2I6U
 
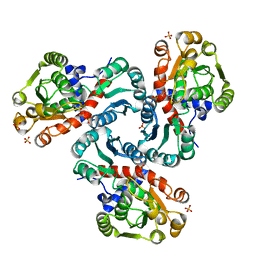 | | Crystal Structure of Ornithine Carbamoyltransferase complexed with Carbamoyl Phosphate and L-Norvaline from Mycobacterium tuberculosis (Rv1656) at 2.2 A | | Descriptor: | NORVALINE, Ornithine carbamoyltransferase, PHOSPHORIC ACID MONO(FORMAMIDE)ESTER, ... | | Authors: | Sankaranarayanan, R, Moradian, F, Cherney, L.T, Garen, C, Cherney, M.M, James, M.N.G, TB Structural Genomics Consortium (TBSGC) | | Deposit date: | 2006-08-29 | | Release date: | 2007-06-26 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The crystal structures of ornithine carbamoyltransferase from Mycobacterium tuberculosis and its ternary complex with carbamoyl phosphate and L-norvaline reveal the enzyme's catalytic mechanism
J.Mol.Biol., 375, 2008
|
|
2GJF
 
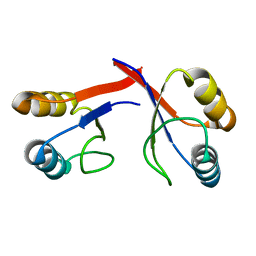 | |
2PLB
 
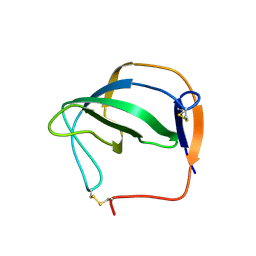
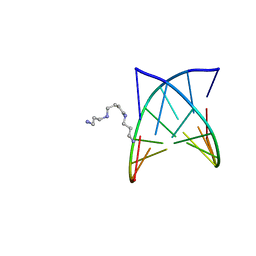 | | D(GTATACC) under hydrostatic pressure of 1.39 GPa | | Descriptor: | 5'-D(*DGP*DGP*DTP*DAP*DTP*DAP*DCP*DC)-3', SPERMINE | | Authors: | Prange, T, Girard, E, Kahn, R, Fourme, R. | | Deposit date: | 2007-04-19 | | Release date: | 2007-07-24 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Adaptation of the base-paired double-helix molecular architecture to extreme pressure.
Nucleic Acids Res., 35, 2007
|
|
2PKV
 
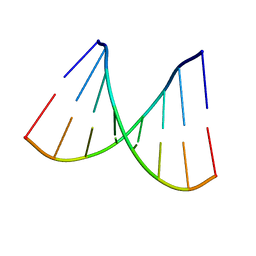 | | D-(GGTATACC) ambient pressure | | Descriptor: | 5'-D(*GP*GP*TP*AP*TP*AP*CP*C)-3' | | Authors: | Girard, E, Prange, T, Kahn, R, Fourme, R. | | Deposit date: | 2007-04-18 | | Release date: | 2007-07-24 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Adaptation of the base-paired double-helix molecular architecture to extreme pressure.
Nucleic Acids Res., 35, 2007
|
|
2PL4
 
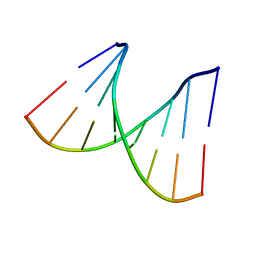 | |
2PL8
 
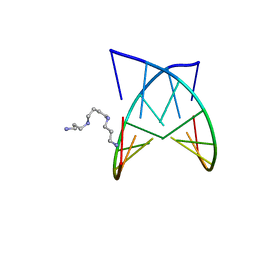 | | D(GTATACC) under hydrostatic pressure of 1.04 GPa | | Descriptor: | 5'-D(*DGP*DGP*DTP*DAP*DTP*DAP*DCP*DC)-3', SPERMINE | | Authors: | Prange, T, Girard, E, Fourme, R, Kahn, R. | | Deposit date: | 2007-04-19 | | Release date: | 2007-07-24 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Adaptation of the base-paired double-helix molecular architecture to extreme pressure.
Nucleic Acids Res., 35, 2007
|
|
11BG
 
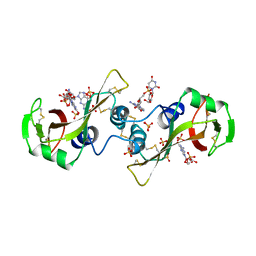 | | A POTENTIAL ALLOSTERIC SUBSITE GENERATED BY DOMAIN SWAPPING IN BOVINE SEMINAL RIBONUCLEASE | | Descriptor: | PROTEIN (BOVINE SEMINAL RIBONUCLEASE), SULFATE ION, URIDYLYL-2'-5'-PHOSPHO-GUANOSINE | | Authors: | Vitagliano, L, Adinolfi, S, Sica, F, Merlino, A, Zagari, A, Mazzarella, L. | | Deposit date: | 1999-03-11 | | Release date: | 1999-11-05 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | A potential allosteric subsite generated by domain swapping in bovine seminal ribonuclease.
J.Mol.Biol., 293, 1999
|
|
1R3H
 
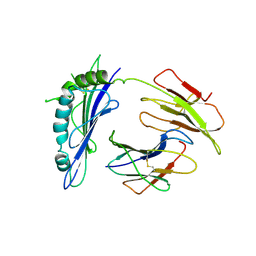 | | Crystal Structure of T10 | | Descriptor: | Beta-2-microglobulin, MHC H2-TL-T10-129 | | Authors: | Rudolph, M.G, Wilson, I.A. | | Deposit date: | 2003-10-02 | | Release date: | 2004-03-30 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Combined pseudo-merohedral twinning, non-crystallographic symmetry and pseudo-translation in a monoclinic crystal form of the gammadelta T-cell ligand T10.
Acta Crystallogr.,Sect.D, 60, 2004
|
|
1HJD
 
 | |
2BTF
 
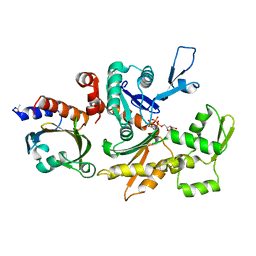 | | THE STRUCTURE OF CRYSTALLINE PROFILIN-BETA-ACTIN | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, BETA-ACTIN, PROFILIN, ... | | Authors: | Schutt, C.E, Myslik, J.C, Rozycki, M.D, Goonesekere, N.C.W. | | Deposit date: | 1994-01-18 | | Release date: | 1994-06-22 | | Last modified: | 2017-11-29 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | The structure of crystalline profilin-beta-actin.
Nature, 365, 1993
|
|
1H87
 
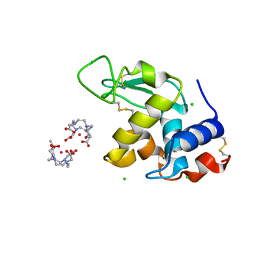 | | Gadolinium derivative of tetragonal Hen Egg-White Lysozyme at 1.7 A resolution | | Descriptor: | 10-((2R)-2-HYDROXYPROPYL)-1,4,7,10-TETRAAZACYCLODODECANE 1,4,7-TRIACETIC ACID, CHLORIDE ION, GADOLINIUM ATOM, ... | | Authors: | Girard, E, Chantalat, L, Vicat, J, Kahn, R. | | Deposit date: | 2001-01-25 | | Release date: | 2002-01-15 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.72 Å) | | Cite: | Gd-Hp-Do3A, a Complex to Obtain High-Phasing-Power Heavy Atom Derivatives for Sad and MAD Experiments. Results with Tetragonal Hen Egg-White Lysozyme
Acta Crystallogr.,Sect.D, 58, 2001
|
|
1F89
 
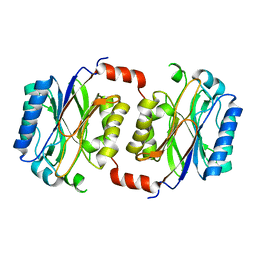 | | Crystal structure of Saccharomyces cerevisiae Nit3, a member of branch 10 of the nitrilase superfamily | | Descriptor: | 32.5 KDA PROTEIN YLR351C | | Authors: | Kumaran, D, Eswaramoorthy, S, Studier, F.W, Swaminathan, S, Burley, S.K, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2000-06-29 | | Release date: | 2001-10-04 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of a putative CN hydrolase from yeast
Proteins, 52, 2003
|
|
7JPU
 
 | |
7JPT
 
 | | Structure of an endocytic receptor | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Lymphocyte antigen 75, beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | | Authors: | Gully, B.S, Rossjohn, J, Berry, R. | | Deposit date: | 2020-08-09 | | Release date: | 2020-12-09 | | Last modified: | 2021-07-14 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | The cryo-EM structure of the endocytic receptor DEC-205.
J.Biol.Chem., 296, 2020
|
|
