2D4A
 
 | | Structure of the malate dehydrogenase from Aeropyrum pernix | | Descriptor: | Malate dehydrogenase | | Authors: | Kawakami, R, Sakuraba, H, Tsuge, H, Ohshima, T. | | Deposit date: | 2005-10-12 | | Release date: | 2006-11-14 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.87 Å) | | Cite: | Refolding, characterization and crystal structure of (S)-malate dehydrogenase from the hyperthermophilic archaeon Aeropyrum pernix.
Biochim.Biophys.Acta, 1794, 2009
|
|
2YY7
 
 | | Crystal structure of thermolabile L-threonine dehydrogenase from Flavobacterium frigidimaris KUC-1 | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, 3,6,9,12,15,18,21-HEPTAOXATRICOSANE-1,23-DIOL, GLYCEROL, ... | | Authors: | Yoneda, K, Sakuraba, H, Oikawa, T, Muraoka, I, Ohshima, T. | | Deposit date: | 2007-04-27 | | Release date: | 2008-04-29 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.061 Å) | | Cite: | Crystal structure of UDP-galactose 4-epimerase-like L-threonine dehydrogenase belonging to the intermediate short-chain dehydrogenase-reductase superfamily
Febs J., 277, 2010
|
|
2YVS
 
 | |
3A9W
 
 | | Crystal structure of L-Threonine bound L-Threonine dehydrogenase (Y137F) from Hyperthermophilic Archaeon Thermoplasma volcanium | | Descriptor: | (4R)-2-METHYLPENTANE-2,4-DIOL, (4S)-2-METHYL-2,4-PENTANEDIOL, NDP-sugar epimerase, ... | | Authors: | Yoneda, K, Sakuraba, H, Ohshima, T. | | Deposit date: | 2009-11-07 | | Release date: | 2010-11-10 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Crystal Structure of Binary and Ternary Complexes of Archaeal UDP-galactose 4-Epimerase-like L-Threonine Dehydrogenase from Thermoplasma volcanium.
J.Biol.Chem., 287, 2012
|
|
2DC0
 
 | | Crystal structure of amidase | | Descriptor: | probable amidase | | Authors: | Ohshima, T, Sakuraba, H, Ebihara, A, Kanagawa, M, Nakagawa, N, Kuroishi, C, Satoh, S, Kuramitsu, S, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2005-12-17 | | Release date: | 2007-01-02 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of amidase
To be Published
|
|
3A4V
 
 | | Crystal structure of pyruvate bound L-Threonine dehydrogenase from Hyperthermophilic Archaeon Thermoplasma volcanium | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, NDP-sugar epimerase, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, ... | | Authors: | Yoneda, K, Sakuraba, H, Ohshima, T. | | Deposit date: | 2009-07-20 | | Release date: | 2010-07-28 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.78 Å) | | Cite: | Crystal Structure of Binary and Ternary Complexes of Archaeal UDP-galactose 4-Epimerase-like L-Threonine Dehydrogenase from Thermoplasma volcanium.
J.Biol.Chem., 287, 2012
|
|
3A1N
 
 | |
3ABI
 
 | | Crystal Structure of L-Lysine Dehydrogenase from Hyperthermophilic Archaeon Pyrococcus horikoshii | | Descriptor: | NICOTINAMIDE-ADENINE-DINUCLEOTIDE, Putative uncharacterized protein PH1688, SULFATE ION | | Authors: | Yoneda, K, Sakuraba, H, Fukuda, J, Ohshima, T. | | Deposit date: | 2009-12-14 | | Release date: | 2009-12-29 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.44 Å) | | Cite: | First crystal structure of L-lysine 6-dehydrogenase as an NAD-dependent amine dehydrogenase.
J.Biol.Chem., 285, 2010
|
|
6JYG
 
 | | Crystal Structure of L-threonine dehydrogenase from Phytophthora infestans | | Descriptor: | 3,6,9,12,15,18,21-HEPTAOXATRICOSANE-1,23-DIOL, CITRATE ANION, L-threonine 3-dehydrogenase, ... | | Authors: | Yoneda, K, Sakuraba, H, Ohshima, T. | | Deposit date: | 2019-04-26 | | Release date: | 2020-04-01 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.31 Å) | | Cite: | Catalytic properties and crystal structure of UDP-galactose 4-epimerase-like l-threonine 3-dehydrogenase from Phytophthora infestans.
Enzyme.Microb.Technol., 140, 2020
|
|
6JXN
 
 | | Crystal Structure of Indigo reductase from Bacillus smithii type strain DSM 4216 | | Descriptor: | 2-[N-CYCLOHEXYLAMINO]ETHANE SULFONIC ACID, 3,6,9,12,15,18,21-HEPTAOXATRICOSANE-1,23-DIOL, FLAVIN MONONUCLEOTIDE, ... | | Authors: | Yoneda, K, Sakuraba, H, Ohshima, T. | | Deposit date: | 2019-04-24 | | Release date: | 2020-04-01 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.97 Å) | | Cite: | Structural and biochemical characterization of an extremely thermostable FMN-dependent NADH-indigo reductase from Bacillus smithii.
Int.J.Biol.Macromol., 164, 2020
|
|
6JXS
 
 | | Crystal Structure of Indigo reductase (Y151F) from Bacillus smithii type strain DSM 4216 | | Descriptor: | 3,6,9,12,15,18,21-HEPTAOXATRICOSANE-1,23-DIOL, FLAVIN MONONUCLEOTIDE, FMN-dependent NADH-azoreductase | | Authors: | Yoneda, K, Sakuraba, H, Ohshima, T. | | Deposit date: | 2019-04-24 | | Release date: | 2020-04-01 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structural and biochemical characterization of an extremely thermostable FMN-dependent NADH-indigo reductase from Bacillus smithii.
Int.J.Biol.Macromol., 164, 2020
|
|
3AJR
 
 | | Crystal structure of L-3-Hydroxynorvaline bound L-Threonine dehydrogenase (Y137F) from Hyperthermophilic Archaeon Thermoplasma volcanium | | Descriptor: | (3R)-3-hydroxy-L-norvaline, (4S)-2-METHYL-2,4-PENTANEDIOL, NDP-sugar epimerase, ... | | Authors: | Yoneda, K, Sakuraba, H, Ohshima, T. | | Deposit date: | 2010-06-14 | | Release date: | 2011-06-15 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | Crystal Structure of Binary and Ternary Complexes of Archaeal UDP-galactose 4-Epimerase-like L-Threonine Dehydrogenase from Thermoplasma volcanium.
J.Biol.Chem., 287, 2012
|
|
3W82
 
 | | Human alpha-L-iduronidase in complex with iduronic acid | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Alpha-L-iduronidase, ... | | Authors: | Maita, N, Tsukimura, T, Taniguchi, T, Saito, S, Ohno, K, Taniguchi, H, Sakuraba, H. | | Deposit date: | 2013-03-11 | | Release date: | 2013-08-21 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.76 Å) | | Cite: | Human alpha-L-iduronidase uses its own N-glycan as a substrate-binding and catalytic module
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
3W81
 
 | | Human alpha-l-iduronidase | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Alpha-L-iduronidase, ... | | Authors: | Maita, N, Tsukimura, T, Taniguchi, T, Saito, S, Ohno, K, Taniguchi, H, Sakuraba, H. | | Deposit date: | 2013-03-11 | | Release date: | 2013-08-21 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Human alpha-L-iduronidase uses its own N-glycan as a substrate-binding and catalytic module
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
3VPX
 
 | | Crystal structure of leucine dehydrogenase from a psychrophilic bacterium Sporosarcina psychrophila. | | Descriptor: | Leucine dehydrogenase | | Authors: | Zhao, Y, Wakamatsu, T, Doi, K, Sakuraba, H, Ohshima, T. | | Deposit date: | 2012-03-14 | | Release date: | 2013-02-06 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | A psychrophilic leucine dehydrogenase from Sporosarcina psychrophila: Purification, characterization, gene sequencing and crystal structure analysis
J.MOL.CATAL., B ENZYM., 83, 2012
|
|
3WXB
 
 | | Crystal structure of NADPH bound carbonyl reductase from chicken fatty liver | | Descriptor: | 1,2-ETHANEDIOL, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, Uncharacterized protein | | Authors: | Yoneda, K, Sakuraba, H, Fukuda, Y, Sone, T, Araki, T, Ohshima, T. | | Deposit date: | 2014-07-29 | | Release date: | 2015-07-29 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | A novel NAD(P)H-dependent carbonyl reductase specifically expressed in the thyroidectomized chicken fatty liver: catalytic properties and crystal structure.
Febs J., 282, 2015
|
|
3VYL
 
 | | Structure of L-ribulose 3-epimerase | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, L-ribulose 3-epimerase, MANGANESE (II) ION | | Authors: | Uechi, K, Sakuraba, H, Takata, G. | | Deposit date: | 2012-09-27 | | Release date: | 2013-10-02 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structural insight into L-ribulose 3-epimerase from Mesorhizobium loti.
Acta Crystallogr.,Sect.D, 69, 2013
|
|
3WQO
 
 | | Crystal structure of D-tagatose 3-epimerase-like protein | | Descriptor: | MANGANESE (II) ION, Uncharacterized protein MJ1311 | | Authors: | Uechi, K, Takata, G, Yoneda, K, Ohshima, T, Sakuraba, H. | | Deposit date: | 2014-01-28 | | Release date: | 2014-07-09 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.64 Å) | | Cite: | Structure of D-tagatose 3-epimerase-like protein from Methanocaldococcus jannaschii
Acta Crystallogr.,Sect.F, 70, 2014
|
|
3W6Z
 
 | |
3W6U
 
 | |
3WS7
 
 | | The 1.18 A resolution structure of L-serine 3-dehydrogenase complexed with NADP+ and sulfate ion from the hyperthermophilic archaeon Pyrobaculum calidifontis | | Descriptor: | 6-phosphogluconate dehydrogenase, NAD-binding protein, ACETIC ACID, ... | | Authors: | Yoneda, K, Sakuraba, H, Ohshima, T. | | Deposit date: | 2014-03-04 | | Release date: | 2015-03-04 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.18 Å) | | Cite: | Crystal structure of the NADP+and tartrate-bound complex of L-serine 3-dehydrogenase from the hyperthermophilic archaeon Pyrobaculum calidifontis.
Extremophiles, 22, 2018
|
|
5BRO
 
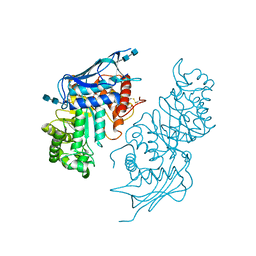 | | Crystal structure of modified HexB (modB) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Beta-hexosaminidase subunit beta, FORMIC ACID, ... | | Authors: | Kitakaze, K, Maita, N, Itoh, K. | | Deposit date: | 2015-06-01 | | Release date: | 2016-05-04 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Protease-resistant modified human beta-hexosaminidase B ameliorates symptoms in GM2 gangliosidosis model.
J.Clin.Invest., 126, 2016
|
|
4NXS
 
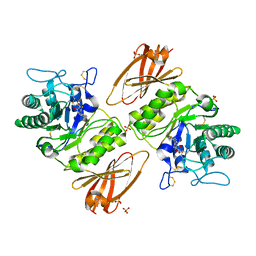 | | Crystal structure of human alpha-galactosidase A in complex with 1-deoxygalactonojirimycin-pFPhT | | Descriptor: | (2R,3S,4R,5S)-N-(4-fluorophenyl)-3,4,5-trihydroxy-2-(hydroxymethyl)piperidine-1-carbothioamide, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Johnson, J.L, Drury, J.E, Lieberman, R.L. | | Deposit date: | 2013-12-09 | | Release date: | 2014-06-11 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.5493 Å) | | Cite: | Molecular Basis of 1-Deoxygalactonojirimycin Arylthiourea Binding to Human alpha-Galactosidase A: Pharmacological Chaperoning Efficacy on Fabry Disease Mutants.
Acs Chem.Biol., 9, 2014
|
|
2DCH
 
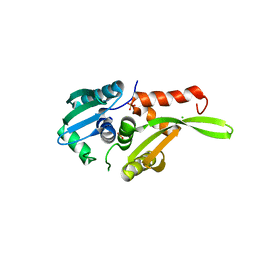 | | Crystal structure of archaeal intron-encoded homing endonuclease I-Tsp061I | | Descriptor: | CHLORIDE ION, SULFATE ION, putative homing endonuclease | | Authors: | Nakayama, H, Tsuge, H, Shimamura, T, Miyano, M, Nomura, N, Sako, Y. | | Deposit date: | 2006-01-06 | | Release date: | 2006-07-06 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.06 Å) | | Cite: | Structure of a hyperthermophilic archaeal homing endonuclease, I-Tsp061I: contribution of cross-domain polar networks to thermostability.
J.Mol.Biol., 365, 2007
|
|
