1UE7
 
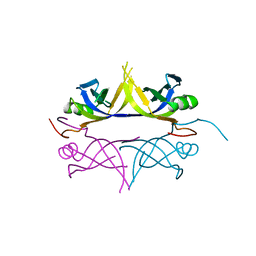 | | Crystal structure of the single-stranded dna-binding protein from mycobacterium tuberculosis | | Descriptor: | Single-strand binding protein | | Authors: | Saikrishnan, K, Jeyakanthan, J, Venkatesh, J, Acharya, N, Sekar, K, Varshney, U, Vijayan, M, TB Structural Genomics Consortium (TBSGC) | | Deposit date: | 2003-05-09 | | Release date: | 2004-02-10 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Structure of Mycobacterium tuberculosis single-stranded DNA-binding protein. Variability in quaternary structure and its implications
J.MOL.BIOL., 331, 2003
|
|
9KYO
 
 | | GES bound mTAUT | | Descriptor: | CHLORIDE ION, Heavy chain of 9D5 fab, Light chain of 9D5 fab, ... | | Authors: | She, J, Wang, M.X, He, J. | | Deposit date: | 2024-12-09 | | Release date: | 2025-07-02 | | Method: | ELECTRON MICROSCOPY (2.84 Å) | | Cite: | Molecular basis for substrate recognition and transport of mammalian taurine transporters
To Be Published
|
|
2V10
 
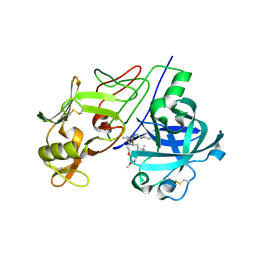 | | Crystal Structure of Renin with Inhibitor 9 | | Descriptor: | (2R,4S,5S,7S)-5-AMINO-N-BUTYL-4-HYDROXY-7-[4-METHOXY-3-(3-METHOXYPROPOXY)BENZYL]-2,8-DIMETHYLNONANAMIDE, RENIN | | Authors: | Rahuel, J, Rasetti, V, Maibaum, J, Rueger, H, Goschke, R, Cohen, N.C, Stutz, S, Cumin, F, Fuhrer, W, Wood, J.M, Grutter, M.G. | | Deposit date: | 2007-05-21 | | Release date: | 2007-07-03 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Structure-Based Drug Design: The Discovery of Novel Nonpeptide Orally Active Inhibitors of Human Renin
Chem.Biol., 7, 2000
|
|
9KMJ
 
 | | Taurine bound hTAUT | | Descriptor: | 2-AMINOETHANESULFONIC ACID, CHLORIDE ION, Heavy chain of 9D5 Fab, ... | | Authors: | She, J, Wang, M, He, J. | | Deposit date: | 2024-11-16 | | Release date: | 2025-07-02 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | Molecular basis for substrate recognition and transport of mammalian taurine transporters
To Be Published
|
|
9KMK
 
 | | apo mTAUT in inward open II state | | Descriptor: | CHLORIDE ION, Sodium- and chloride-dependent taurine transporter, heavy chain of 9D5 fab, ... | | Authors: | She, J, Wang, M, He, J. | | Deposit date: | 2024-11-16 | | Release date: | 2025-07-02 | | Method: | ELECTRON MICROSCOPY (3.06 Å) | | Cite: | Molecular basis for substrate recognition and transport of mammalian taurine transporters
To Be Published
|
|
9KMM
 
 | | IAA bound mTAUT | | Descriptor: | 2H-IMIDAZOL-4-YLACETIC ACID, CHLORIDE ION, SODIUM ION, ... | | Authors: | She, J, Wang, M, He, J. | | Deposit date: | 2024-11-16 | | Release date: | 2025-07-02 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Molecular basis for substrate recognition and transport of mammalian taurine transporters
To Be Published
|
|
9KMI
 
 | | Taurine bound mTAUT | | Descriptor: | 2-AMINOETHANESULFONIC ACID, CHLORIDE ION, Heavy chain of 9D5 Fab, ... | | Authors: | She, J, Wang, M, He, J. | | Deposit date: | 2024-11-16 | | Release date: | 2025-07-02 | | Method: | ELECTRON MICROSCOPY (3.05 Å) | | Cite: | Molecular basis for substrate recognition and transport of mammalian taurine transporters
To Be Published
|
|
4V6M
 
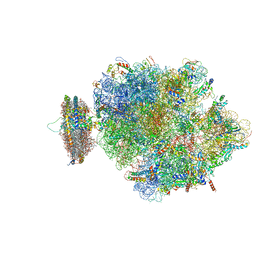 | | Structure of the ribosome-SecYE complex in the membrane environment | | Descriptor: | (1R)-2-{[{[(2S)-2,3-DIHYDROXYPROPYL]OXY}(HYDROXY)PHOSPHORYL]OXY}-1-[(PALMITOYLOXY)METHYL]ETHYL (11E)-OCTADEC-11-ENOATE, (1S)-2-{[(2-AMINOETHOXY)(HYDROXY)PHOSPHORYL]OXY}-1-[(PALMITOYLOXY)METHYL]ETHYL STEARATE, 16S RIBOSOMAL RNA, ... | | Authors: | Frauenfeld, J, Gumbart, J, van der Sluis, E.O, Funes, S, Gartmann, M, Beatrix, B, Mielke, T, Berninghausen, O, Becker, T, Schulten, K, Beckmann, R. | | Deposit date: | 2011-02-08 | | Release date: | 2014-07-09 | | Last modified: | 2024-02-28 | | Method: | ELECTRON MICROSCOPY (7.1 Å) | | Cite: | Cryo-EM structure of the ribosome-SecYE complex in the membrane environment.
Nat.Struct.Mol.Biol., 18, 2011
|
|
1KAO
 
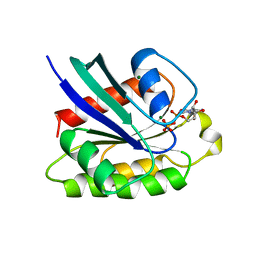 | | CRYSTAL STRUCTURE OF THE SMALL G PROTEIN RAP2A WITH GDP | | Descriptor: | GUANOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, RAP2A | | Authors: | Cherfils, J, Menetrey, J, Le Bras, G. | | Deposit date: | 1997-08-01 | | Release date: | 1997-12-24 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal structures of the small G protein Rap2A in complex with its substrate GTP, with GDP and with GTPgammaS.
EMBO J., 16, 1997
|
|
3SDX
 
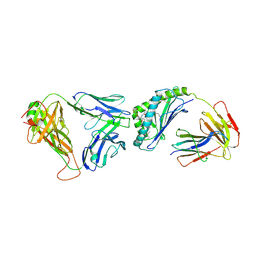 | | Crystal structure of human autoreactive-Valpha24 NKT TCR in complex with CD1d-beta-galactosylceramide | | Descriptor: | Antigen-presenting glycoprotein CD1d, Beta-2-microglobulin, N-[(2S,3R)-1-(beta-D-galactopyranosyloxy)-3-hydroxyoctadec-4-en-2-yl]tetracosanamide, ... | | Authors: | Clarke, A.J, Patel, O, Rossjohn, J. | | Deposit date: | 2011-06-09 | | Release date: | 2011-10-05 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (3.12 Å) | | Cite: | Recognition of beta-linked self glycolipids mediated by natural killer T cell antigen receptors
Nat.Immunol., 12, 2011
|
|
4UZC
 
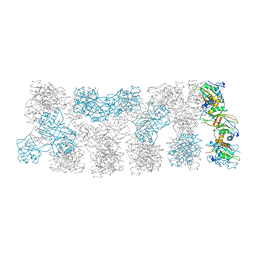 | |
1TQX
 
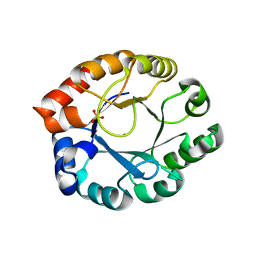 | | Crystal Structure of Pfal009167 A Putative D-Ribulose 5-Phosphate 3-Epimerase from P.falciparum | | Descriptor: | D-ribulose-5-phosphate 3-epimerase, putative, SULFATE ION, ... | | Authors: | Caruthers, J, Bosch, J, Hol, W.G.J, Structural Genomics of Pathogenic Protozoa Consortium (SGPP) | | Deposit date: | 2004-06-18 | | Release date: | 2004-12-21 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure of a ribulose 5-phosphate 3-epimerase from Plasmodium falciparum.
Proteins, 62, 2006
|
|
9MGY
 
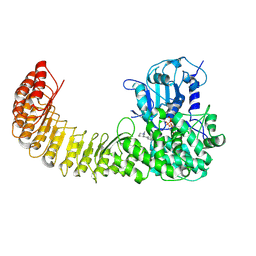 | | Cryo-EM structure of Human NLRP3 complex with compound 1 | | Descriptor: | (2M)-2-(6-{[(3R)-1-methylpiperidin-3-yl]amino}pyridazin-3-yl)-5-(trifluoromethyl)phenol, ADENOSINE-5'-TRIPHOSPHATE, NACHT, ... | | Authors: | Mammoliti, O, Carbajo, R.J, Perez-Benito, L, Yu, X, Prieri, M.L.C, Bontempi, L, Embrechts, S, Paesmans, I, Bassi, M, Bhattacharya, A, Roman, S.C, Hoog, S.D, Demin, S, Gijsen, H.J.M, Hache, G, Jacobs, T, Jerhaoui, S, Leenaerts, J, Lutter, F.H, Matico, R, Oehlrich, D, Perrier, M, Ryabchuk, P, Schepens, W, Sharma, S, Somers, M, Suarez, J, Surkyn, M, Opdenbosch, N.V, Verhulst, T, Bottelbergs, A. | | Deposit date: | 2024-12-11 | | Release date: | 2025-02-26 | | Last modified: | 2025-03-12 | | Method: | ELECTRON MICROSCOPY (2.9 Å) | | Cite: | Discovery of Potent and Brain-Penetrant Bicyclic NLRP3 Inhibitors with Peripheral and Central In Vivo Activity.
J.Med.Chem., 68, 2025
|
|
9MIG
 
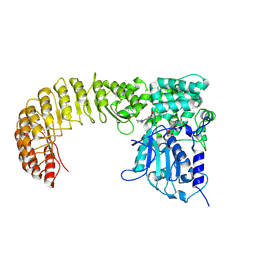 | | Cryo-EM structure of Human NLRP3 complex with compound 3 | | Descriptor: | (2P)-2-(4-{[(3R)-1-methylpiperidin-3-yl]amino}-5,6,7,8-tetrahydrophthalazin-1-yl)-5-(trifluoromethyl)phenol, ADENOSINE-5'-TRIPHOSPHATE, NACHT, ... | | Authors: | Mammoliti, O, Carbajo, R.J, Perez-Benito, L, Yu, X, Prieri, M.L.C, Bontempi, L, Embrechts, S, Paesmans, I, Bassi, M, Bhattacharya, A, Roman, S.C, Hoog, S.D, Demin, S, Gijsen, H.J.M, Hache, G, Jacobs, T, Jerhaoui, S, Leenaerts, J, Lutter, F.H, Matico, R, Oehlrich, D, Perrier, M, Ryabchuk, P, Schepens, W, Sharma, S, Somers, M, Suarez, J, Surkyn, M, Opdenbosch, N.V, Verhulst, T, Bottelbergs, A. | | Deposit date: | 2024-12-12 | | Release date: | 2025-02-26 | | Last modified: | 2025-03-12 | | Method: | ELECTRON MICROSCOPY (3.6 Å) | | Cite: | Discovery of Potent and Brain-Penetrant Bicyclic NLRP3 Inhibitors with Peripheral and Central In Vivo Activity.
J.Med.Chem., 68, 2025
|
|
4V96
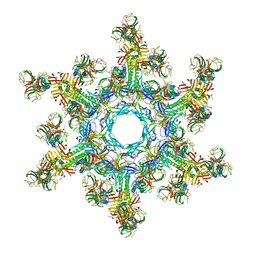 
 | | The structure of a 1.8 MDa viral genome injection device suggests alternative infection mechanisms | | Descriptor: | BPP, ORF46, ORF48 | | Authors: | Veesler, D, Spinelli, S, Mahony, J, Lichiere, J, Blangy, S, Bricogne, G, Legrand, P, Ortiz-Lombardia, M, Campanacci, V, van Sinderen, D, Cambillau, C. | | Deposit date: | 2012-02-01 | | Release date: | 2014-07-09 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (3.8 Å) | | Cite: | Structure of the phage TP901-1 1.8 MDa baseplate suggests an alternative host adhesion mechanism.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
4V6N
 
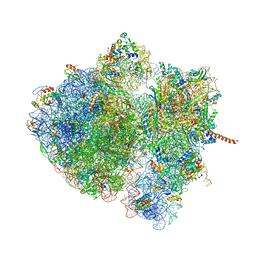 | | Structural characterization of mRNA-tRNA translocation intermediates (50S ribosome of class2 of the six classes) | | Descriptor: | 16S ribosomal RNA, 23S ribosomal RNA, 30S ribosomal protein S10, ... | | Authors: | Agirrezabala, X, Liao, H, Schreiner, E, Fu, J, Ortiz-Meoz, R.F, Schulten, K, Green, R, Frank, J. | | Deposit date: | 2011-12-07 | | Release date: | 2014-07-09 | | Last modified: | 2025-03-19 | | Method: | ELECTRON MICROSCOPY (12.1 Å) | | Cite: | Structural characterization of mRNA-tRNA translocation intermediates.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
1UE5
 
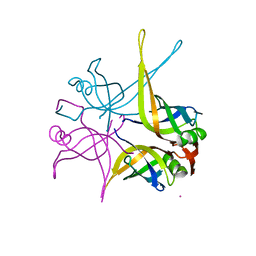 | | Crystal structure of the single-stranded dna-binding protein from mycobacterium tuberculosis | | Descriptor: | CADMIUM ION, Single-strand binding protein | | Authors: | Saikrishnan, K, Jeyakanthan, J, Venkatesh, J, Acharya, N, Sekar, K, Varshney, U, Vijayan, M, TB Structural Genomics Consortium (TBSGC) | | Deposit date: | 2003-05-09 | | Release date: | 2004-02-10 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structure of Mycobacterium tuberculosis single-stranded DNA-binding protein. Variability in quaternary structure and its implications
J.MOL.BIOL., 331, 2003
|
|
5XWR
 
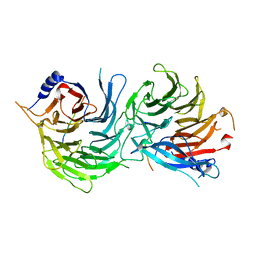 | | Crystal Structure of RBBP4-peptide complex | | Descriptor: | Histone-binding protein RBBP4, MET-SER-ARG-ARG-LYS-GLN-ALA-LYS-PRO-GLN-HIS-ILE | | Authors: | Jobichen, C, Lui, B.H, Daniel, G.T, Sivaraman, J. | | Deposit date: | 2017-06-30 | | Release date: | 2018-07-11 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.69 Å) | | Cite: | Targeting cancer addiction for SALL4 by shifting its transcriptome with a pharmacologic peptide.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
1UE6
 
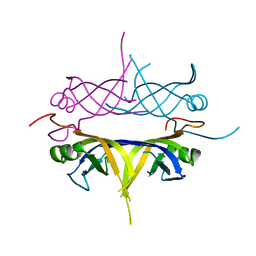 | | Crystal structure of the single-stranded dna-binding protein from mycobacterium tuberculosis | | Descriptor: | Single-strand binding protein | | Authors: | Saikrishnan, K, Jeyakanthan, J, Venkatesh, J, Acharya, N, Sekar, K, Varshney, U, Vijayan, M, TB Structural Genomics Consortium (TBSGC) | | Deposit date: | 2003-05-09 | | Release date: | 2004-02-10 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structure of Mycobacterium tuberculosis single-stranded DNA-binding protein. Variability in quaternary structure and its implications
J.MOL.BIOL., 331, 2003
|
|
6TFW
 
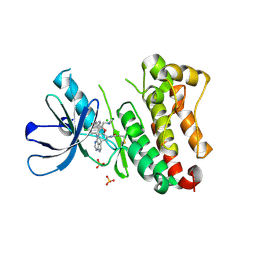 | | Crystal Structure of EGFR T790M/V948R in Complex with Covalent Pyrrolopyrimidine 18d | | Descriptor: | CHLORIDE ION, Epidermal growth factor receptor, MAGNESIUM ION, ... | | Authors: | Niggenaber, J, Mueller, M.P, Rauh, D. | | Deposit date: | 2019-11-14 | | Release date: | 2020-09-30 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Targeting Her2-insYVMA with Covalent Inhibitors-A Focused Compound Screening and Structure-Based Design Approach.
J.Med.Chem., 63, 2020
|
|
9XIM
 
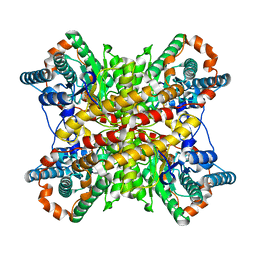 | |
4L8D
 
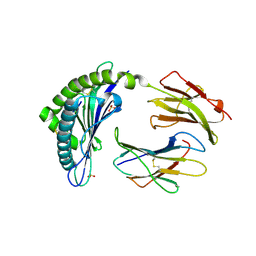 | | Crystal structure of the H2Db in complex with the NP-N5D peptide | | Descriptor: | Beta-2-microglobulin, H-2 class I histocompatibility antigen, D-B alpha chain, ... | | Authors: | Rossjohn, J, Gras, S. | | Deposit date: | 2013-06-16 | | Release date: | 2013-10-16 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Acute emergence and reversion of influenza A virus quasispecies within CD8(+) T cell antigenic peptides.
Nat Commun, 4, 2013
|
|
4W2E
 
 | | Crystal structure of Elongation Factor 4 (EF4/LepA) bound to the Thermus thermophilus 70S ribosome | | Descriptor: | 16S Ribosomal RNA, 23S Ribosomal RNA, 30S ribosomal protein S10, ... | | Authors: | Gagnon, M.G, Lin, J, Steitz, T.A. | | Deposit date: | 2014-06-04 | | Release date: | 2014-10-01 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Crystal structure of elongation factor 4 bound to a clockwise ratcheted ribosome.
Science, 345, 2014
|
|
4W5S
 
 | | Tankyrase in complex with compound | | Descriptor: | 8-(hydroxymethyl)-2-[4-(1-methyl-1H-pyrazol-4-yl)phenyl]quinazolin-4(3H)-one, GLYCEROL, Tankyrase-1, ... | | Authors: | Johannes, J, Kazmirski, S.L, Boriack-Sjodin, P.A, Howard, T. | | Deposit date: | 2014-08-18 | | Release date: | 2015-05-13 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Pyrimidinone nicotinamide mimetics as selective tankyrase and wnt pathway inhibitors suitable for in vivo pharmacology.
Acs Med.Chem.Lett., 6, 2015
|
|
4UUW
 
 | |
