3T81
 
 | |
3PZL
 
 | |
3EXQ
 
 | |
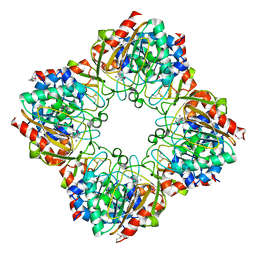
3FIJ
 | |
3F6A
 
 | |
3TFW
 
 | |
3H5O
 
 | |
3GRC
 
 | |
3FIE
 
 | |
3FND
 
 | |
3GV1
 
 | |
3GL3
 | |
3GV0
 
 | |
3OR5
 | |
3OXN
 | |
3DEC
 
 | | Crystal structure of a glycosyl hydrolases family 2 protein from Bacteroides thetaiotaomicron | | Descriptor: | Beta-galactosidase, POTASSIUM ION | | Authors: | Kumaran, D, Bonanno, J, Romero, R, Burley, S.K, Swaminathan, S, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2008-06-09 | | Release date: | 2008-06-17 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal Structure of a Glycosyl Hydrolases Family 2 protein from Bacteroides thetaiotaomicron.
To be Published
|
|
3SMD
 
 | |
3DDA
 
 | |
3H75
 | | Crystal Structure of a Periplasmic Sugar-binding protein from the Pseudomonas fluorescens | | Descriptor: | GLYCEROL, Periplasmic sugar-binding domain protein, SULFATE ION | | Authors: | Kumaran, D, Mahmood, A, Burley, S.K, Swaminathan, S, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2009-04-24 | | Release date: | 2009-05-12 | | Last modified: | 2021-02-10 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystal Structure of a Periplasmic Sugar-binding protein from the Pseudomonas fluorescens
To be Published
|
|
3H6E
 
 | |
3H74
 
 | | Crystal structure of pyridoxal kinase from Lactobacillus plantarum | | Descriptor: | GLYCEROL, Pyridoxal kinase, SULFATE ION | | Authors: | Bagaria, A, Kumaran, D, Burley, S.K, Swaminathan, S, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2009-04-24 | | Release date: | 2009-05-26 | | Last modified: | 2021-02-10 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Crystal structure of pyridoxal kinase from Lactobacillus plantarum
To be Published
|
|
3HH0
 
 | | Crystal structure of a transcriptional regulator, MerR family from Bacillus cereus | | Descriptor: | Transcriptional regulator, MerR family | | Authors: | Palani, K, Zhang, Z, Burley, S.K, Swaminathan, S, New York SGX Research Center for Structural Genomics, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2009-05-14 | | Release date: | 2009-05-26 | | Last modified: | 2021-02-10 | | Method: | X-RAY DIFFRACTION (2.67 Å) | | Cite: | Crystal structure of a transcriptional regulator, MerR family from Bacillus cereus
To be Published
|
|
3HIC
 
 | |
3HDO
 
 | |
3HD5
 
 | |
