5QUP
 
 | |
5QUJ
 
 | |
5QUK
 
 | | Structure of unliganded HumRadA1.2 | | Descriptor: | PHOSPHATE ION, RadA | | Authors: | Marsh, M, Hyvonen, M. | | Deposit date: | 2020-01-27 | | Release date: | 2021-03-03 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.16 Å) | | Cite: | Optimisation of crystal forms for structure-guided drug discovery
To be published
|
|
5QUN
 
 | | Structure of unliganded HumRadA1.6 | | Descriptor: | PHOSPHATE ION, RadA | | Authors: | Marsh, M, Hyvonen, M. | | Deposit date: | 2020-01-27 | | Release date: | 2021-03-03 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.24 Å) | | Cite: | Optimisation of crystal forms for structure-guided drug discovery
To be published
|
|
5QUM
 
 | | Structure of unliganded HumRadA1.4 | | Descriptor: | RadA | | Authors: | Marsh, M, Hyvonen, M. | | Deposit date: | 2020-01-27 | | Release date: | 2021-03-03 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | Optimisation of crystal forms for structure-guided drug discovery
To be published
|
|
5QUL
 
 | | Structure of unliganded HumRadA1.3 | | Descriptor: | PHOSPHATE ION, RadA | | Authors: | Marsh, M, Hyvonen, M. | | Deposit date: | 2020-01-27 | | Release date: | 2021-03-03 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.278 Å) | | Cite: | Optimisation of crystal forms for structure-guided drug discovery
To be published
|
|
5QUT
 
 | |
5QUS
 
 | | HumRadA1 soaked with 5 mM indazole in 10% DMSO | | Descriptor: | 1H-indazole, DIMETHYL SULFOXIDE, PHOSPHATE ION, ... | | Authors: | Marsh, M, Hyvonen, M. | | Deposit date: | 2020-01-27 | | Release date: | 2021-03-03 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.247 Å) | | Cite: | Optimisation of crystal forms for structure-guided drug discovery
To be published
|
|
5QUU
 
 | | HumRadA1 as soaking control with 10% DMSO | | Descriptor: | DIMETHYL SULFOXIDE, PHOSPHATE ION, RadA | | Authors: | Marsh, M, Hyvonen, M. | | Deposit date: | 2020-01-27 | | Release date: | 2021-03-03 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.248 Å) | | Cite: | Optimisation of crystal forms for structure-guided drug discovery
To be published
|
|
5QUR
 
 | |
6WMI
 
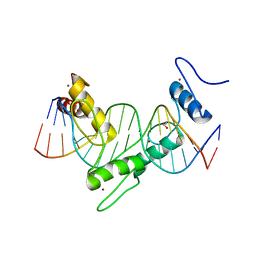 | | ZNF410 zinc fingers 1-5 with 17 mer blunt DNA Oligonucleotide | | Descriptor: | 1,2-ETHANEDIOL, DNA (5'-D(*CP*AP*CP*AP*TP*CP*CP*CP*AP*TP*AP*AP*TP*AP*AP*TP*G)-3'), DNA (5'-D(*CP*AP*TP*TP*AP*TP*TP*AP*TP*GP*GP*GP*AP*TP*GP*TP*G)-3'), ... | | Authors: | Ren, R, Horton, J.R, Cheng, X. | | Deposit date: | 2020-04-21 | | Release date: | 2020-12-09 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | ZNF410 Uniquely Activates the NuRD Component CHD4 to Silence Fetal Hemoglobin Expression.
Mol.Cell, 81, 2021
|
|
6Y0K
 
 | |
1RJW
 
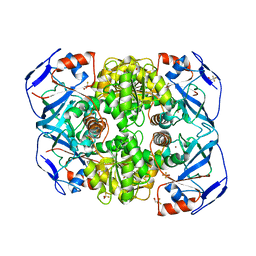 | |
6TT2
 
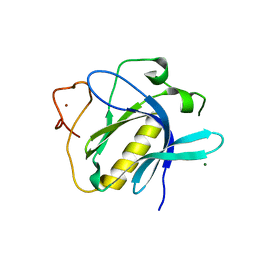 | |
7AYS
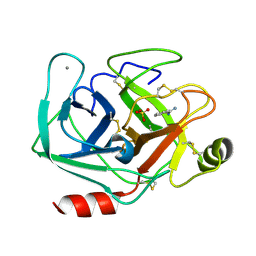 
 | |
6SIU
 
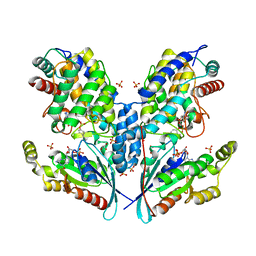 | | Crystal structure of IbpAFic2 covalently tethered to Cdc42 | | Descriptor: | Cell division control protein 42 homolog, GLYCEROL, GUANOSINE-5'-DIPHOSPHATE, ... | | Authors: | Gulen, B, Roselin, M, Albers, M, Hedberg, C, Itzen, A, Pogenberg, V. | | Deposit date: | 2019-08-12 | | Release date: | 2020-03-18 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.49 Å) | | Cite: | Identification of targets of AMPylating Fic enzymes by co-substrate-mediated covalent capture.
Nat.Chem., 12, 2020
|
|
5BQ0
 
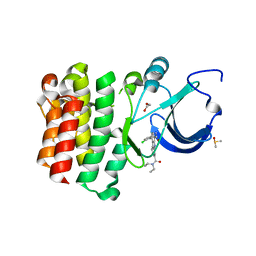 | |
5BPY
 
 | |
1JN7
 
 | |
5KWQ
 
 | | Two Tandem RRM Domains of FBP-Interacting Repressor (FIR), also Known as PUF60 | | Descriptor: | Poly(U)-binding-splicing factor PUF60 | | Authors: | Crichlow, G.V, Yang, Y, Zhou, H, Lolis, E.J, Braddock, D.T. | | Deposit date: | 2016-07-18 | | Release date: | 2017-08-23 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Unraveling the mechanism of recognition of the 3' splice site of the adenovirus major late promoter intron by the alternative splicing factor PUF60.
Plos One, 15, 2020
|
|
5KW6
 
 | | Two Tandem RRM Domains of PUF60 Bound to an AdML Pre-mRNA 3' Splice Site Analogue with a Modified Binding-Site Nucleic Acid Base | | Descriptor: | DNA (30-MER), Poly(U)-binding-splicing factor PUF60 | | Authors: | Crichlow, G.V, Hsiao, H.-H, Albright, R, Lolis, E.J, Braddock, D.T. | | Deposit date: | 2016-07-15 | | Release date: | 2017-08-23 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.91 Å) | | Cite: | Unraveling the mechanism of recognition of the 3' splice site of the adenovirus major late promoter intron by the alternative splicing factor PUF60.
Plos One, 15, 2020
|
|
2QFJ
 
 | | Crystal Structure of First Two RRM Domains of FIR Bound to ssDNA from a Portion of FUSE | | Descriptor: | DNA (5'-D(*DTP*DCP*DGP*DGP*DGP*DAP*DTP*DTP*DTP*DTP*DTP*DTP*DAP*DTP*DTP*DTP*DTP*DGP*DTP*DGP*DTP*DTP*DAP*DTP*DT)-3'), FBP-interacting repressor | | Authors: | Crichlow, G.V, Yang, Y, Fan, C, Lolis, E, Braddock, D. | | Deposit date: | 2007-06-27 | | Release date: | 2008-03-04 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Dimerization of FIR upon FUSE DNA binding suggests a mechanism of c-myc inhibition
EMBO J., 27, 2007
|
|
2MPL
 
 | |
2HLS
 
 | |
1JKW
 
 | |
