3Q8D
 
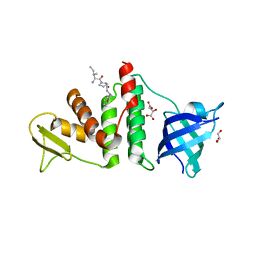 | | E. coli RecO complex with SSB C-terminus | | Descriptor: | 3-[(3-CHOLAMIDOPROPYL)DIMETHYLAMMONIO]-1-PROPANESULFONATE, D-MALATE, DNA repair protein recO, ... | | Authors: | Koroleva, O, Ryzhikov, M, Korolev, S. | | Deposit date: | 2011-01-06 | | Release date: | 2011-05-04 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Mechanism of RecO recruitment to DNA by single-stranded DNA binding protein.
Nucleic Acids Res., 39, 2011
|
|
2O5V
 
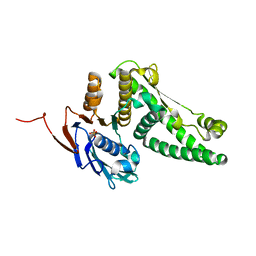 | | Recombination mediator RecF | | Descriptor: | DNA replication and repair protein recF, SULFATE ION | | Authors: | Korolev, S, Koroleva, O. | | Deposit date: | 2006-12-06 | | Release date: | 2007-02-06 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.61 Å) | | Cite: | Structural conservation of RecF and Rad50: implications for DNA recognition and RecF function.
Embo J., 26, 2007
|
|
4X57
 
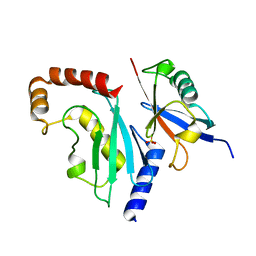 | |
6AUN
 
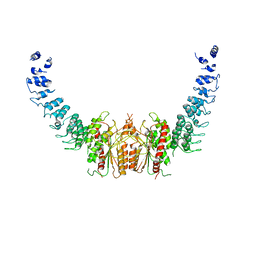 | | calcium-independent phospholipase A2 beta | | Descriptor: | PLA2G6, iPLA2beta | | Authors: | Malley, K, Koroleva, O, Miller, I, Sanishvili, R, Jenkins, C.M, Gross, R.W, Korolev, S. | | Deposit date: | 2017-09-01 | | Release date: | 2018-03-07 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (3.951 Å) | | Cite: | The structure of iPLA2beta reveals dimeric active sites and suggests mechanisms of regulation and localization.
Nat Commun, 9, 2018
|
|
1I6N
 
 | | 1.8 A Crystal structure of IOLI protein with a binding zinc atom | | Descriptor: | IOLI PROTEIN, ZINC ION | | Authors: | Zhang, R.G, Dementiva, I, Collart, F, Quaite-Randall, E, Joachimiak, A, Alkire, R, Maltsev, N, Korolev, O, Dieckman, L, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2001-03-02 | | Release date: | 2002-03-13 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of Bacillus subtilis ioli shows endonuclase IV fold with altered Zn binding.
Proteins, 48, 2002
|
|
1U5K
 
 | | Recombinational repair protein RecO | | Descriptor: | ZINC ION, hypothetical protein | | Authors: | Makharashvili, N, Koroleva, O, Bera, S, Grandgenett, D.P, Korolev, S. | | Deposit date: | 2004-07-27 | | Release date: | 2004-10-26 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A Novel Structure of DNA Repair Protein RecO from Deinococcus radiodurans
STRUCTURE, 12, 2004
|
|
1NPY
 
 | | Structure of shikimate 5-dehydrogenase-like protein HI0607 | | Descriptor: | ACETYL GROUP, Hypothetical shikimate 5-dehydrogenase-like protein HI0607 | | Authors: | Korolev, S, Koroleva, O, Zarembinski, T, Collart, F, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2003-01-20 | | Release date: | 2003-07-29 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Crystal Structure of a Novel Shikimate Dehydrogenase from Haemophilus influenzae.
J.Biol.Chem., 280, 2005
|
|
1K6D
 
 | | CRYSTAL STRUCTURE OF ACETATE COA-TRANSFERASE ALPHA SUBUNIT | | Descriptor: | ACETATE COA-TRANSFERASE ALPHA SUBUNIT, MAGNESIUM ION | | Authors: | Korolev, S, Koroleva, O, Petterson, K, Collart, F, Dementieva, I, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2001-10-15 | | Release date: | 2002-06-26 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Autotracing of Escherichia coli acetate CoA-transferase alpha-subunit structure using 3.4 A MAD and 1.9 A native data.
Acta Crystallogr.,Sect.D, 58, 2002
|
|
1NJH
 
 | |
1PVT
 
 | | Crystal structure of sugar-phosphate aldolase from Thermotoga maritima | | Descriptor: | sugar-phosphate aldolase | | Authors: | Osipiuk, J, Cuff, M.E, Korolev, O, Skarina, T, Savchenko, A, Edwards, A, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2003-06-28 | | Release date: | 2003-09-23 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure of sugar-phosphate aldolase from Thermotoga maritima.
To be Published
|
|
1L7A
 
 | |
1I60
 
 | | Structural genomics, IOLI protein | | Descriptor: | IOLI PROTEIN | | Authors: | Zhang, R, Dementieva, I, Collart, F, Quaite-Randall, E, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2001-03-01 | | Release date: | 2002-03-13 | | Last modified: | 2017-10-04 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystal structure of Bacillus subtilis ioli shows endonuclase IV fold with altered Zn binding.
Proteins, 48, 2002
|
|
