4K40
 
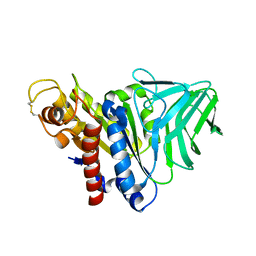 | | Peptidoglycan O-acetylesterase in action, 0 min | | Descriptor: | GDSL-like Lipase/Acylhydrolase family protein | | Authors: | Williams, A.H, Gompert Boneca, I. | | Deposit date: | 2013-04-11 | | Release date: | 2014-09-03 | | Last modified: | 2017-06-21 | | Method: | X-RAY DIFFRACTION (2.634 Å) | | Cite: | Visualization of a substrate-induced productive conformation of the catalytic triad of the Neisseria meningitidis peptidoglycan O-acetylesterase reveals mechanistic conservation in SGNH esterase family members.
Acta Crystallogr.,Sect.D, 70, 2014
|
|
6NXD
 
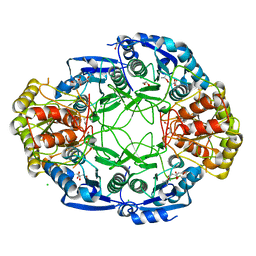 | |
1QD7
 
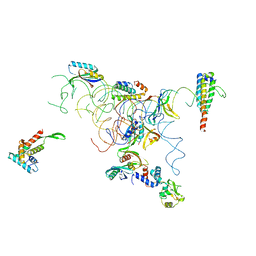 | | PARTIAL MODEL FOR 30S RIBOSOMAL SUBUNIT | | Descriptor: | CENTRAL FRAGMENT OF 16 S RNA, END FRAGMENT OF 16 S RNA, S15 RIBOSOMAL PROTEIN, ... | | Authors: | Clemons Jr, W.M, May, J.L.C, Wimberly, B.T, McCutcheon, J.P, Capel, M.S, Ramakrishnan, V. | | Deposit date: | 1999-07-09 | | Release date: | 1999-08-31 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (5.5 Å) | | Cite: | Structure of a bacterial 30S ribosomal subunit at 5.5 A resolution.
Nature, 400, 1999
|
|
1BM9
 
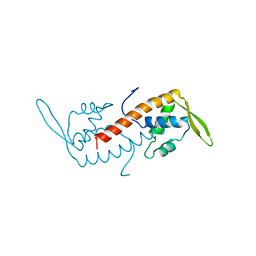 | |
1CQU
 
 | |
6NXC
 | |
2EWS
 
 | |
2AQ7
 
 | | Structure-activity relationships at the 5-posiiton of thiolactomycin: an intact 5(R)-isoprene unit is required for activity against the condensing enzymes from Mycobacterium tuberculosis and Escherichia coli | | Descriptor: | (5R)-4-HYDROXY-3,5-DIMETHYL-5-[(1E,3E)-2-METHYLPENTA-1,3-DIENYL]THIOPHEN-2(5H)-ONE, 3-oxoacyl-[acyl-carrier-protein] synthase I | | Authors: | Kim, P, Zhang, Y.M, Shenoy, G, Nguyen, Q.A, Boshoff, H.I, Manjunatha, U.H, Goodwin, M.B, Lonsdale, J, Price, A.C, Miller, D.J. | | Deposit date: | 2005-08-17 | | Release date: | 2006-01-17 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure-Activity Relationships at the 5-Position of Thiolactomycin: An Intact (5R)-Isoprene Unit Is Required for Activity against the Condensing Enzymes from Mycobacterium tuberculosis and Escherichia coli
J.Med.Chem., 49, 2006
|
|
2AQB
 
 | | Structure-activity relationships at the 5-position of thiolactomycin: an intact 5(R)-isoprene unit is required for activity against the condensing enzymes from Mycobacterium tuberculosis and Escherchia coli | | Descriptor: | (5R)-5-[(1E)-BUTA-1,3-DIENYL]-4-HYDROXY-3,5-DIMETHYLTHIOPHEN-2(5H)-ONE, 3-oxoacyl-[acyl-carrier-protein] synthase I | | Authors: | Kim, P, Zhang, Y.M, Shenoy, G, Nguyen, Q.A, Boshoff, H.I, Manjunatha, U.H, Goodwin, M.B, Lonsdale, J, Price, A.C, Miller, D.J. | | Deposit date: | 2005-08-17 | | Release date: | 2006-01-17 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.19 Å) | | Cite: | Structure-Activity Relationships at the 5-Position of Thiolactomycin: An Intact (5R)-Isoprene Unit Is Required for Activity against the Condensing Enzymes from Mycobacterium tuberculosis and Escherichia coli
J.Med.Chem., 49, 2006
|
|
2M03
 
 | |
6NX9
 
 | |
6NXB
 
 | | ECAII IN COMPLEX WITH CITRATE AT PH 7 | | Descriptor: | CITRIC ACID, GLYCEROL, L-asparaginase 2 | | Authors: | Lubkowski, J, Wlodawer, A. | | Deposit date: | 2019-02-08 | | Release date: | 2019-08-07 | | Last modified: | 2019-11-20 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Opportunistic complexes of E. coli L-asparaginases with citrate anions.
Sci Rep, 9, 2019
|
|
6NX6
 
 | |
6NXA
 
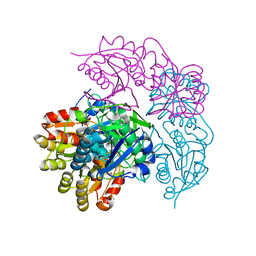 | | ECAII(D90T,K162T) MUTANT AT PH 7 | | Descriptor: | ACETIC ACID, GLYCEROL, L-asparaginase 2 | | Authors: | Lubkowski, J, Wlodawer, A. | | Deposit date: | 2019-02-08 | | Release date: | 2019-08-07 | | Last modified: | 2020-08-19 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | Opportunistic complexes of E. coli L-asparaginases with citrate anions.
Sci Rep, 9, 2019
|
|
6NX8
 
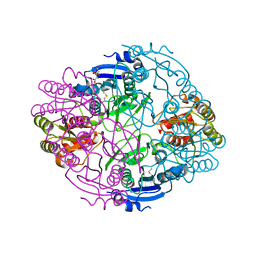 | |
6NX7
 
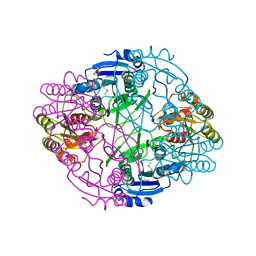 | |
1FKA
 
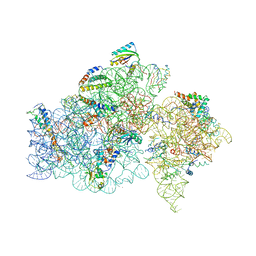 | | STRUCTURE OF FUNCTIONALLY ACTIVATED SMALL RIBOSOMAL SUBUNIT AT 3.3 A RESOLUTION | | Descriptor: | 16S RIBOSOMAL RNA, 30S RIBOSOMAL PROTEIN S10, 30S RIBOSOMAL PROTEIN S11, ... | | Authors: | Schluenzen, F, Tocilj, A, Zarivach, R, Harms, J, Gluehmann, M, Janell, D, Bashan, A, Bartels, H, Agmon, I, Franceschi, F, Yonath, A. | | Deposit date: | 2000-08-09 | | Release date: | 2000-09-04 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Structure of functionally activated small ribosomal subunit at 3.3 angstroms resolution.
Cell(Cambridge,Mass.), 102, 2000
|
|
1RSS
 
 | |
3TZN
 
 | |
3TZF
 
 | |
3TYZ
 
 | |
3TYU
 
 | |
2DPM
 
 | | DPNM DNA ADENINE METHYLTRANSFERASE FROM STREPTOCCOCUS PNEUMONIAE COMPLEXED WITH S-ADENOSYLMETHIONINE | | Descriptor: | MERCURY (II) ION, PROTEIN (ADENINE-SPECIFIC METHYLTRANSFERASE DPNII 1), S-ADENOSYLMETHIONINE | | Authors: | Tran, P.H, Korszun, Z.R, Cerritelli, S, Springhorn, S.S, Lacks, S.A. | | Deposit date: | 1998-09-03 | | Release date: | 1998-12-23 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of the DpnM DNA adenine methyltransferase from the DpnII restriction system of streptococcus pneumoniae bound to S-adenosylmethionine.
Structure, 6, 1998
|
|
1WTU
 | | TRANSCRIPTION FACTOR 1, NMR, MINIMIZED AVERAGE STRUCTURE | | Descriptor: | TRANSCRIPTION FACTOR 1 | | Authors: | Jia, X, Grove, A, Ivancic, M, Hsu, V.L, Geiduschek, E.P, Kearns, D.R. | | Deposit date: | 1996-07-29 | | Release date: | 1997-02-12 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structure of the Bacillus subtilis phage SPO1-encoded type II DNA-binding protein TF1 in solution.
J.Mol.Biol., 263, 1996
|
|
2BCW
 
 | | Coordinates of the N-terminal domain of ribosomal protein L11,C-terminal domain of ribosomal protein L7/L12 and a portion of the G' domain of elongation factor G, as fitted into cryo-em map of an Escherichia coli 70S*EF-G*GDP*fusidic acid complex | | Descriptor: | 50S ribosomal protein L11, 50S ribosomal protein L7/L12, Elongation factor G | | Authors: | Datta, P.P, Sharma, M.R, Qi, L, Frank, J, Agrawal, R.K. | | Deposit date: | 2005-10-19 | | Release date: | 2005-12-20 | | Last modified: | 2024-02-14 | | Method: | ELECTRON MICROSCOPY (11.2 Å) | | Cite: | Interaction of the G' Domain of Elongation Factor G and the C-Terminal Domain of Ribosomal Protein L7/L12 during Translocation as Revealed by Cryo-EM.
Mol.Cell, 20, 2005
|
|
