5YA6
 
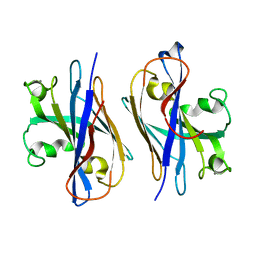 | |
5Z1L
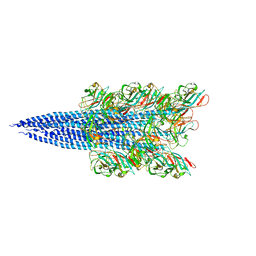 
 | | Cryo-EM structure of Methanoccus maripaludis archaellum | | Descriptor: | Flagellin | | Authors: | Meshcheryakov, V.A, Shibata, S, Schreiber, M.T, Villar-Briones, A, Jarrell, K.F, Aizawa, S, Wolf, M. | | Deposit date: | 2017-12-26 | | Release date: | 2019-02-13 | | Last modified: | 2024-03-27 | | Method: | ELECTRON MICROSCOPY (4 Å) | | Cite: | High-resolution archaellum structure reveals a conserved metal-binding site.
Embo Rep., 20, 2019
|
|
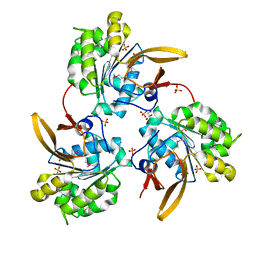
4WIA
 
 | |
3B0Z
 
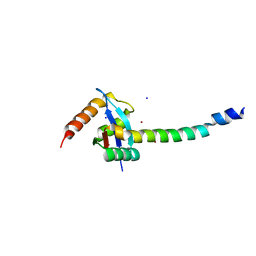 | |
3B1S
 
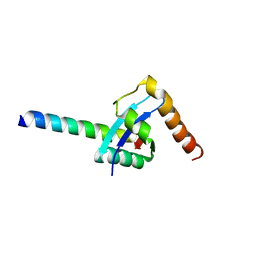 | |
3AZD
 
 | | Crystal structure of tropomyosin N-terminal fragment at 0.98A resolution | | Descriptor: | short alpha-tropomyosin,transcription factor GCN4 | | Authors: | Meshcheryakov, V.A, Krieger, I, Kostyukova, A.S, Samatey, F.A. | | Deposit date: | 2011-05-23 | | Release date: | 2011-10-19 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (0.98 Å) | | Cite: | Structure of a tropomyosin N-terminal fragment at 0.98 A resolution
Acta Crystallogr.,Sect.D, 67, 2011
|
|
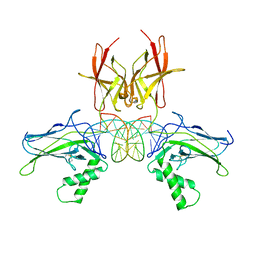
7VUP
 
 | |
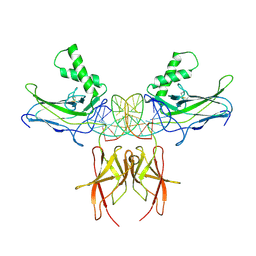
7VUQ
 
 | |
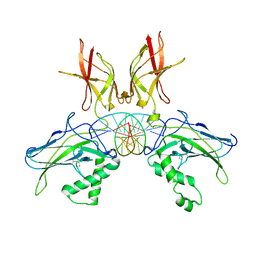
7W7L
 
 | |
7CLI
 
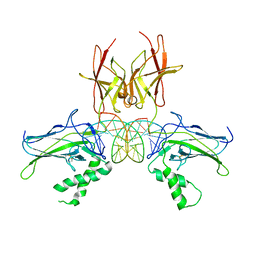 | | Structure of NF-kB p52 homodimer bound to P-Selectin kB DNA fragment | | Descriptor: | DNA (5'-D(*CP*AP*AP*GP*GP*GP*GP*TP*CP*AP*CP*CP*CP*CP*CP*TP*TP*C)-3'), DNA (5'-D(*GP*AP*AP*GP*GP*GP*GP*GP*TP*GP*AP*CP*CP*CP*CP*TP*TP*G)-3'), Nuclear factor NF-kappa-B p52 subunit | | Authors: | Meshcheryakov, V.A, Wang, V.Y.-F. | | Deposit date: | 2020-07-21 | | Release date: | 2021-07-21 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structures of NF-kappa B p52 homodimer-DNA complexes rationalize binding mechanisms and transcription activation.
Elife, 12, 2023
|
|
1FEU
 
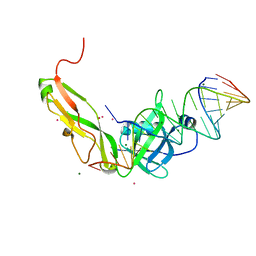 | | CRYSTAL STRUCTURE OF RIBOSOMAL PROTEIN TL5, ONE OF THE CTC FAMILY PROTEINS, COMPLEXED WITH A FRAGMENT OF 5S RRNA. | | Descriptor: | 19 NT FRAGMENT OF 5S RRNA, 21 NT FRAGMENT OF 5S RRNA, 50S RIBOSOMAL PROTEIN L25, ... | | Authors: | Fedorov, R.V, Meshcheryakov, V.A, Gongadze, G.M, Fomenkova, N.P, Nevskaya, N.A, Selmer, M, Laurberg, M, Kristensen, O, Al-Karadaghi, S, Liljas, A, Garber, M.B, Nikonov, S.V. | | Deposit date: | 2000-07-23 | | Release date: | 2001-06-25 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure of ribosomal protein TL5 complexed with RNA provides new insights into the CTC family of stress proteins.
Acta Crystallogr.,Sect.D, 57, 2001
|
|
3VKI
 
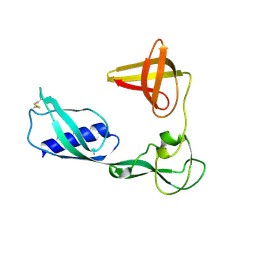 | |
3VJP
 
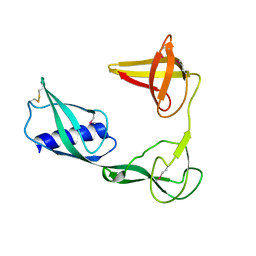 | |
3TEE
 
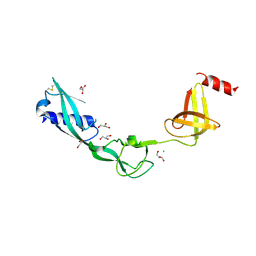 | | Crystal Structure of Salmonella FlgA in open form | | Descriptor: | CHLORIDE ION, Flagella basal body P-ring formation protein flgA, GLYCEROL | | Authors: | Matsunami, H, Samatey, F.A, Namba, K. | | Deposit date: | 2011-08-12 | | Release date: | 2012-08-15 | | Last modified: | 2016-07-06 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structural flexibility of the periplasmic protein, FlgA, regulates flagellar P-ring assembly in Salmonella enterica
Sci Rep, 6, 2016
|
|
