Novacek, J, Fuzik, T, Pridal, A, Paxton, R, Plevka, P. | | Deposit date: | 2016-05-18 | | Release date: | 2017-03-22 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (3.802 Å) | | Cite: | Structure of deformed wing virus, a major honey bee pathogen.
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
4Z92
 
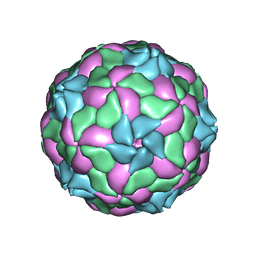 | | crystal structure of parechovirus-1 virion | | Descriptor: | Capsid subunit VP3, RNA (5'-R(*AP*UP*UP*UP*UP*U)-3'), capsid subunit VP0, ... | | Authors: | Kalynych, S, Palkova, L, Plevka, P. | | Deposit date: | 2015-04-09 | | Release date: | 2015-11-18 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | The Structure of Human Parechovirus 1 Reveals an Association of the RNA Genome with the Capsid.
J.Virol., 90, 2015
|
|
7QW9
 
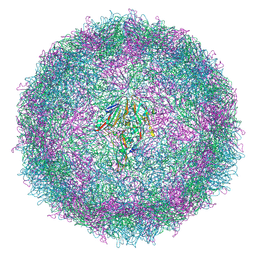 | | Cryo-EM structure of coxsackievirus A6 mature virion | | Descriptor: | Capsid protein VP1, Capsid protein VP2, Capsid protein VP3, ... | | Authors: | Buttner, C.R, Spurny, R, Fuzik, T, Plevka, P. | | Deposit date: | 2022-01-25 | | Release date: | 2022-09-07 | | Last modified: | 2022-09-14 | | Method: | ELECTRON MICROSCOPY (2.7 Å) | | Cite: | Cryo-electron microscopy and image classification reveal the existence and structure of the coxsackievirus A6 virion.
Commun Biol, 5, 2022
|
|
5G51
 
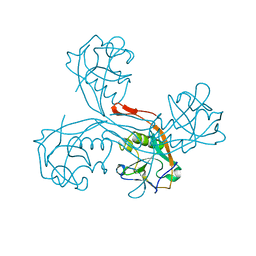 | | High resolution structure of the part of VP3 protein of Deformed Wing Virus forming P-domain | | Descriptor: | DWV-VP3-P-DOMAIN | | Authors: | Skubnik, K, Novacek, J, Fuzik, T, Pridal, A, Paxton, R, Plevka, P. | | Deposit date: | 2016-05-18 | | Release date: | 2017-03-22 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Structure of deformed wing virus, a major honey bee pathogen.
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
5J96
 
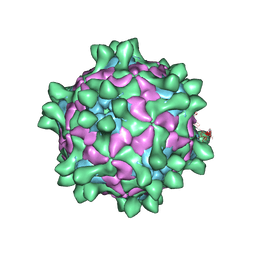 | | Crystal structure of Slow Bee Paralysis Virus at 3.4A resolution | | Descriptor: | Genome polyprotein, VP1, VP2 | | Authors: | Kalynych, S, Levdansky, Y, Palkova, L, Plevka, P. | | Deposit date: | 2016-04-08 | | Release date: | 2016-06-08 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (3.41 Å) | | Cite: | Virion Structure of Iflavirus Slow Bee Paralysis Virus at 2.6-Angstrom Resolution.
J.Virol., 90, 2016
|
|
5AOO
 
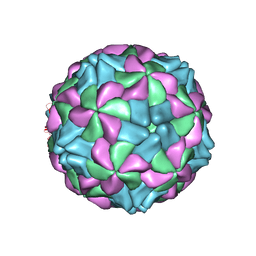 | | X-ray structure of a human Kobuvirus: Aichi virus A (AiV) | | Descriptor: | VP0, VP1, VP3 | | Authors: | Sabin, C, Palkova, L, Plevka, P. | | Deposit date: | 2015-09-11 | | Release date: | 2016-03-09 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The Use of Noncrystallographic Symmetry Averaging to Solve Structures from Data Affected by Perfect Hemihedral Twinning
Acta Crystallogr.,Sect.F, 72, 2016
|
|
6IAB
 
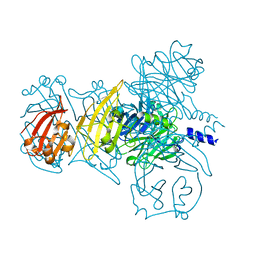 | |
6IAT
 
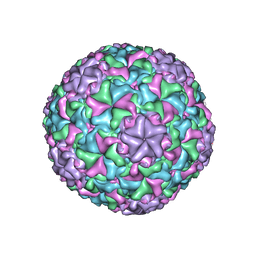 | | Icosahedrally averaged capsid of bacteriophage P68 | | Descriptor: | Arstotzka protein, Major head protein | | Authors: | Hrebik, D, Skubnik, K, Fuzik, T, Plevka, P. | | Deposit date: | 2018-11-27 | | Release date: | 2019-11-06 | | Last modified: | 2024-05-15 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Structure and genome ejection mechanism ofStaphylococcus aureusphage P68.
Sci Adv, 5, 2019
|
|
6IAC
 
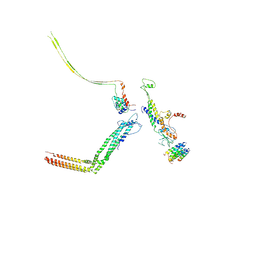 | | Portal and tail of native bacteriophage P68 | | Descriptor: | Lower collar protein, Minor structural protein, Portal protein, ... | | Authors: | Hrebik, D, Skubnik, K, Fuzik, T, Plevka, P. | | Deposit date: | 2018-11-26 | | Release date: | 2019-11-06 | | Last modified: | 2024-05-15 | | Method: | ELECTRON MICROSCOPY (3.9 Å) | | Cite: | Structure and genome ejection mechanism ofStaphylococcus aureusphage P68.
Sci Adv, 5, 2019
|
|
6IAW
 
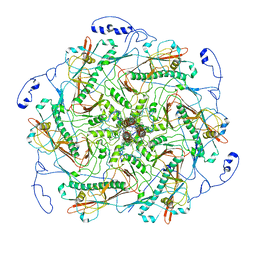 | | Structure of head fiber and inner core protein gp22 of native bacteriophage P68 | | Descriptor: | Arstotzka protein, Head fiber, Major head protein, ... | | Authors: | Hrebik, D, Skubnik, K, Fuzik, T, Plevka, P. | | Deposit date: | 2018-11-27 | | Release date: | 2019-11-06 | | Last modified: | 2024-05-15 | | Method: | ELECTRON MICROSCOPY (3.8 Å) | | Cite: | Structure and genome ejection mechanism ofStaphylococcus aureusphage P68.
Sci Adv, 5, 2019
|
|
4K7H
 
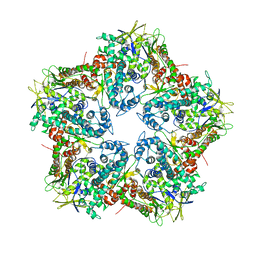 | | Major capsid protein P1 of the Pseudomonas phage phi6 | | Descriptor: | Major inner protein P1 | | Authors: | Boura, E, Nemecek, D, Plevka, P, Steven, C.A, Hurley, J.H. | | Deposit date: | 2013-04-17 | | Release date: | 2013-08-14 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (3.5964 Å) | | Cite: | Subunit Folds and Maturation Pathway of a dsRNA Virus Capsid.
Structure, 21, 2013
|
|
7OZK
 
 | | CryoEM structure of human enterovirus 70 in complex with Pleconaril | | Descriptor: | 3-{3,5-DIMETHYL-4-[3-(3-METHYL-ISOXAZOL-5-YL)-PROPOXY]-PHENYL}-5-TRIFLUOROMETHYL-[1,2,4]OXADIAZOLE, Capsid protein VP1, Capsid protein VP2, ... | | Authors: | Fuzik, T, Plevka, P, Moravcova, J. | | Deposit date: | 2021-06-28 | | Release date: | 2022-07-27 | | Last modified: | 2022-09-28 | | Method: | ELECTRON MICROSCOPY (2.31 Å) | | Cite: | Structure of Human Enterovirus 70 and Its Inhibition by Capsid-Binding Compounds.
J.Virol., 96, 2022
|
|
7OZL
 
 | | CryoEM structure of human enterovirus 70 in complex with WIN51711 | | Descriptor: | 5-(7-(4-(4,5-DIHYDRO-2-OXAZOLYL)PHENOXY)HEPTYL)-3-METHYL ISOXAZOLE, Capsid protein VP1, Capsid protein VP2, ... | | Authors: | Fuzik, T, Plevka, P, Moravcova, J. | | Deposit date: | 2021-06-28 | | Release date: | 2022-07-27 | | Last modified: | 2022-09-28 | | Method: | ELECTRON MICROSCOPY (2.74 Å) | | Cite: | Structure of Human Enterovirus 70 and Its Inhibition by Capsid-Binding Compounds.
J.Virol., 96, 2022
|
|
6F5J
 
 | |
6HBG
 
 | | Echovirus 18 native particle | | Descriptor: | Echovirus 18 viral protein 1, Echovirus 18 viral protein 2, Echovirus 18 viral protein 3, ... | | Authors: | Buchta, D, Fuzik, T, Hrebik, D, Levdansky, Y, Moravcova, J, Plevka, P. | | Deposit date: | 2018-08-10 | | Release date: | 2019-03-20 | | Last modified: | 2024-05-15 | | Method: | ELECTRON MICROSCOPY (3.16 Å) | | Cite: | Enterovirus particles expel capsid pentamers to enable genome release.
Nat Commun, 10, 2019
|
|
6HBL
 
 | | Echovirus 18 Open particle without three pentamers | | Descriptor: | Echovirus 18 capsid protein 1, Echovirus 18 capsid protein 2, Echovirus 18 capsid protein 3 | | Authors: | Buchta, D, Fuzik, T, Hrebik, D, Levdansky, Y, Moravcova, J, Plevka, P. | | Deposit date: | 2018-08-10 | | Release date: | 2019-03-20 | | Method: | ELECTRON MICROSCOPY (3.7 Å) | | Cite: | Enterovirus particles expel capsid pentamers to enable genome release.
Nat Commun, 10, 2019
|
|
5O6A
 
 | |
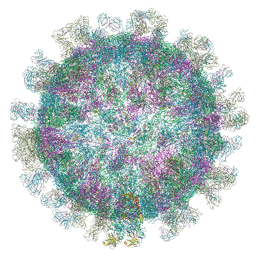
5O6V
 | |
6TEA
 
 | | Tail of native GTA particle computed with helical refinement, C6 symmetry | | Descriptor: | Phage major tail protein, TP901-1 family | | Authors: | Bardy, P, Fuzik, T, Hrebik, D, Pantucek, R, Beatty, J.T, Plevka, P. | | Deposit date: | 2019-11-11 | | Release date: | 2020-07-22 | | Last modified: | 2024-05-22 | | Method: | ELECTRON MICROSCOPY (3.89 Å) | | Cite: | Structure and mechanism of DNA delivery of a gene transfer agent.
Nat Commun, 11, 2020
|
|
