3K2G
 
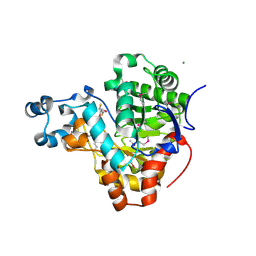 | | Crystal structure of a Resiniferatoxin-binding protein from Rhodobacter sphaeroides | | Descriptor: | (2S,3S)-1,4-DIMERCAPTOBUTANE-2,3-DIOL, MAGNESIUM ION, Resiniferatoxin-binding, ... | | Authors: | Kumaran, D, Burley, S.K, Swaminathan, S, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2009-09-30 | | Release date: | 2009-10-13 | | Last modified: | 2021-02-10 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of a Resiniferatoxin-binding protein from Rhodobacter sphaeroides
To be Published
|
|
3JUL
 
 | |
3K17
 
 | |
3K85
 
 | |
3K9E
 
 | |
3K5W
 
 | |
3KZH
 
 | |
3KZB
 
 | |
3K9C
 
 | |
3KZG
 
 | |
3L0Q
 
 | |
3HZ4
 
 | |
3I3V
 
 | |
3HS3
 
 | |
3HZ6
 
 | |
3IBQ
 
 | | Crystal structure of pyridoxal kinase from Lactobacillus plantarum in complex with ATP | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, MAGNESIUM ION, Pyridoxal kinase | | Authors: | Bagaria, A, Kumaran, D, Burley, S.K, Swaminathan, S, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2009-07-16 | | Release date: | 2009-07-28 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of pyridoxal kinase from Lactobacillus plantarum in complex with ATP
To be Published
|
|
3I8B
 
 | |
3LJS
 
 | |
3LM4
 | | Crystal Structure of 2,3-Dihydroxy Biphenyl dioxygenase from Rhodococcus sp. (strain RHA1) | | Descriptor: | (2Z,4E)-2-HYDROXY-6-OXO-6-PHENYLHEXA-2,4-DIENOIC ACID, Catechol 2,3-dioxygenase, FE (III) ION, ... | | Authors: | Syed Ibrahim, B, Kumaran, D, Burley, S.K, Swaminathan, S, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2010-01-29 | | Release date: | 2010-02-23 | | Last modified: | 2021-02-10 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal Structure of 2,3-Dihydroxy Biphenyl dioxygenase from Rhodococcus sp. (strain RHA1)
To be Published
|
|
3HUU
 
 | |
3I3Y
 
 | |
3HV1
 
 | |
3HYO
 
 | | Crystal structure of pyridoxal kinase from Lactobacillus plantarum in complex with ADP | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, Pyridoxal kinase | | Authors: | Bagaria, A, Kumaran, D, Burley, S.K, Swaminathan, S, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2009-06-22 | | Release date: | 2009-06-30 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Crystal structure of pyridoxal kinase from Lactobacillus plantarum in complex with ADP
To be Published
|
|
3HUT
 
 | |
3LDT
 
 | |
