4YHE
 
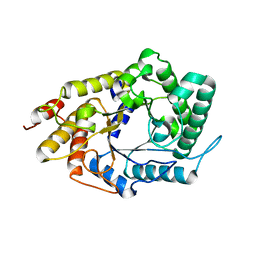 | | NATIVE BACTEROIDETES-AFFILIATED GH5 CELLULASE LINKED WITH A POLYSACCHARIDE UTILIZATION LOCUS | | Descriptor: | GH5 | | Authors: | Naas, A.E, MacKenzie, A.K, Dalhus, B, Eijsink, V.G.H, Pope, P.B. | | Deposit date: | 2015-02-27 | | Release date: | 2015-05-20 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Structural Features of a Bacteroidetes-Affiliated Cellulase Linked with a Polysaccharide Utilization Locus.
Sci Rep, 5, 2015
|
|
4OY7
 
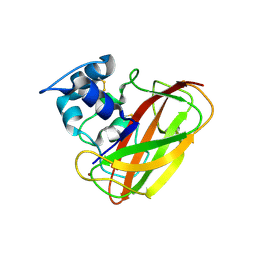 | | Structure of cellulose active LPMO CelS2 (ScLPMO10C) in complex with Copper. | | Descriptor: | CALCIUM ION, COPPER (II) ION, Putative secreted cellulose binding protein | | Authors: | Forsberg, Z, Mackenzie, A.K, Sorlie, M, Rohr, A.K, Helland, R, Arvai, A.S, Vaaje-Kolstad, G, Eijsink, V.G.H. | | Deposit date: | 2014-02-11 | | Release date: | 2014-05-28 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Structural and functional characterization of a conserved pair of bacterial cellulose-oxidizing lytic polysaccharide monooxygenases.
Proc.Natl.Acad.Sci.USA, 111, 2014
|
|
4YHG
 
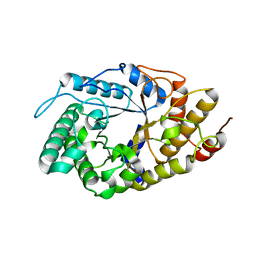 | | NATIVE BACTEROIDETES-AFFILIATED GH5 CELLULASE LINKED WITH A POLYSACCHARIDE UTILIZATION LOCUS | | Descriptor: | GH5, beta-D-glucopyranose-(1-4)-beta-D-glucopyranose-(1-4)-beta-D-glucopyranose | | Authors: | Naas, A.E, MacKenzie, A.K, Dalhus, B, Eijsink, V.G.H, Pope, P.B. | | Deposit date: | 2015-02-27 | | Release date: | 2015-07-15 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural Features of a Bacteroidetes-Affiliated Cellulase Linked with a Polysaccharide Utilization Locus.
Sci Rep, 5, 2015
|
|
6TWE
 
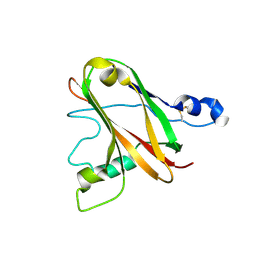 | |
8C10
 
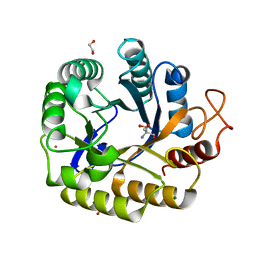 | | Biochemical and structural characterisation of an alkaline family GH5 cellulase from a shipworm symbiont | | Descriptor: | 1,2-ETHANEDIOL, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, GH5 Cellulase, ... | | Authors: | Leiros, I, Vaaje-Kolstad, G. | | Deposit date: | 2022-12-19 | | Release date: | 2023-04-19 | | Last modified: | 2024-06-19 | | Method: | X-RAY DIFFRACTION (1 Å) | | Cite: | Biochemical and structural characterisation of a family GH5 cellulase from endosymbiont of shipworm P. megotara.
Biotechnol Biofuels Bioprod, 16, 2023
|
|
2Y8U
 
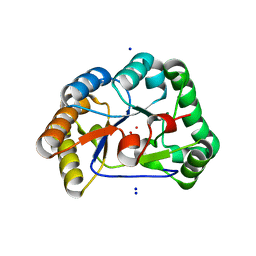 | | A. nidulans chitin deacetylase | | Descriptor: | CHITIN DEACETYLASE, CHLORIDE ION, COBALT (II) ION, ... | | Authors: | Penman, G, Gay, L.M, van Aalten, D.M.F. | | Deposit date: | 2011-02-10 | | Release date: | 2012-02-22 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.99 Å) | | Cite: | Structure and function of a broad-specificity chitin deacetylase from Aspergillus nidulans FGSC A4.
Sci Rep, 7, 2017
|
|
9FO1
 
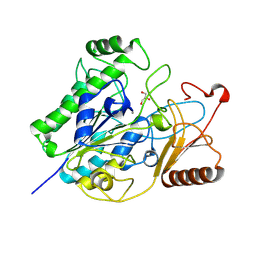 | | Family S1_16 sulfatase from metagenomic dataset AMOR | | Descriptor: | CALCIUM ION, GLYCEROL, k-carrageenan sulfatase S1_16 | | Authors: | Rhein-Knudsen, N, Horn, S.J, Jeudy, A, Czjzek, M. | | Deposit date: | 2024-06-11 | | Release date: | 2025-01-22 | | Last modified: | 2025-02-05 | | Method: | X-RAY DIFFRACTION (3.11 Å) | | Cite: | Identification and Characterization of a New Thermophilic kappa-Carrageenan Sulfatase.
J.Agric.Food Chem., 73, 2025
|
|
1O6I
 
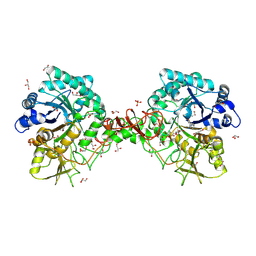 | | Chitinase B from Serratia marcescens complexed with the catalytic intermediate mimic cyclic dipeptide CI4. | | Descriptor: | Chitinase, GLYCEROL, SULFATE ION, ... | | Authors: | Houston, D.R, Eggleston, I, Synstad, B, Eijsink, V.G.H, van Aalten, D.M.F. | | Deposit date: | 2002-10-03 | | Release date: | 2003-03-30 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | The cyclic dipeptide CI-4 [cyclo-(l-Arg-d-Pro)] inhibits family 18 chitinases by structural mimicry of a reaction intermediate.
Biochem. J., 368, 2002
|
|
2VYO
 
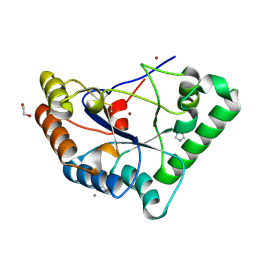 | | Chitin deacetylase family member from Encephalitozoon cuniculi | | Descriptor: | 1,2-ETHANEDIOL, ACETATE ION, CALCIUM ION, ... | | Authors: | Urch, J.E, Hurtado-Guerrero, R, Texier, C, Van Aalten, D.M.F. | | Deposit date: | 2008-07-25 | | Release date: | 2008-08-12 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Structural and Functional Characterization of a Putative Polysaccharide Deacetylase of the Human Parasite Encephalitozoon Cuniculi.
Protein Sci., 18, 2009
|
|
5VG1
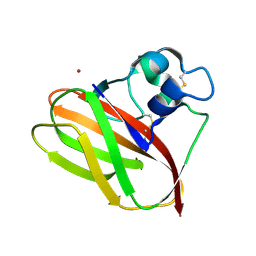 
 | |
5VG0
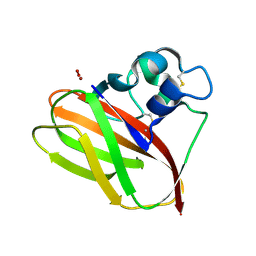 
 | |
5LW4
 
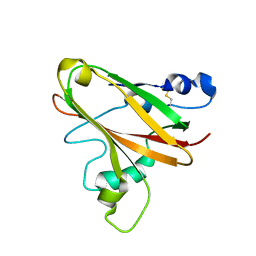 | |
5WH8
 
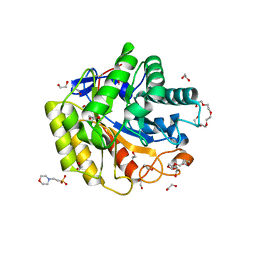 | | Cellulase Cel5C_n | | Descriptor: | 1,2-ETHANEDIOL, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, PENTAETHYLENE GLYCOL, ... | | Authors: | Koropatkin, N.M, Pope, P.B, Naas, A.E. | | Deposit date: | 2017-07-15 | | Release date: | 2018-03-14 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (1.57 Å) | | Cite: | "Candidatus Paraporphyromonas polyenzymogenes" encodes multi-modular cellulases linked to the type IX secretion system.
Microbiome, 6, 2018
|
|
5LFZ
 
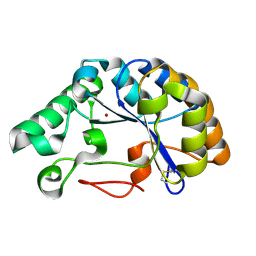 | | T48 deacetylase | | Descriptor: | ArCE4A, NICKEL (II) ION | | Authors: | Rothweiler, U. | | Deposit date: | 2016-07-05 | | Release date: | 2017-08-02 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (1.561 Å) | | Cite: | Structure and function of a CE4 deacetylase isolated from a marine environment.
PLoS ONE, 12, 2017
|
|
5LGC
 
 | | T48 deacetylase with substrate | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ArCE4A | | Authors: | Rothweiler, U. | | Deposit date: | 2016-07-06 | | Release date: | 2017-08-02 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.09 Å) | | Cite: | Structure and function of a CE4 deacetylase isolated from a marine environment.
PLoS ONE, 12, 2017
|
|
