5WPS
 
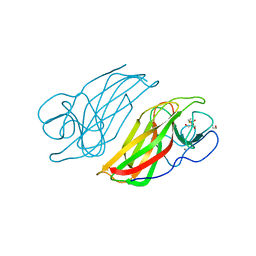 | | Crystal structure HpiC1 Y101F | | Descriptor: | 1,2-ETHANEDIOL, 12-epi-hapalindole C/U synthase, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, ... | | Authors: | Newmister, S.A, Li, S, Garcia-Borras, M, Sanders, J.N, Yang, S, Lowell, A.N, Yu, F, Smith, J.L, Williams, R.M, Houk, K.N, Sherman, D.H. | | Deposit date: | 2017-08-07 | | Release date: | 2018-04-04 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.389 Å) | | Cite: | Structural basis of the Cope rearrangement and cyclization in hapalindole biogenesis.
Nat. Chem. Biol., 14, 2018
|
|
8G7X
 
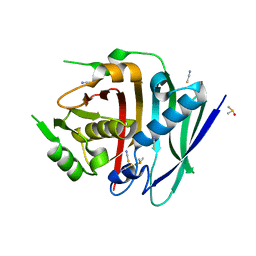 | | Human fatty acid synthase dehydratase domain | | Descriptor: | 3-hydroxyacyl-[acyl-carrier-protein] dehydratase, ACETATE ION, DIMETHYL SULFOXIDE, ... | | Authors: | Akey, D.L, Konwerski, J.R, McCullough, T.M, Smith, J.L. | | Deposit date: | 2023-02-17 | | Release date: | 2023-06-07 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.815 Å) | | Cite: | Structure of a modular polyketide synthase reducing region.
Structure, 31, 2023
|
|
8G7W
 
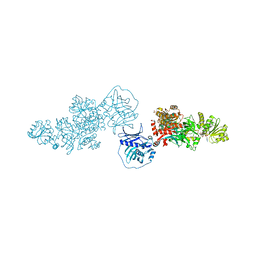 | | Type I modPKS reducing region | | Descriptor: | GLYCEROL, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, SULFATE ION, ... | | Authors: | McCullough, T.M, Smith, J.L. | | Deposit date: | 2023-02-17 | | Release date: | 2023-06-07 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (3.4 Å) | | Cite: | Structure of a modular polyketide synthase reducing region.
Structure, 31, 2023
|
|
7UCH
 
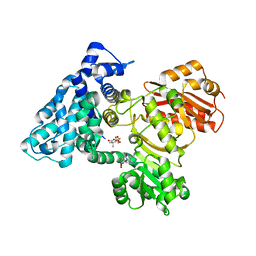 | | AprA Methyltransferase 1 - GNAT in complex with Mn2+ , SAM, and Di-methyl-malonate | | Descriptor: | AprA Methyltransferase 1, GLYCEROL, MANGANESE (II) ION, ... | | Authors: | Skiba, M.A, Lao, Y, Smith, J.L. | | Deposit date: | 2022-03-16 | | Release date: | 2022-06-01 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.18 Å) | | Cite: | Structural Basis for Control of Methylation Extent in Polyketide Synthase Metal-Dependent C -Methyltransferases.
Acs Chem.Biol., 17, 2022
|
|
5IOQ
 
 | | Flavin-dependent thymidylate synthase in complex with FAD and deoxyuridine | | Descriptor: | 2'-DEOXYURIDINE, FLAVIN-ADENINE DINUCLEOTIDE, TRIETHYLENE GLYCOL, ... | | Authors: | Bernard, S.M, Stull, F.W, Smith, J.L. | | Deposit date: | 2016-03-08 | | Release date: | 2016-06-08 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | Deprotonations in the Reaction of Flavin-Dependent Thymidylate Synthase.
Biochemistry, 55, 2016
|
|
5IOR
 
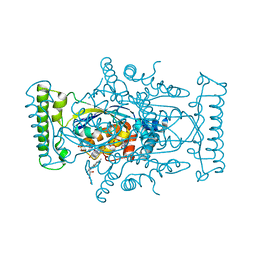 | | Flavin-dependent thymidylate synthase in complex with FAD and 2'-deoxyuridine-5'-monosulfate | | Descriptor: | 2'-deoxy-5'-O-sulfouridine, FLAVIN-ADENINE DINUCLEOTIDE, RIBOFLAVIN, ... | | Authors: | Bernard, S.M, Stull, F.W, Smith, J.L. | | Deposit date: | 2016-03-08 | | Release date: | 2016-06-08 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Deprotonations in the Reaction of Flavin-Dependent Thymidylate Synthase.
Biochemistry, 55, 2016
|
|
5K6K
 
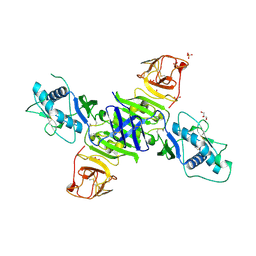 | | Zika virus non-structural protein 1 (NS1) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, GLYCEROL, SULFATE ION, ... | | Authors: | Akey, D.L, Brown, W.C, Smith, J.L. | | Deposit date: | 2016-05-24 | | Release date: | 2016-07-06 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.89 Å) | | Cite: | Extended surface for membrane association in Zika virus NS1 structure.
Nat.Struct.Mol.Biol., 23, 2016
|
|
5KP8
 
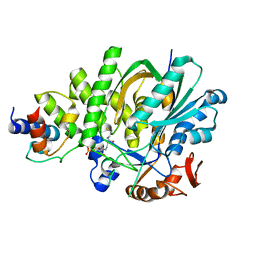 | |
5KP5
 
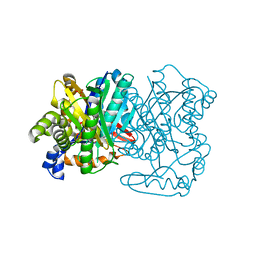 | |
5KP7
 
 | |
5KP6
 
 | |
1VF5
 
 | | Crystal Structure of Cytochrome b6f Complex from M.laminosus | | Descriptor: | (7R,17E)-4-HYDROXY-N,N,N,7-TETRAMETHYL-7-[(8E)-OCTADEC-8-ENOYLOXY]-10-OXO-3,5,9-TRIOXA-4-PHOSPHAHEPTACOS-17-EN-1-AMINIUM 4-OXIDE, 2,3-DIMETHYL-5-(3,7,11,15,19,23,27,31,35-NONAMETHYL-2,6,10,14,18,22,26,30,34-HEXATRIACONTANONAENYL-2,5-CYCLOHEXADIENE-1,4-DIONE-2,3-DIMETHYL-5-SOLANESYL-1,4-BENZOQUINONE, 8-HYDROXY-5,7-DIMETHOXY-3-METHYL-2-TRIDECYL-4H-CHROMEN-4-ONE, ... | | Authors: | Kurisu, G, Zhang, H, Smith, J.L, Cramer, W.A. | | Deposit date: | 2004-04-08 | | Release date: | 2004-04-20 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structure of the Cytochrome B6F Complex of Oxygenic Photosynthesis: Tuning the Cavity
Science, 302, 2003
|
|
6ECW
 
 | | StiD O-MT residues 956-1266 | | Descriptor: | S-ADENOSYL-L-HOMOCYSTEINE, StiD protein | | Authors: | Skiba, M.A, Bivins, M.M, Smith, J.L. | | Deposit date: | 2018-08-08 | | Release date: | 2018-12-12 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural Basis of Polyketide Synthase O-Methylation.
ACS Chem. Biol., 13, 2018
|
|
6ECX
 
 | | StiE O-MT residues 942-1257 | | Descriptor: | GLYCEROL, S-ADENOSYLMETHIONINE, StiE protein | | Authors: | Skiba, M.A, Bivins, M.B, Smith, J.L. | | Deposit date: | 2018-08-08 | | Release date: | 2018-12-12 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural Basis of Polyketide Synthase O-Methylation.
ACS Chem. Biol., 13, 2018
|
|
6ECV
 
 | | StiD O-MT residues 976-1266 | | Descriptor: | CHLORIDE ION, S-ADENOSYL-L-HOMOCYSTEINE, StiD protein | | Authors: | Skiba, M.A, Bivins, M.M, Smith, J.L. | | Deposit date: | 2018-08-08 | | Release date: | 2018-12-12 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.798 Å) | | Cite: | Structural Basis of Polyketide Synthase O-Methylation.
ACS Chem. Biol., 13, 2018
|
|
6ECT
 
 | | StiE O-MT residues 961-1257 | | Descriptor: | S-ADENOSYLMETHIONINE, StiE protein | | Authors: | Skiba, M.A, Bivins, M.M, Smith, J.L. | | Deposit date: | 2018-08-08 | | Release date: | 2018-12-12 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.42 Å) | | Cite: | Structural Basis of Polyketide Synthase O-Methylation.
ACS Chem. Biol., 13, 2018
|
|
6ECU
 
 | |
8SY3
 | |
8SY2
 
 | |
7UCL
 
 | |
7UCI
 
 | | SxtA Methyltransferase and decarboxylase didomain in complex with Mn2+ and SAH | | Descriptor: | GLYCEROL, MANGANESE (II) ION, Polyketide synthase-related protein, ... | | Authors: | Lao, Y, Skiba, M.A, Smith, J.L. | | Deposit date: | 2022-03-16 | | Release date: | 2022-06-01 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structural Basis for Control of Methylation Extent in Polyketide Synthase Metal-Dependent C -Methyltransferases.
Acs Chem.Biol., 17, 2022
|
|
6MBG
 
 | |
6MBF
 | | GphF Dehydratase 1 | | Descriptor: | GphF Dehydratase 1, MAGNESIUM ION | | Authors: | Dodge, G.J, Smith, J.L. | | Deposit date: | 2018-08-29 | | Release date: | 2018-09-19 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.543 Å) | | Cite: | Molecular Basis for Olefin Rearrangement in the Gephyronic Acid Polyketide Synthase.
ACS Chem. Biol., 13, 2018
|
|
6MBH
 
 | |
6MFD
 
 | | GphF GNAT-like decarboxylase in complex with isobutyryl-CoA | | Descriptor: | ACETATE ION, GLYCEROL, GphF, ... | | Authors: | Skiba, M.A, Tran, C.L, Smith, J.L. | | Deposit date: | 2018-09-10 | | Release date: | 2019-09-18 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.794 Å) | | Cite: | Repurposing the GNAT Fold in the Initiation of Polyketide Biosynthesis.
Structure, 28, 2020
|
|
