6A24
 
 | | The crystal structure of Mandelate oxidase with 3-fluoropyruvate | | Descriptor: | 4-hydroxymandelate oxidase, FLAVIN MONONUCLEOTIDE, FLUORIDE ION, ... | | Authors: | Li, T.L, Lin, K.H. | | Deposit date: | 2018-06-08 | | Release date: | 2019-06-19 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.39 Å) | | Cite: | Structural and chemical trapping of flavin-oxide intermediates reveals substrate-directed reaction multiplicity.
Protein Sci., 29, 2020
|
|
6A0D
 
 | |
6A0O
 
 | | Mandelate oxidase mutant-Y128F with benzaldehyde | | Descriptor: | 4-hydroxymandelate oxidase, FLAVIN MONONUCLEOTIDE, benzaldehyde | | Authors: | Li, T.L, Lin, K.H. | | Deposit date: | 2018-06-06 | | Release date: | 2019-06-19 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Biochemical and structural explorations of alpha-hydroxyacid oxidases reveal a four-electron oxidative decarboxylation reaction.
Acta Crystallogr D Struct Biol, 75, 2019
|
|
6A0Y
 
 | | Mandelate oxidase mutant-Y128F with Benzoic acid | | Descriptor: | 4-hydroxymandelate oxidase, BENZOIC ACID, FLAVIN MONONUCLEOTIDE | | Authors: | Li, T.L, Lin, K.H. | | Deposit date: | 2018-06-06 | | Release date: | 2019-06-19 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Biochemical and structural explorations of alpha-hydroxyacid oxidases reveal a four-electron oxidative decarboxylation reaction.
Acta Crystallogr D Struct Biol, 75, 2019
|
|
6A1H
 
 | | Mandelate oxidase mutant-Y128F with 5-deazariboflavin mononucleotide | | Descriptor: | 1-deoxy-1-(7,8-dimethyl-2,4-dioxo-3,4-dihydropyrimido[4,5-b]quinolin-10(2H)-yl)-5-O-phosphono-D-ribitol, 4-hydroxymandelate oxidase | | Authors: | Li, T.L, Lin, K.H. | | Deposit date: | 2018-06-07 | | Release date: | 2019-06-19 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.36 Å) | | Cite: | The flavin mononucleotide cofactor in alpha-hydroxyacid oxidases exerts its electrophilic/nucleophilic duality in control of the substrate-oxidation level.
Acta Crystallogr D Struct Biol, 75, 2019
|
|
6A0M
 
 | | Mandelate oxidase mutant-Y128F with 2-Phenylacetic acid | | Descriptor: | 2-PHENYLACETIC ACID, 4-hydroxymandelate oxidase, FLAVIN MONONUCLEOTIDE | | Authors: | Li, T.L, Lin, K.H. | | Deposit date: | 2018-06-05 | | Release date: | 2019-06-19 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Biochemical and structural explorations of alpha-hydroxyacid oxidases reveal a four-electron oxidative decarboxylation reaction.
Acta Crystallogr D Struct Biol, 75, 2019
|
|
6A1R
 
 | | Mandelate oxidase mutant-Y128F with the N5-phenylacetyl-FMN adduct | | Descriptor: | 1-deoxy-1-[7,8-dimethyl-2,4-dioxo-5-(phenylacetyl)-1,3,4,5-tetrahydrobenzo[g]pteridin-10(2H)-yl]-5-O-phosphono-D-ribitol, 4-hydroxymandelate oxidase, MAGNESIUM ION | | Authors: | Li, T.L, Lin, K.H. | | Deposit date: | 2018-06-08 | | Release date: | 2019-06-19 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.652 Å) | | Cite: | The flavin mononucleotide cofactor in alpha-hydroxyacid oxidases exerts its electrophilic/nucleophilic duality in control of the substrate-oxidation level.
Acta Crystallogr D Struct Biol, 75, 2019
|
|
6A1L
 
 | | Mandelate oxidase mutant-Y128F with 5-deazariboflavin mononucleotide and benzoic acid | | Descriptor: | 1-deoxy-1-(7,8-dimethyl-2,4-dioxo-3,4-dihydropyrimido[4,5-b]quinolin-10(2H)-yl)-5-O-phosphono-D-ribitol, 4-hydroxymandelate oxidase, BENZOIC ACID, ... | | Authors: | Li, T.L, Lin, K.H. | | Deposit date: | 2018-06-07 | | Release date: | 2019-06-19 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | The flavin mononucleotide cofactor in alpha-hydroxyacid oxidases exerts its electrophilic/nucleophilic duality in control of the substrate-oxidation level.
Acta Crystallogr D Struct Biol, 75, 2019
|
|
6A3D
 | | The crystal structure of Mandelate oxidase Y128F with 4-Br-2-hydroxy-methylphenylacetate | | Descriptor: | 1-deoxy-1-[(4aS)-4a-[(methoxycarbonyl)peroxy]-7,8-dimethyl-2,4-dioxo-3,4,4a,5-tetrahydrobenzo[g]pteridin-10(2H)-yl]-5-O-phosphono-D-ribitol, 4-hydroxymandelate oxidase | | Authors: | Li, T.L, Lin, K.H. | | Deposit date: | 2018-06-15 | | Release date: | 2019-06-19 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.923 Å) | | Cite: | Structural and chemical trapping of flavin-oxide intermediates reveals substrate-directed reaction multiplicity.
Protein Sci., 29, 2020
|
|
6A3T
 
 | | The crystal structure of Mandelate oxidase R163L with 2-hydroxy-phenylacetamide | | Descriptor: | 1-{5-[(1S)-2-amino-1-hydroxy-2-oxo-1-phenylethyl]-7,8-dimethyl-2,4-dioxo-1,2,3,4-tetrahydrobenzo[g]pteridine-5,10-diium-10-yl}-1-deoxy-5-O-phosphono-D-ribitol, 4-hydroxymandelate oxidase | | Authors: | Li, T.L, Lin, K.H. | | Deposit date: | 2018-06-16 | | Release date: | 2019-06-19 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.511 Å) | | Cite: | The flavin mononucleotide cofactor in alpha-hydroxyacid oxidases exerts its electrophilic/nucleophilic duality in control of the substrate-oxidation level.
Acta Crystallogr D Struct Biol, 75, 2019
|
|
7F0A
 
 | |
7F0B
 
 | | Crystal structure of capreomycin phosphotransferase in complex with ATP | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, Capreomycin phosphotransferase | | Authors: | Chang, C.Y, Pan, Y.C, Wang, Y.L, Toh, S.I. | | Deposit date: | 2021-06-03 | | Release date: | 2022-05-11 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.14 Å) | | Cite: | Dual-Mechanism Confers Self-Resistance to the Antituberculosis Antibiotic Capreomycin.
Acs Chem.Biol., 17, 2022
|
|
7F0F
 
 | | Crystal structure of capreomycin phosphotransferase in complex with CMN IIB | | Descriptor: | Capreomycin phosphotransferase, DPP-ALA-DPP-UAL-MYN-KBE | | Authors: | Chang, C.Y, Pan, Y.C, Wang, Y.L, Toh, S.I. | | Deposit date: | 2021-06-03 | | Release date: | 2022-05-11 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Dual-Mechanism Confers Self-Resistance to the Antituberculosis Antibiotic Capreomycin.
Acs Chem.Biol., 17, 2022
|
|
7F0C
 
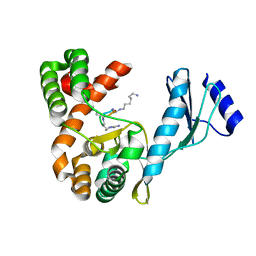 | | Crystal structure of capreomycin phosphotransferase in complex with CMN IIA | | Descriptor: | Capreomycin phosphotransferase, DPP-SER-DPP-UAL-MYN-KBE | | Authors: | Chang, C.Y, Pan, Y.C, Wang, Y.L, Toh, S.I. | | Deposit date: | 2021-06-03 | | Release date: | 2022-05-11 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.07 Å) | | Cite: | Dual-Mechanism Confers Self-Resistance to the Antituberculosis Antibiotic Capreomycin.
Acs Chem.Biol., 17, 2022
|
|
7EUE
 
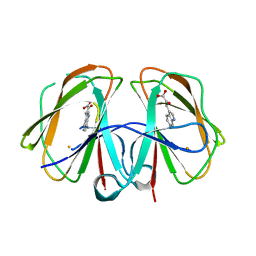 | |
7F6X
 
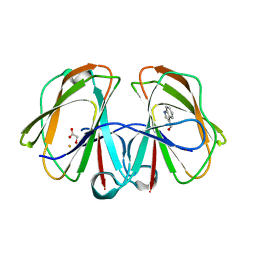 | | Structural and mechanistic studies of a novel non-heme iron epimerase/lyase and its utilization in chemoselective synthesis. | | Descriptor: | 1H-INDOLE-3-CARBALDEHYDE, Cupin domain-containing protein, FE (III) ION, ... | | Authors: | Li, T.L, Li, Y.S, Chen, M.H. | | Deposit date: | 2021-06-26 | | Release date: | 2022-03-16 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.16 Å) | | Cite: | Structural and Mechanistic Bases for StnK3 and Its Mutant-MediatedLewis-Acid-Dependent Epimerization and Retro-Aldol Reactions.
Acs Catalysis, 12, 2022
|
|
7EUZ
 
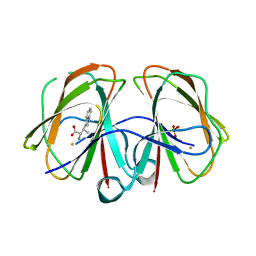 | | Structural and mechanistic studies of a novel non-heme iron epimerase/lyase and its utilization in chemoselective synthesis. | | Descriptor: | (2S,3R)-2-azanyl-3-(1H-indol-3-yl)butanoic acid, Cupin domain-containing protein, FE (III) ION, ... | | Authors: | Li, T.L, Li, Y.S, Chen, M.H. | | Deposit date: | 2021-05-19 | | Release date: | 2022-03-16 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.91028166 Å) | | Cite: | Structural and Mechanistic Bases for StnK3 and Its Mutant-Mediated Lewis-Acid-Dependent Epimerization and Retro-Aldol Reactions.
Acs Catalysis, 12, 2022
|
|
7EUP
 
 | | Structural and mechanistic studies of a novel non-heme iron epimerase/lyase and its utilization in chemoselective synthesis. | | Descriptor: | (2S,3R)-2-azanyl-3-phenyl-butanoic acid, Cupin domain-containing protein, FE (III) ION | | Authors: | Li, T.L, Li, Y.S, Chen, M.H. | | Deposit date: | 2021-05-18 | | Release date: | 2022-03-16 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.11043429 Å) | | Cite: | Structural and Mechanistic Bases for StnK3 and Its Mutant-Mediated Lewis-Acid-Dependent Epimerization and Retro-Aldol Reactions.
Acs Catalysis, 12, 2022
|
|
7EQK
 
 | | Structural and mechanistic studies of a novel non-heme iron epimerase/lyase and its utilization in chemoselective synthesis. | | Descriptor: | (E)-3-(1H-indol-3-yl)-2-oxidanyl-but-2-enoic acid, 1-(1~{H}-indol-3-yl)ethanone, Cupin domain-containing protein, ... | | Authors: | Li, T.L, Li, Y.S, Chen, M.H. | | Deposit date: | 2021-05-03 | | Release date: | 2022-03-16 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.04001021 Å) | | Cite: | Structural and Mechanistic Bases for StnK3 and Its Mutant-Mediated Lewis-Acid-Dependent Epimerization and Retro-Aldol Reactions.
Acs Catalysis, 12, 2022
|
|
7EU6
 
 | |
7V4M
 
 | |
7V4O
 
 | | Unique non-heme hydroxylase in biosynthesis of nucleoside antibiotic pathway uncover mechanism of reaction | | Descriptor: | 3,6,9,12,15-PENTAOXAHEPTADECANE, Beta-hydroxylase | | Authors: | Li, T.L, Saeid, M.Z. | | Deposit date: | 2021-08-13 | | Release date: | 2022-02-23 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | beta-Hydroxylation of alpha-amino-beta-hydroxylbutanoyl-glycyluridine catalyzed by a nonheme hydroxylase ensures the maturation of caprazamycin
Commun Chem, 5, 2022
|
|
7V4N
 
 | |
7V4P
 | |
7VYY
 
 | | The crystal structure of Non-hydrolyzing UDPGlcNAc 2-epimerase | | Descriptor: | Putative UDP-N-acetylglucosamine 2-epimerase, SODIUM ION | | Authors: | Li, T.L, Rattinam, R. | | Deposit date: | 2021-11-15 | | Release date: | 2022-11-16 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.443 Å) | | Cite: | KasQ an Epimerase Primes the Biosynthesis of Aminoglycoside Antibiotic Kasugamycin and KasF/H Acetyltransferases Inactivate Its Activity.
Biomedicines, 10, 2022
|
|
