2F77
 
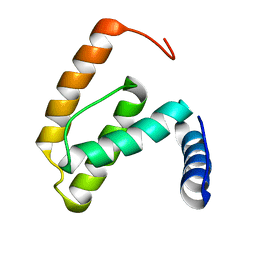 | | Solution structure of the R55F mutant of M-PMV matrix protein (p10) | | Descriptor: | Core protein p10 | | Authors: | Vlach, J, Lipov, J, Veverka, V, Lang, J, Srb, P, Rumlova, M, Hunter, E, Ruml, T, Hrabal, R. | | Deposit date: | 2005-11-30 | | Release date: | 2006-12-05 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | D-retrovirus morphogenetic switch driven by the targeting signal accessibility to Tctex-1 of dynein.
Proc.Natl.Acad.Sci.USA, 105, 2008
|
|
2F76
 
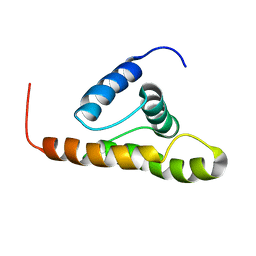 | | Solution structure of the M-PMV wild type matrix protein (p10) | | Descriptor: | Core protein p10 | | Authors: | Vlach, J, Lipov, J, Veverka, V, Lang, J, Srb, P, Rumlova, M, Hunter, E, Ruml, T, Hrabal, R. | | Deposit date: | 2005-11-30 | | Release date: | 2006-12-05 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | D-retrovirus morphogenetic switch driven by the targeting signal accessibility to Tctex-1 of dynein.
Proc.Natl.Acad.Sci.USA, 105, 2008
|
|
2LYA
 
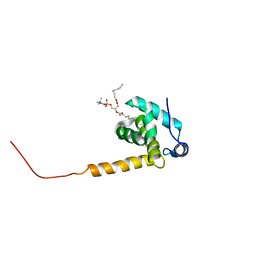 | |
2LYB
 
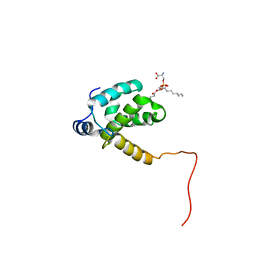 | |
2MGU
 
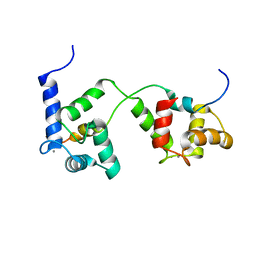 | |
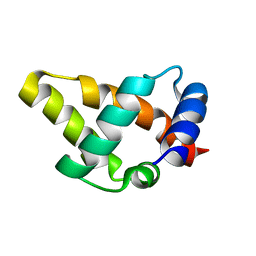
6CCJ
 
 | |
6CV8
 
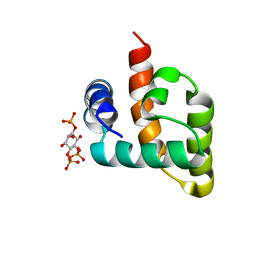 | | HADDOCK structure of the Rous sarcoma virus matrix protein (M-domain) in complex with inositol 1,4,5-trisphosphate | | Descriptor: | D-MYO-INOSITOL-1,4,5-TRIPHOSPHATE, Matrix protein p19 | | Authors: | Vlach, J, Saad, J.S. | | Deposit date: | 2018-03-27 | | Release date: | 2018-10-24 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structural basis for targeting avian sarcoma virus Gag polyprotein to the plasma membrane for virus assembly.
J. Biol. Chem., 293, 2018
|
|
6CE5
 
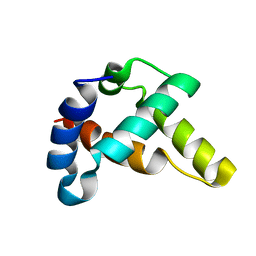 | |
6CUS
 
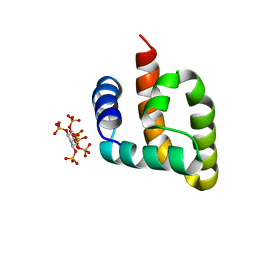 | |
6CW4
 
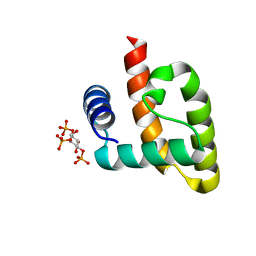 | | HADDOCK structure of the Rous sarcoma virus matrix protein (M-domain) in complex with inositol 1,3,5-trisphosphate | | Descriptor: | (1R,2S,3r,4R,5S,6s)-2,4,6-trihydroxycyclohexane-1,3,5-triyl tris[dihydrogen (phosphate)], Matrix protein p19 | | Authors: | Vlach, J, Saad, J.S. | | Deposit date: | 2018-03-29 | | Release date: | 2018-10-24 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structural basis for targeting avian sarcoma virus Gag polyprotein to the plasma membrane for virus assembly.
J. Biol. Chem., 293, 2018
|
|
5VWL
 
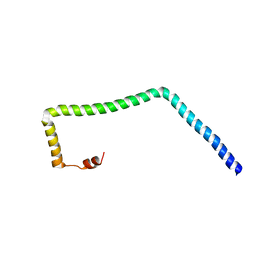 | |
8DKU
 
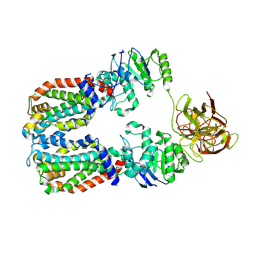 | | CryoEM structure of the A. aeolicus WzmWzt transporter bound to the native O antigen | | Descriptor: | 6-deoxy-3-O-methyl-alpha-D-mannopyranose-(1-3)-beta-D-rhamnopyranose-(1-2)-alpha-D-rhamnopyranose, ABC transporter, Transport permease protein | | Authors: | Spellmon, N, Muszynski, A, Vlach, J, Zimmer, J. | | Deposit date: | 2022-07-06 | | Release date: | 2022-09-21 | | Last modified: | 2024-06-12 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | Molecular basis for polysaccharide recognition and modulated ATP hydrolysis by the O antigen ABC transporter.
Nat Commun, 13, 2022
|
|
8DNC
 
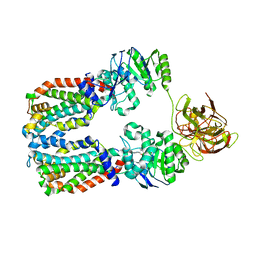 | | CryoEM structure of the A. aeolicus WzmWzt transporter bound to the native O antigen and ADP | | Descriptor: | 6-deoxy-3-O-methyl-alpha-D-mannopyranose-(1-3)-beta-D-rhamnopyranose-(1-2)-alpha-D-rhamnopyranose, ABC transporter, ADENOSINE-5'-DIPHOSPHATE, ... | | Authors: | Spellmon, N, Muszynski, A, Vlach, J, Zimmer, J. | | Deposit date: | 2022-07-11 | | Release date: | 2022-09-21 | | Last modified: | 2024-06-12 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Molecular basis for polysaccharide recognition and modulated ATP hydrolysis by the O antigen ABC transporter.
Nat Commun, 13, 2022
|
|
6ZJZ
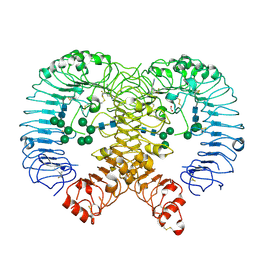 
 | | Discovery of M5049: a novel selective TLR7/8 inhibitor for treatment of autoimmunity | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 5-[(3~{R},5~{S})-3-azanyl-5-(trifluoromethyl)piperidin-1-yl]quinoline-8-carbonitrile, FORMIC ACID, ... | | Authors: | Musil, D, Lehman, M, Strauss, J. | | Deposit date: | 2020-06-29 | | Release date: | 2020-12-30 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2.489 Å) | | Cite: | Discovery of M5049: A Novel Selective Toll-Like Receptor 7/8 Inhibitor for Treatment of Autoimmunity.
J.Pharmacol.Exp.Ther., 376, 2021
|
|
8DN8
 
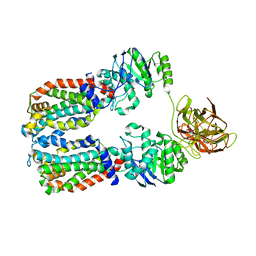 | |
8DOU
 
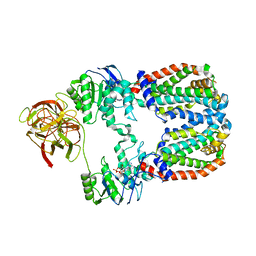 | |
8DL0
 
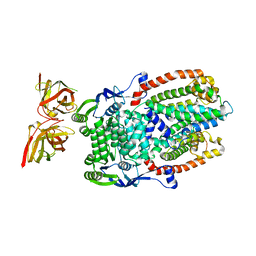 | |
8DKY
 
 | |
8DNE
 
 | |
