2CB1
 
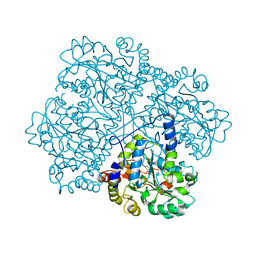 | | Crystal Structure of O-actetyl Homoserine Sulfhydrylase From Thermus Thermophilus HB8,OAH2. | | Descriptor: | O-ACETYL HOMOSERINE SULFHYDRYLASE, PYRIDOXAL-5'-PHOSPHATE | | Authors: | Imagawa, T, Utsunomiya, H, Tsuge, H, Ebihara, A, Kanagawa, M, Nakagawa, N, Kuroishi, C, Agari, Y, Kuramitsu, S, Yokoyama, S. | | Deposit date: | 2005-12-28 | | Release date: | 2007-01-03 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The Crystal Structure of O-Acetyl Homoserine Sulfhydrylase
To be Published
|
|
6JF2
 
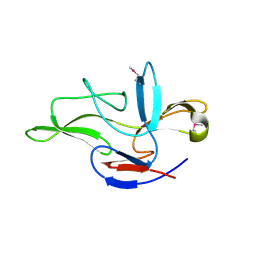 | |
2CTZ
 
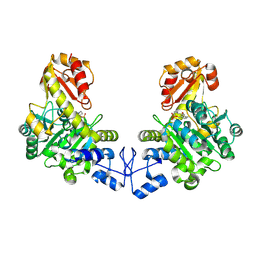 | | Crystal structure of o-acetyl homoserine sulfhydrylase from Thermus thermophilus HB8 | | Descriptor: | O-acetyl-L-homoserine sulfhydrylase, PYRIDOXAL-5'-PHOSPHATE | | Authors: | Imagawa, T, Kousumi, Y, Tsuge, H, Utsunomiya, H, Ebihara, A, Nakagawa, N, Yokoyama, S, Kuramitsu, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2005-05-24 | | Release date: | 2005-11-24 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Crystal structure of o-acetyl homoserine sulfhydrylase from Thermus thermophilus HB8
To be Published
|
|
6JZT
 
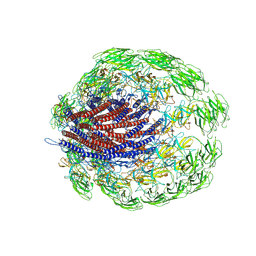 | |
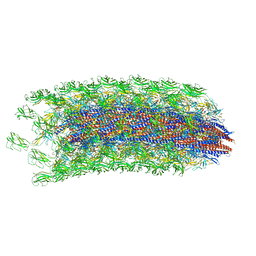
6K3I
 | |
6JZR
 
 | |
7VFE
 
 | |
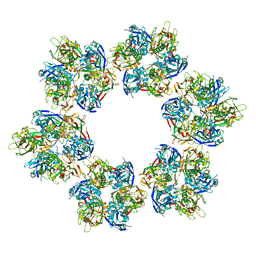
7VFH
 | |
7VFD
 
 | |
7VFF
 
 | |
7VFG
 
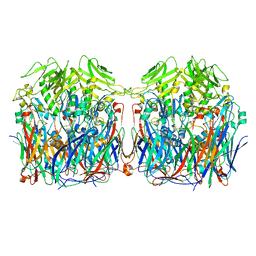 | |
5Z9W
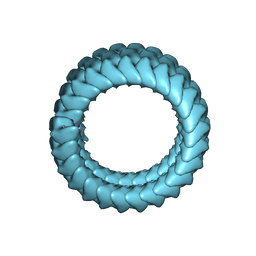 
 | | Ebola virus nucleoprotein-RNA complex | | Descriptor: | Ebolavirus nucleoprotein (residues 19-406), RNA (6-MER) | | Authors: | Sugita, Y, Matsunami, H, Kawaoka, Y, Noda, T, Wolf, M. | | Deposit date: | 2018-02-05 | | Release date: | 2018-10-24 | | Last modified: | 2024-03-27 | | Method: | ELECTRON MICROSCOPY (3.6 Å) | | Cite: | Cryo-EM structure of the Ebola virus nucleoprotein-RNA complex at 3.6 angstrom resolution.
Nature, 563, 2018
|
|
2D4Y
 
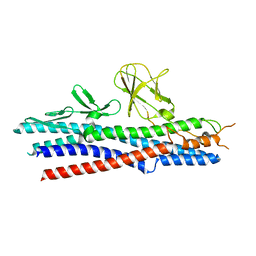 | |
2D4X
 
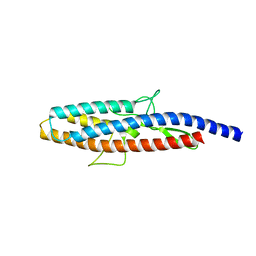 | |
8JUB
 
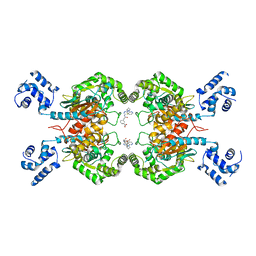 | | Crystal structure of glutaminase C in complex with compound 27 | | Descriptor: | 3-[2-oxidanylidene-2-[[5-[[(3R)-1-pyridazin-3-ylpyrrolidin-3-yl]amino]-1,3,4-thiadiazol-2-yl]amino]ethyl]benzoic acid, Glutaminase kidney isoform, mitochondrial | | Authors: | Wang, X, Hanyu, S, Tingting, D. | | Deposit date: | 2023-06-26 | | Release date: | 2023-10-11 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | Targeting the Subpocket Enables the Discovery of Thiadiazole-Pyridazine Derivatives as Glutaminase C Inhibitors.
Acs Med.Chem.Lett., 14, 2023
|
|
8JUE
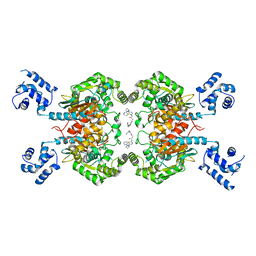 
 | | Crystal structure of glutaminase C in complex with compound 11 | | Descriptor: | 2-(3-phenoxyphenyl)-N-[5-[[(3R)-1-pyridazin-3-ylpyrrolidin-3-yl]amino]-1,3,4-thiadiazol-2-yl]ethanamide, Glutaminase kidney isoform, mitochondrial | | Authors: | Wang, X, Hanyu, S, Tingting, D. | | Deposit date: | 2023-06-26 | | Release date: | 2023-10-11 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.39 Å) | | Cite: | Targeting the Subpocket Enables the Discovery of Thiadiazole-Pyridazine Derivatives as Glutaminase C Inhibitors.
Acs Med.Chem.Lett., 14, 2023
|
|
5B6I
 
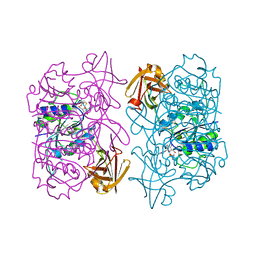 | | Structure of fluorinase from Streptomyces sp. MA37 | | Descriptor: | ADENOSINE, Fluorinase, METHIONINE | | Authors: | Xue, B, Robinson, R.C. | | Deposit date: | 2016-05-29 | | Release date: | 2016-10-26 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Directed Evolution of a Fluorinase for Improved Fluorination Efficiency with a Non-native Substrate
Angew.Chem.Int.Ed.Engl., 55, 2016
|
|
8HTU
 
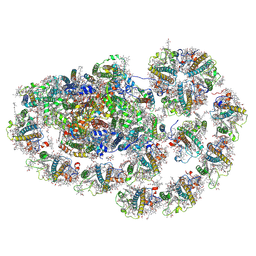 | | Cryo-EM structure of PpPSI-L | | Descriptor: | (1R,3R)-6-{(3E,5E,7E,9E,11E,13E,15E,17E)-18-[(1S,4R,6R)-4-HYDROXY-2,2,6-TRIMETHYL-7-OXABICYCLO[4.1.0]HEPT-1-YL]-3,7,12,16-TETRAMETHYLOCTADECA-1,3,5,7,9,11,13,15,17-NONAENYLIDENE}-1,5,5-TRIMETHYLCYCLOHEXANE-1,3-DIOL, (3R,3'R,6S)-4,5-DIDEHYDRO-5,6-DIHYDRO-BETA,BETA-CAROTENE-3,3'-DIOL, (3S,5R,6S,3'S,5'R,6'S)-5,6,5',6'-DIEPOXY-5,6,5',6'- TETRAHYDRO-BETA,BETA-CAROTENE-3,3'-DIOL, ... | | Authors: | Li, M, Pan, X.W, Sun, H.Y. | | Deposit date: | 2022-12-21 | | Release date: | 2023-08-02 | | Last modified: | 2023-08-30 | | Method: | ELECTRON MICROSCOPY (2.87 Å) | | Cite: | Structural insights into the assembly and energy transfer of the Lhcb9-dependent photosystem I from moss Physcomitrium patens.
Nat.Plants, 9, 2023
|
|
2JK7
 
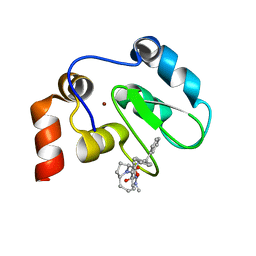 | | XIAP BIR3 bound to a Smac Mimetic | | Descriptor: | (3S,6S,7Z,10AS)-N-(DIPHENYLMETHYL)-6-{[(2S)-2-(METHYLIDENEAMINO)BUTANOYL]AMINO}-5-OXO-1,2,3,5,6,9,10,10A-OCTAHYDROPYRROLO[1,2-A]AZOCINE-3-CARBOXAMIDE, BACULOVIRAL IAP REPEAT-CONTAINING PROTEIN 4, ZINC ION | | Authors: | Saito, N.G, Meagher, J.L, Stuckey, J.A. | | Deposit date: | 2008-08-21 | | Release date: | 2008-11-04 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.82 Å) | | Cite: | Structure-Based Design, Synthesis, Evaluation, and Crystallographic Studies of Conformationally Constrained Smac Mimetics as Inhibitors of the X-Linked Inhibitor of Apoptosis Protein (Xiap).
J.Med.Chem., 51, 2008
|
|
5EC9
 
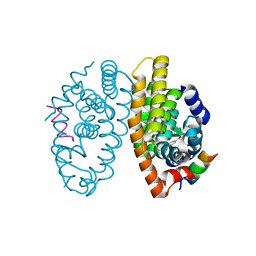 | | Retinoic acid receptor alpha in complex with chiral dihydrobenzofuran benzoic acid 9a and a fragment of the coactivator TIF2 | | Descriptor: | 4-[(11S,15R)-4,4,7,7-Tetramethyl-16-oxatetracyclo[8.6.0.03,8.011,15]hexadeca-1(10),2,8-trien-11-yl]benzoic acid, LYS-HIS-LYS-ILE-LEU-HIS-ARG-LEU-LEU-GLN-ASP, Retinoic acid receptor RXR-alpha | | Authors: | Leysen, S, Ottmann, C, Schafer, A, Scheepstra, M, Brunsveld, L, Sunden, R, Ma, J.N, Burnstein, E.S, Olsson, R. | | Deposit date: | 2015-10-20 | | Release date: | 2016-03-09 | | Last modified: | 2024-06-19 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Chiral Dihydrobenzofuran Acids Show Potent Retinoid X Receptor-Nuclear Receptor Related 1 Protein Dimer Activation.
J.Med.Chem., 59, 2016
|
|
3HH8
 
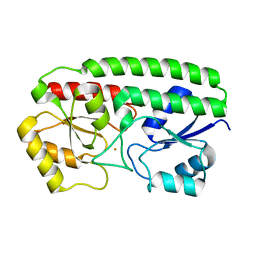 | | Crystal Structure and metal binding properties of the lipoprotein MtsA | | Descriptor: | FE (III) ION, Metal ABC transporter substrate-binding lipoprotein | | Authors: | Baker, E.N, Baker, H.M, Sun, X, Ye, Q.-Y. | | Deposit date: | 2009-05-15 | | Release date: | 2009-06-09 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.87 Å) | | Cite: | Crystal structure and metal binding properties of the lipoprotein MtsA, responsible for iron transport in Streptococcus pyogenes.
Biochemistry, 48, 2009
|
|
2XEM
 
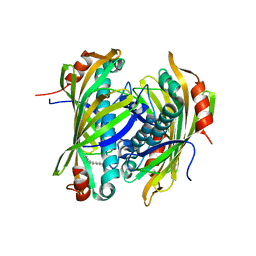 | | Induced-fit and allosteric effects upon polyene binding revealed by crystal structures of the Dynemicin thioesterase | | Descriptor: | (3E,5E,7E,9E,11E,13E)-pentadeca-3,5,7,9,11,13-hexaen-2-one, DYNE7 | | Authors: | Liew, C.W, Sharff, A, Kotaka, M, Kong, R, Bricogne, G, Liang, Z.X, Lescar, J. | | Deposit date: | 2010-05-17 | | Release date: | 2010-10-13 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Induced-Fit Upon Ligand Binding Revealed by Crystal Structures of the Hot-Dog Fold Thioesterase in Dynemicin Biosynthesis.
J.Mol.Biol., 404, 2010
|
|
7MBO
 
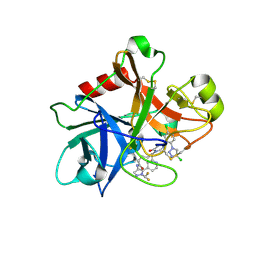 | | FACTOR XIA (PICHIA PASTORIS; C500S [C122S]) IN COMPLEX WITH THE INHIBITOR Milvexian (BMS-986177), IUPAC NAME:(6R,10S)-10-{4-[5-chloro-2-(4-chloro-1H-1,2,3-triazol-1-yl)phenyl]-6- oxopyrimidin-1(6H)-yl}-1-(difluoromethyl)-6-methyl-1,4,7,8,9,10-hexahydro-15,11- (metheno)pyrazolo[4,3-b][1,7]diazacyclotetradecin-5(6H)-one | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Coagulation factor XIa light chain, Milvexian | | Authors: | Sheriff, S. | | Deposit date: | 2021-04-01 | | Release date: | 2021-09-15 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (0.924 Å) | | Cite: | Discovery of Milvexian, a High-Affinity, Orally Bioavailable Inhibitor of Factor XIa in Clinical Studies for Antithrombotic Therapy.
J.Med.Chem., 65, 2022
|
|
5QTT
 
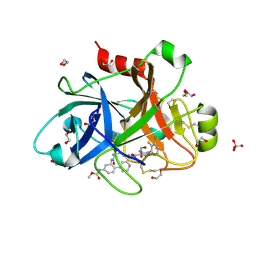 | | FACTOR XIA IN COMPLEX WITH THE INHIBITOR methyl [(3R,7S)-7-{[5-amino-1-(3-chloro-2-fluorophenyl)-1H-pyrazole-4-carbonyl]amino}-3-methyl-2-oxo-2,3,4,5,6,7-hexahydro-1H-12,8-(metheno)-1,9-benzodiazacyclotetradecin-15-yl]carbamate | | Descriptor: | 1,2-ETHANEDIOL, Coagulation factor XI, SULFATE ION, ... | | Authors: | Sheriff, S. | | Deposit date: | 2019-10-16 | | Release date: | 2019-12-25 | | Last modified: | 2021-05-12 | | Method: | X-RAY DIFFRACTION (2.23 Å) | | Cite: | Potent, Orally Bioavailable and Efficacious Macrocyclic Inhibitors of Factor XIa. Discovery of Pyridine-Based Macrocycles Possessing Phenylazole Carboxamide P1 Groups.
J.Med.Chem., 63, 2019
|
|
5QTU
 
 | | FACTOR XIA IN COMPLEX WITH THE INHIBITOR methyl [(3R,7S)-7-{[1-(3-chloro-2-fluorophenyl)-5-methyl-1H-imidazole-4-carbonyl]amino}-3-methyl-2-oxo-2,3,4,5,6,7-hexahydro-1H-12,8-(metheno)-1,9-benzodiazacyclotetradecin-15-yl]carbamate | | Descriptor: | 1,2-ETHANEDIOL, Coagulation factor XI, SULFATE ION, ... | | Authors: | Sheriff, S. | | Deposit date: | 2019-10-16 | | Release date: | 2019-12-25 | | Last modified: | 2021-05-12 | | Method: | X-RAY DIFFRACTION (2.53 Å) | | Cite: | Potent, Orally Bioavailable and Efficacious Macrocyclic Inhibitors of Factor XIa. Discovery of Pyridine-Based Macrocycles Possessing Phenylazole Carboxamide P1 Groups.
J.Med.Chem., 63, 2019
|
|
