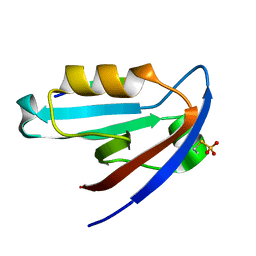
4Q6H
 | |
4QAO
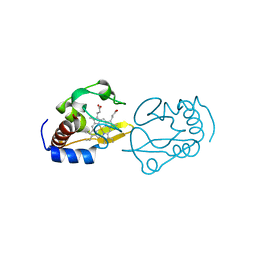 
 | | Lysine-ligated cytochrome c with F82H | | Descriptor: | Cytochrome c iso-1, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Amacher, J.F, Zhu, M.Q, Zhong, F, Pletneva, E.K, Madden, D.R. | | Deposit date: | 2014-05-05 | | Release date: | 2015-08-05 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.103 Å) | | Cite: | Understanding PDZ Affinity and Selectivity: All Residues Considered
Thesis, 2014
|
|
4Q6S
 
 | |
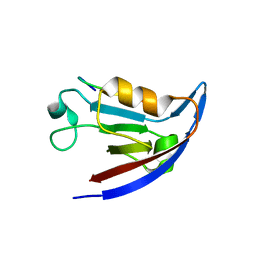
3SFJ
 | |
4NNL
 
 | |
4NNM
 
 | |
7S51
 
 | | Structure of C208A Sortase A from Streptococcus pyogenes bound to LPATA peptide | | Descriptor: | LEU-PRO-ALA-THR-ALA, Sortase | | Authors: | Johnson, D.A, Svendsen, J.E, Antos, J.M, Amacher, J.F. | | Deposit date: | 2021-09-09 | | Release date: | 2022-09-07 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Structures of Streptococcus pyogenes class A sortase in complex with substrate and product mimics provide key details of target recognition.
J.Biol.Chem., 298, 2022
|
|
7S4O
 
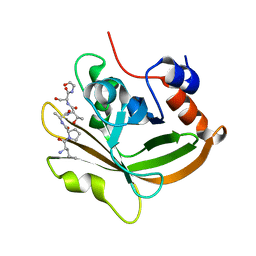 | | Structure of C208A Sortase A from Streptococcus pyogenes bound to LPATS peptide | | Descriptor: | LEU-PRO-ALA-THR-SER-GLY, Sortase | | Authors: | Johnson, D.A, Svendsen, J.E, Antos, J.M, Amacher, J.F. | | Deposit date: | 2021-09-09 | | Release date: | 2022-09-07 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.396 Å) | | Cite: | Structures of Streptococcus pyogenes class A sortase in complex with substrate and product mimics provide key details of target recognition.
J.Biol.Chem., 298, 2022
|
|
8SNQ
 
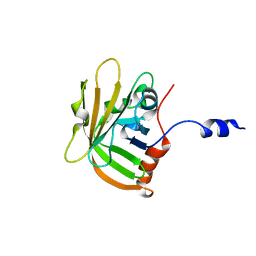 | |
8T8G
 
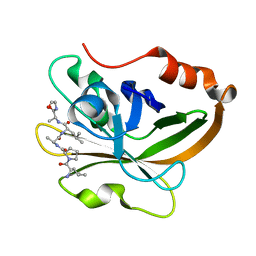 | |
7T8Y
 
 | | Structure of Class A sortase from Streptococcus pyogenes bound to lipid II mimetic, LPATA: Thr-in conformation | | Descriptor: | ACE-ALA-ZGL-LYS-DAL-DAL, BE2-LEU-PRO-ALA-THR-ALA-ALA, Sortase | | Authors: | Piper, I.M, Antos, J.M, Amacher, J.F. | | Deposit date: | 2021-12-17 | | Release date: | 2022-09-07 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structures of Streptococcus pyogenes class A sortase in complex with substrate and product mimics provide key details of target recognition.
J.Biol.Chem., 298, 2022
|
|
7T8Z
 
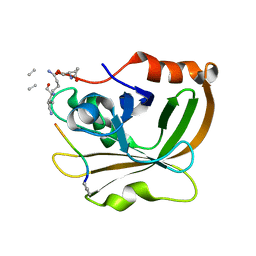 | | Structure of Class A sortase from Streptococcus pyogenes bound to lipid II mimetic, LPATA: Thr-out conformation | | Descriptor: | ACE-ALA-ZGL-LYS-DAL-DAL, BE2-LEU-PRO-ALA-THR-ALA-ALA, Sortase | | Authors: | Piper, I.M, Antos, J.M, Amacher, J.F. | | Deposit date: | 2021-12-17 | | Release date: | 2022-09-07 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structures of Streptococcus pyogenes class A sortase in complex with substrate and product mimics provide key details of target recognition.
J.Biol.Chem., 298, 2022
|
|
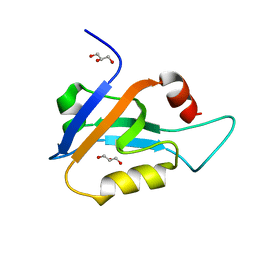
6X1N
 
 | |
6X20
 
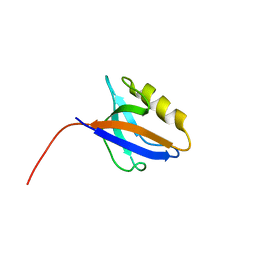 | |
6X1X
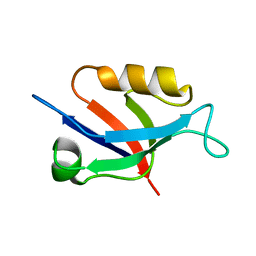
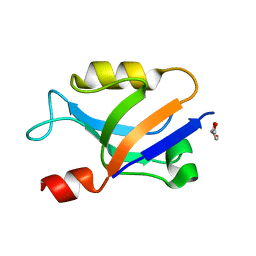
 | | PDZ domain from choanoflagellate GIPC (mbGIPC) | | Descriptor: | GLYCEROL, mbGIPC protein | | Authors: | Gao, M, Amacher, J.F. | | Deposit date: | 2020-05-19 | | Release date: | 2020-11-25 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.198 Å) | | Cite: | Structural characterization and computational analysis of PDZ domains in Monosiga brevicollis.
Protein Sci., 29, 2020
|
|
6X1P
 
 | |
6X23
 | |
6X1R
 
 | |
6X22
 
 | |
7S56
 
 | | Sortase A from Streptococcus agalactiae, residues 79-247 | | Descriptor: | Class A sortase | | Authors: | Gao, M, Kodama, H.M, Antos, J.M, Amacher, J.F. | | Deposit date: | 2021-09-09 | | Release date: | 2022-01-12 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Structural and biochemical analyses of selectivity determinants in chimeric Streptococcus Class A sortase enzymes.
Protein Sci., 31, 2022
|
|
7S54
 
 | | Sortase A from Streptococcus agalactiae with the deltaN188 b7-b8 loop sequence from Staphylococcus aureus Sortase A | | Descriptor: | Class A sortase, sortase A chimera | | Authors: | Gao, M, Kodama, H.M, Antos, J.M, Amacher, J.F. | | Deposit date: | 2021-09-09 | | Release date: | 2022-01-12 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.794 Å) | | Cite: | Structural and biochemical analyses of selectivity determinants in chimeric Streptococcus Class A sortase enzymes.
Protein Sci., 31, 2022
|
|
7S57
 
 | | Structure of Sortase A from Streptococcus pyogenes with the b7-b8 loop sequence of Enterococcus faecalis Sortase A | | Descriptor: | Class A sortase, sortase A chimera | | Authors: | Svendsen, J.E, Johnson, D.A, Gao, M, Antos, J.M, Amacher, J.F. | | Deposit date: | 2021-09-09 | | Release date: | 2022-01-12 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural and biochemical analyses of selectivity determinants in chimeric Streptococcus Class A sortase enzymes.
Protein Sci., 31, 2022
|
|
7S53
 
 | | Structure of Sortase A from Streptococcus pyogenes with the b7-b8 loop sequence from Listeria monocytogenes Sortase A | | Descriptor: | Class A sortase, sortase A chimera | | Authors: | Johnson, D.A, Svendsen, J.E, Antos, J.M, Amacher, J.F. | | Deposit date: | 2021-09-09 | | Release date: | 2022-01-12 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structural and biochemical analyses of selectivity determinants in chimeric Streptococcus Class A sortase enzymes.
Protein Sci., 31, 2022
|
|
4DLN
 
 | |
