1A0O
 
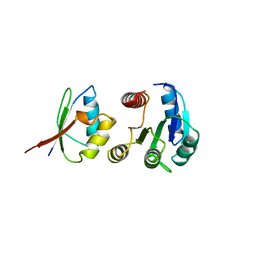 | | CHEY-BINDING DOMAIN OF CHEA IN COMPLEX WITH CHEY | | Descriptor: | CHEA, CHEY, MANGANESE (II) ION | | Authors: | Chinardet, N, Welch, M, Mourey, L, Birck, C, Samama, J.P. | | Deposit date: | 1997-12-05 | | Release date: | 1998-12-30 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Structure of the CheY-binding domain of histidine kinase CheA in complex with CheY.
Nat.Struct.Biol., 5, 1998
|
|
7OO1
 
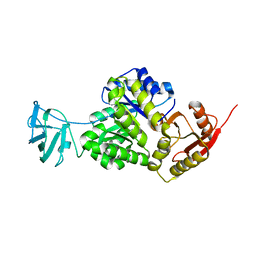 | | Structure, function and characterization of a second pyruvate kinase isozyme in Pseudomonas aeruginosa. | | Descriptor: | Pyruvate kinase | | Authors: | Abdelhamid, Y, Wang, M, Parkhill, S, Brear, P, Welch, M. | | Deposit date: | 2021-05-26 | | Release date: | 2021-11-03 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (3.01 Å) | | Cite: | Structure, Function and Regulation of a Second Pyruvate Kinase Isozyme in Pseudomonas aeruginosa
Front Microbiol, 12, 2021
|
|
1FFS
 
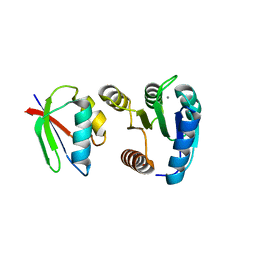 | | CHEY-BINDING DOMAIN OF CHEA IN COMPLEX WITH CHEY FROM CRYSTALS SOAKED IN ACETYL PHOSPHATE | | Descriptor: | CHEMOTAXIS PROTEIN CHEA, CHEMOTAXIS PROTEIN CHEY, MANGANESE (II) ION | | Authors: | Gouet, P, Chinardet, N, Welch, M, Guillet, V, Birck, C, Mourey, L, Samama, J.-P. | | Deposit date: | 2000-07-26 | | Release date: | 2001-01-17 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Further insights into the mechanism of function of the response regulator CheY from crystallographic studies of the CheY--CheA(124--257) complex.
Acta Crystallogr.,Sect.D, 57, 2001
|
|
1FFW
 
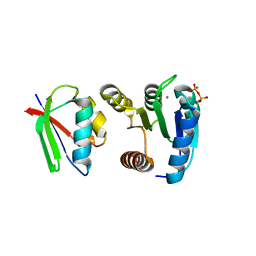 | | CHEY-BINDING DOMAIN OF CHEA IN COMPLEX WITH CHEY WITH A BOUND IMIDO DIPHOSPHATE | | Descriptor: | CHEMOTAXIS PROTEIN CHEA, CHEMOTAXIS PROTEIN CHEY, IMIDO DIPHOSPHATE, ... | | Authors: | Gouet, P, Chinardet, N, Welch, M, Guillet, V, Birck, C, Mourey, L, Samama, J.-P. | | Deposit date: | 2000-07-26 | | Release date: | 2001-01-17 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Further insights into the mechanism of function of the response regulator CheY from crystallographic studies of the CheY--CheA(124--257) complex.
Acta Crystallogr.,Sect.D, 57, 2001
|
|
1FFG
 
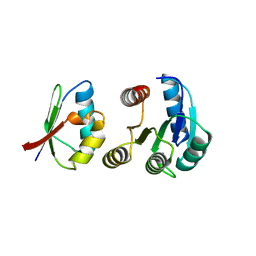 | | CHEY-BINDING DOMAIN OF CHEA IN COMPLEX WITH CHEY AT 2.1 A RESOLUTION | | Descriptor: | CHEMOTAXIS PROTEIN CHEA, CHEMOTAXIS PROTEIN CHEY, MANGANESE (II) ION | | Authors: | Gouet, P, Chinardet, N, Welch, M, Guillet, V, Birck, C, Mourey, L, Samama, J.-P. | | Deposit date: | 2000-07-25 | | Release date: | 2001-01-17 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Further insights into the mechanism of function of the response regulator CheY from crystallographic studies of the CheY--CheA(124--257) complex.
Acta Crystallogr.,Sect.D, 57, 2001
|
|
6ZU0
 
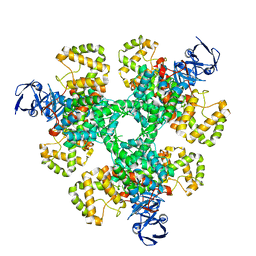 | |
6G3U
 
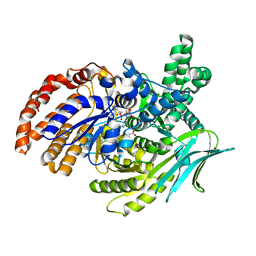 | | Structure of Pseudomonas aeruginosa Isocitrate Dehydrogenase, IDH | | Descriptor: | 2-OXOGLUTARIC ACID, Isocitrate dehydrogenase, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | Crousilles, A, Welch, M. | | Deposit date: | 2018-03-26 | | Release date: | 2018-08-01 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.707 Å) | | Cite: | Gluconeogenic precursor availability regulates flux through the glyoxylate shunt inPseudomonas aeruginosa.
J. Biol. Chem., 293, 2018
|
|
5OAS
 
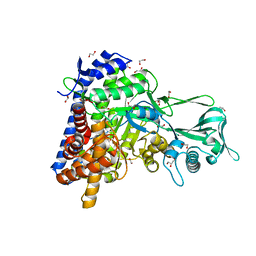 | |
5M2E
 
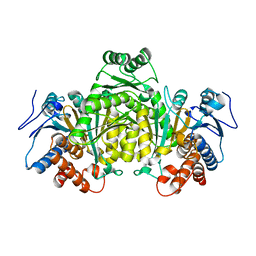 | |
8PU5
 
 | | Crystal structure of the Acyl-CoA dehydrogenase FadE1(PA0506) E441A from Pseudomonas aeruginosa complexed with C16CoA | | Descriptor: | 1,2-ETHANEDIOL, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, NITRATE ION, ... | | Authors: | Wang, M, Brear, P, Welch, M. | | Deposit date: | 2023-07-16 | | Release date: | 2025-02-12 | | Last modified: | 2025-05-21 | | Method: | X-RAY DIFFRACTION (1.44 Å) | | Cite: | Pseudomonas aeruginosa acyl-CoA dehydrogenases and structure-guided inversion of their substrate specificity.
Nat Commun, 16, 2025
|
|
8PNG
 
 | | Crystal structure of the apo acyl-CoA dehydrogenase FadE2 (PA0508) from Pseudomonas aeruginosa | | Descriptor: | DI(HYDROXYETHYL)ETHER, FLAVIN-ADENINE DINUCLEOTIDE, GLYCEROL, ... | | Authors: | Wang, M, Brear, P, Welch, M. | | Deposit date: | 2023-06-30 | | Release date: | 2025-01-15 | | Last modified: | 2025-05-21 | | Method: | X-RAY DIFFRACTION (1.94 Å) | | Cite: | Pseudomonas aeruginosa acyl-CoA dehydrogenases and structure-guided inversion of their substrate specificity.
Nat Commun, 16, 2025
|
|
8PNS
 
 | | Crystal structure of the acyl-CoA dehydrogenase PA0506 (FadE1) from Pseudomonas aeruginosa | | Descriptor: | Acyl-CoA dehydrogenase, DI(HYDROXYETHYL)ETHER, FLAVIN-ADENINE DINUCLEOTIDE, ... | | Authors: | Wang, M, Brear, P, Welch, M. | | Deposit date: | 2023-06-30 | | Release date: | 2025-01-15 | | Last modified: | 2025-05-21 | | Method: | X-RAY DIFFRACTION (2.08 Å) | | Cite: | Pseudomonas aeruginosa acyl-CoA dehydrogenases and structure-guided inversion of their substrate specificity.
Nat Commun, 16, 2025
|
|
8R1E
 
 | | crystal structure of acyl-CoA dehydrogenase FadE2 mutant (PA0508 M130G E296A Q303A) from Pseudomonas aeruginosa | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, GLYCEROL, Probable acyl-CoA dehydrogenase, ... | | Authors: | Wang, M, Brear, P, Welch, M. | | Deposit date: | 2023-11-01 | | Release date: | 2025-02-19 | | Last modified: | 2025-05-21 | | Method: | X-RAY DIFFRACTION (1.62 Å) | | Cite: | Pseudomonas aeruginosa acyl-CoA dehydrogenases and structure-guided inversion of their substrate specificity.
Nat Commun, 16, 2025
|
|
6S6F
 
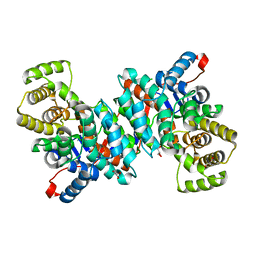 | |
6T5M
 | |
6T4V
 
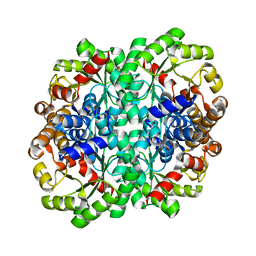 | |
6G1O
 
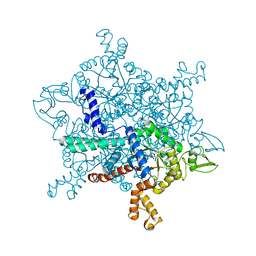 | |
6QXL
 
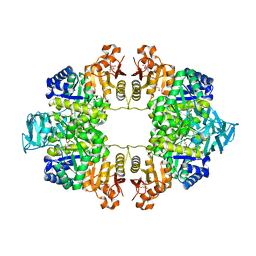 | | Crystal Structure of Pyruvate Kinase II (PykA) from Pseudomonas aeruginosa in complex with sodium malonate, magnesium and glucose-6-phosphate | | Descriptor: | 6-O-phosphono-alpha-D-glucopyranose, GLYCEROL, MAGNESIUM ION, ... | | Authors: | Abdelhamid, Y, Brear, P, Welch, M. | | Deposit date: | 2019-03-07 | | Release date: | 2019-09-11 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.43 Å) | | Cite: | Evolutionary plasticity in the allosteric regulator-binding site of pyruvate kinase isoform PykA fromPseudomonas aeruginosa.
J.Biol.Chem., 294, 2019
|
|
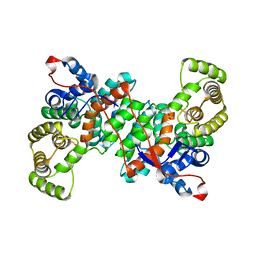
6S87
 
 | |
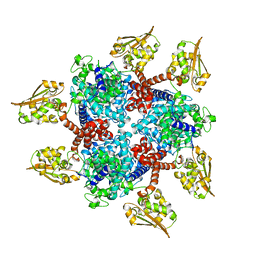
6S62
 
 | |
2GFS
 
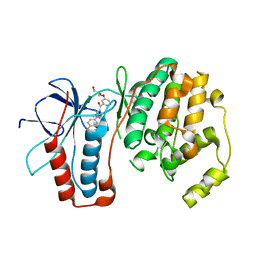 | | P38 Kinase Crystal Structure in complex with RO3201195 | | Descriptor: | Mitogen-Activated Protein Kinase 14, [5-AMINO-1-(4-FLUOROPHENYL)-1H-PYRAZOL-4-YL](3-{[(2R)-2,3-DIHYDROXYPROPYL]OXY}PHENYL)METHANONE | | Authors: | Harris, S.F, Bertrand, J, Villasenor, A. | | Deposit date: | 2006-03-23 | | Release date: | 2006-04-18 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.752 Å) | | Cite: | Discovery of S-[5-Amino-1-(4-fluorophenyl)-1H-pyrazol-4-yl]-[3-(2,3-dihydroxypropoxy)phenyl]-methanone (RO3201195), and Orally Bioavailable and Highly Selective Inhibitor of p38 Map Kinase
J.Med.Chem., 49, 2006
|
|
3D4Q
 
 | | Pyrazole-based inhibitors of B-Raf kinase | | Descriptor: | (1E)-5-(1-piperidin-4-yl-3-pyridin-4-yl-1H-pyrazol-4-yl)-2,3-dihydro-1H-inden-1-one oxime, B-Raf proto-oncogene serine/threonine-protein kinase | | Authors: | Morales, T, Vigers, G.P.A, Brandhuber, B.J. | | Deposit date: | 2008-05-14 | | Release date: | 2008-08-19 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Potent and selective pyrazole-based inhibitors of B-Raf kinase.
Bioorg.Med.Chem.Lett., 18, 2008
|
|
3PSB
 
 | | Furo[2,3-c]pyridine-based Indanone Oximes as Potent and Selective B-Raf Inhibitors | | Descriptor: | B-RAF PROTO-ONCOGENE SERINE/THREONINE-PROTEIN KINASE, ethyl 3-{[1-(hydroxyamino)-2H-inden-5-yl]amino}thieno[2,3-c]pyridine-2-carboxylate | | Authors: | Morales, T, Vigers, G.P.A, Brandhuber, B.J. | | Deposit date: | 2010-12-01 | | Release date: | 2011-01-19 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (3.4 Å) | | Cite: | The Discovery of furo[2,3-c]pyridine-based indanone oximes as potent and selective B-Raf inhibitors.
Bioorg.Med.Chem.Lett., 21, 2011
|
|
3PSD
 
 | |
1OZ1
 
 | | P38 MITOGEN-ACTIVATED KINASE IN COMPLEX WITH 4-AZAINDOLE INHIBITOR | | Descriptor: | 3-(4-FLUOROPHENYL)-2-PYRIDIN-4-YL-1H-PYRROLO[3,2-B]PYRIDIN-1-OL, Mitogen-activated protein kinase 14 | | Authors: | Lovejoy, B, Villasenor, A, Browner, M, Dunten, P. | | Deposit date: | 2003-04-07 | | Release date: | 2003-09-23 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Design and synthesis of 4-azaindoles as inhibitors of p38 MAP kinase.
J.Med.Chem., 46, 2003
|
|
