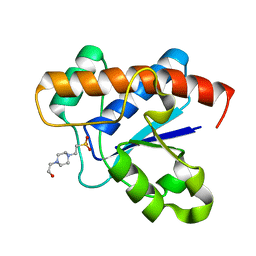
3JS5
 
 | |
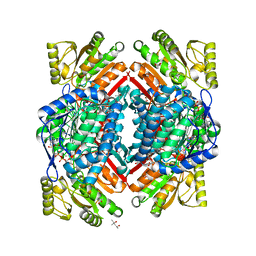
5KF0
 
 | |
3K2H
 
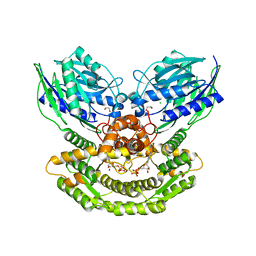
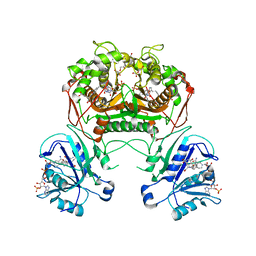 | | Co-crystal structure of dihydrofolate reductase/thymidylate synthase from Babesia bovis with dUMP, Pemetrexed and NADP | | Descriptor: | 1,2-ETHANEDIOL, 2'-DEOXYURIDINE 5'-MONOPHOSPHATE, 2-{4-[2-(2-AMINO-4-OXO-4,7-DIHYDRO-3H-PYRROLO[2,3-D]PYRIMIDIN-5-YL)-ETHYL]-BENZOYLAMINO}-PENTANEDIOIC ACID, ... | | Authors: | Seattle Structural Genomics Center for Infectious Disease (SSGCID) | | Deposit date: | 2009-09-30 | | Release date: | 2009-10-13 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Inhibitor-bound complexes of dihydrofolate reductase-thymidylate synthase from Babesia bovis.
Acta Crystallogr.,Sect.F, 67, 2011
|
|
7RZM
 
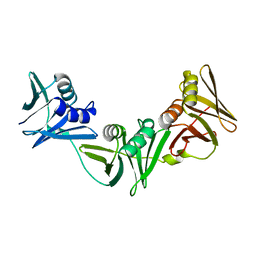 | |
7RZO
 
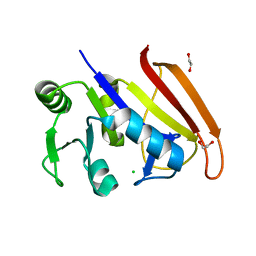 | |
5K9F
 
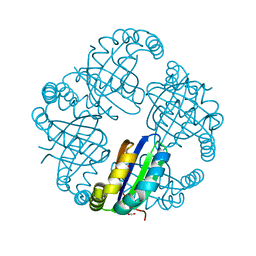 | |
3K9W
 
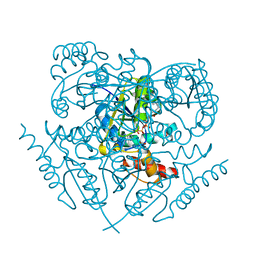 | |
5JSC
 
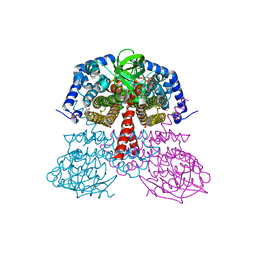 | |
5KHA
 
 | |
6CK0
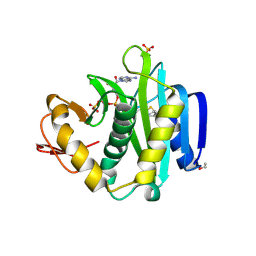 
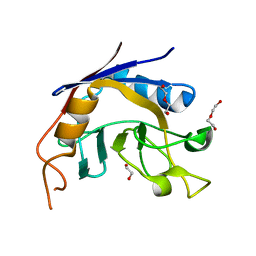
 | | Crystal Structure of Biotin Acetyl Coenzyme A Carboxylase Synthetase from Helicobacter pylori with bound Biotinylated ATP | | Descriptor: | 1,2-ETHANEDIOL, 5'-O-[(S)-({5-[(2R,3aS,4S,6aR)-2-hydroxyhexahydro-1H-thieno[3,4-d]imidazol-4-yl]pentanoyl}oxy){[(S)-hydroxy(phosphonooxy)phosphoryl]oxy}phosphoryl]adenosine, Biotin acetyl coenzyme A carboxylase synthetase, ... | | Authors: | Seattle Structural Genomics Center for Infectious Disease (SSGCID) | | Deposit date: | 2018-02-27 | | Release date: | 2018-04-04 | | Last modified: | 2025-01-15 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Co-crystal structure of Helicobacter pylori biotin protein ligase with biotinyl-5-ATP.
Acta Crystallogr.,Sect.F, 81, 2025
|
|
3S6M
 
 | |
5KIA
 | |
4WHX
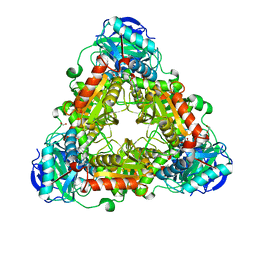 
 | | X-ray Crystal Structure of an Amino Acid Aminotransferase from Burkholderia pseudomallei Bound to the Co-factor Pyridoxal Phosphate | | Descriptor: | 1,2-ETHANEDIOL, ALANINE, Branched-chain-amino-acid transaminase | | Authors: | Seattle Structural Genomics Center for Infectious Disease (SSGCID), Fairman, J.W, Dranow, D.M, Taylor, B.M, Lorimer, D, Edwards, T.E. | | Deposit date: | 2014-09-23 | | Release date: | 2014-10-08 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | X-ray Crystal Structure of an Amino Acid Aminotransferase from Burkholderia pseudomallei Bound to the Co-factor Pyridoxal Phosphate
to be published
|
|
7S5N
 
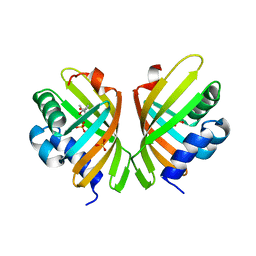 | |
7S5M
 
 | |
6CNZ
 
 | |
4W5K
 
 | |
4WJM
 
 | |
4WGJ
 
 | |
6V91
 
 | |
6V3M
 
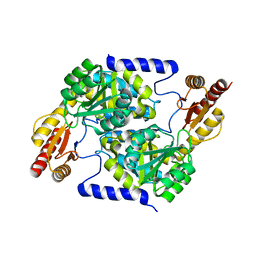
 | | Crystal structure of 2C-methyl-D-erythritol 2,4-cyclodiphosphate synthase (IspF) Burkholderia pseudomallei in compomplex with ligand HGN-0961 (BSI110840) | | Descriptor: | 1,2-ETHANEDIOL, 2-C-methyl-D-erythritol 2,4-cyclodiphosphate synthase, 5-{[(propan-2-yl)carbamoyl]amino}-1,3,4-thiadiazole-2-sulfonamide, ... | | Authors: | Seattle Structural Genomics Center for Infectious Disease (SSGCID) | | Deposit date: | 2019-11-26 | | Release date: | 2020-12-09 | | Last modified: | 2025-04-09 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Analysis of Burkholderia pseudomallei IspF in complex with sulfapyridine, sulfamonomethoxine, ethoxzolamide and acetazolamide.
Acta Crystallogr.,Sect.F, 81, 2025
|
|
6V77
 
 | |
6UZI
 
 | |
6UYH
 
 | |
6VH5
 
 | |
