2B24
 
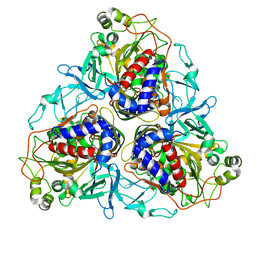 | | Crystal structure of naphthalene 1,2-dioxygenase from Rhodococcus sp. bound to indole | | Descriptor: | FE (III) ION, FE2/S2 (INORGANIC) CLUSTER, INDOLE, ... | | Authors: | Gakhar, L, Malik, Z.A, Allen, C.C, Lipscomb, D.A, Larkin, M.J, Ramaswamy, S. | | Deposit date: | 2005-09-16 | | Release date: | 2005-10-04 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structure and Increased Thermostability of Rhodococcus sp. Naphthalene 1,2-Dioxygenase.
J.Bacteriol., 187, 2005
|
|
2B1X
 
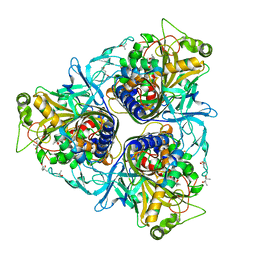 | | Crystal structure of naphthalene 1,2-dioxygenase from Rhodococcus sp. | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, FE (III) ION, FE2/S2 (INORGANIC) CLUSTER, ... | | Authors: | Gakhar, L, Malik, Z.A, Allen, C.C, Lipscomb, D.A, Larkin, M.J, Ramaswamy, S. | | Deposit date: | 2005-09-16 | | Release date: | 2005-10-04 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure and Increased Thermostability of Rhodococcus sp. Naphthalene 1,2-Dioxygenase.
J.Bacteriol., 187, 2005
|
|
5JEN
 
 | |
4RJD
 
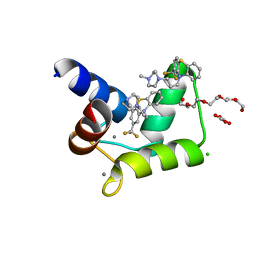 | | TFP bound in alternate orientations to calcium-saturated Calmodulin C-Domains | | Descriptor: | 10-[3-(4-METHYL-PIPERAZIN-1-YL)-PROPYL]-2-TRIFLUOROMETHYL-10H-PHENOTHIAZINE, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Feldkamp, M.D, Gakhar, L, Pandey, N, Shea, M.A. | | Deposit date: | 2014-10-08 | | Release date: | 2015-08-26 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Opposing orientations of the anti-psychotic drug trifluoperazine selected by alternate conformations of M144 in calmodulin.
Proteins, 83, 2015
|
|
3T95
 
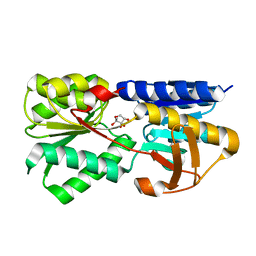 | | Crystal structure of LsrB from Yersinia pestis complexed with autoinducer-2 | | Descriptor: | (2R,4S)-2-methyl-2,3,3,4-tetrahydroxytetrahydrofuran, Autoinducer 2-binding protein lsrB | | Authors: | Kavanaugh, J.S, Gakhar, L, Horswill, A.R. | | Deposit date: | 2011-08-02 | | Release date: | 2011-12-07 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | The structure of LsrB from Yersinia pestis complexed with autoinducer-2.
Acta Crystallogr.,Sect.F, 67, 2011
|
|
3PGE
 
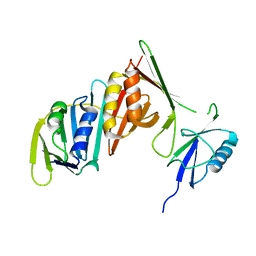 | | Structure of sumoylated PCNA | | Descriptor: | Proliferating cell nuclear antigen, SUMO-modified proliferating cell nuclear antigen | | Authors: | Freudenthal, B.D, Brogie, J.E, Gakhar, L, Washington, T. | | Deposit date: | 2010-11-01 | | Release date: | 2010-12-29 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal Structure of SUMO-Modified Proliferating Cell Nuclear Antigen.
J.Mol.Biol., 406, 2011
|
|
2HQW
 
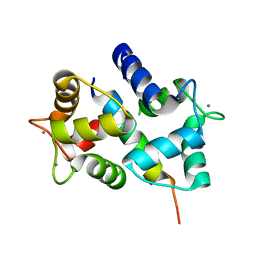 | | Crystal Structure of Ca2+/Calmodulin bound to NMDA Receptor NR1C1 peptide | | Descriptor: | CALCIUM ION, Calmodulin, Glutamate NMDA receptor subunit zeta 1 | | Authors: | Akyol, Z, Gakhar, L, Sorensen, B.R, Hell, J.H, Shea, M.A. | | Deposit date: | 2006-07-19 | | Release date: | 2007-11-13 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The NMDA Receptor NR1 C1 Region Bound to Calmodulin: Structural Insights into Functional Differences between Homologous Domains.
Structure, 15, 2007
|
|
6D4B
 
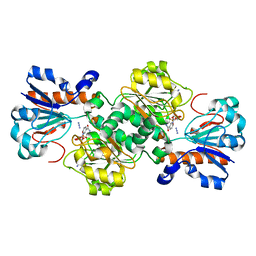 | | Crystal structure of Candida boidinii formate dehydrogenase V123A mutant complexed with NAD+ and azide | | Descriptor: | AZIDE ION, CHLORIDE ION, Formate dehydrogenase, ... | | Authors: | Guo, Q, Ye, H, Gakhar, L, Cheatum, C.M, Kohen, A. | | Deposit date: | 2018-04-17 | | Release date: | 2019-04-24 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Oscillatory Active-site Motions Correlate with Kinetic Isotope Effects in Formate Dehydrogenase
Acs Catalysis, 2019
|
|
7JXV
 
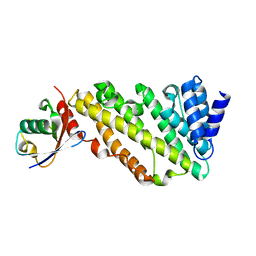 | | ANTH domain of CALM (clathrin-assembly lymphoid myeloid leukemia protein) bound to ubiquitin | | Descriptor: | Phosphatidylinositol-binding clathrin assembly protein, Ubiquitin | | Authors: | Pashkova, N, Gakhar, L, Schnicker, N.J, Piper, R.C. | | Deposit date: | 2020-08-28 | | Release date: | 2021-08-04 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | ANTH domains within CALM, HIP1R, and Sla2 recognize ubiquitin internalization signals.
Elife, 10, 2021
|
|
6D4C
 
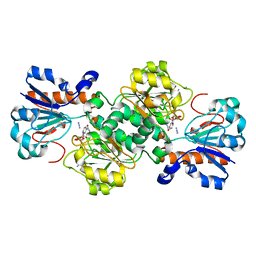 | | Crystal structure of Candida boidinii formate dehydrogenase V123G mutant complexed with NAD+ and azide | | Descriptor: | AZIDE ION, CHLORIDE ION, Formate dehydrogenase, ... | | Authors: | Guo, Q, Ye, H, Gakhar, L, Cheatum, C.M, Kohen, A. | | Deposit date: | 2018-04-17 | | Release date: | 2019-04-24 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Oscillatory Active-site Motions Correlate with Kinetic Isotope Effects in Formate Dehydrogenase
Acs Catalysis, 2019
|
|
8TOH
 
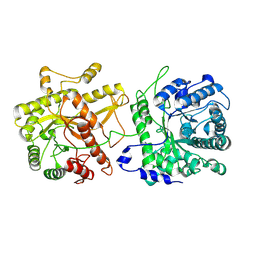 | |
8UF8
 
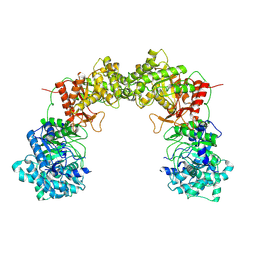 | |
5DNA
 
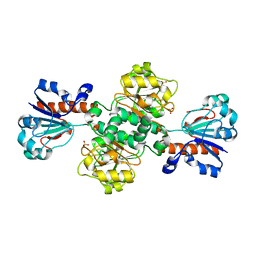 | | Crystal structure of Candida boidinii formate dehydrogenase | | Descriptor: | FORMATE DEHYDROGENASE, SULFATE ION | | Authors: | Guo, Q, Gakhar, L, Wichersham, K, Francis, K, Vardi-Kilshtain, A, Major, D.T, Cheatum, C.M, Kohen, A. | | Deposit date: | 2015-09-09 | | Release date: | 2016-05-04 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Structural and Kinetic Studies of Formate Dehydrogenase from Candida boidinii.
Biochemistry, 55, 2016
|
|
5DN9
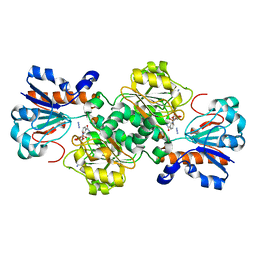 
 | | Crystal structure of Candida boidinii formate dehydrogenase complexed with NAD+ and azide | | Descriptor: | AZIDE ION, CHLORIDE ION, FDH, ... | | Authors: | Guo, Q, Gakhar, L, Wichersham, K, Francis, K, Vardi-Kilshtain, A, Major, D.T, Cheatum, C.M, Kohen, A. | | Deposit date: | 2015-09-09 | | Release date: | 2016-05-04 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Structural and Kinetic Studies of Formate Dehydrogenase from Candida boidinii.
Biochemistry, 55, 2016
|
|
3ODT
 
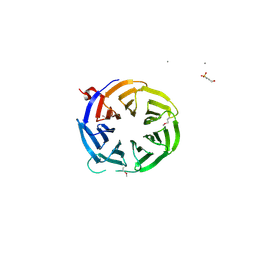 | | Crystal structure of WD40 beta propeller domain of Doa1 | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, CALCIUM ION, Protein DOA1 | | Authors: | Pashkova, N, Gakhar, L, Winistorfer, S.C, Yu, L, Ramaswamy, S, Piper, R.C. | | Deposit date: | 2010-08-11 | | Release date: | 2010-12-01 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | WD40 Repeat Propellers Define a Ubiquitin-Binding Domain that Regulates Turnover of F Box Proteins.
Mol.Cell, 40, 2010
|
|
4JJY
 
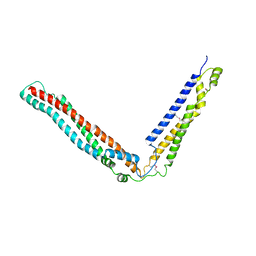 | | Alix V domain | | Descriptor: | Programmed cell death 6-interacting protein | | Authors: | Pashkova, N, Gakhar, L, Yu, L, Piper, R.C. | | Deposit date: | 2013-03-08 | | Release date: | 2013-06-19 | | Last modified: | 2013-07-10 | | Method: | X-RAY DIFFRACTION (6.503 Å) | | Cite: | The yeast alix homolog bro1 functions as a ubiquitin receptor for protein sorting into multivesicular endosomes.
Dev.Cell, 25, 2013
|
|
3L0X
 
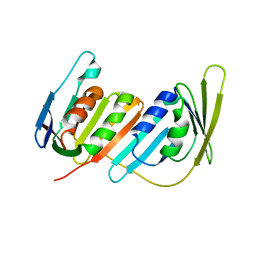 | |
3L0W
 
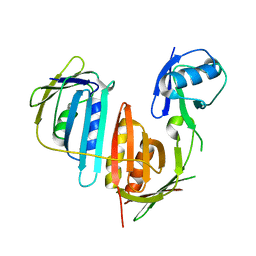 | | Structure of split monoubiquitinated PCNA with ubiquitin in position two | | Descriptor: | Monoubiquitinated Proliferating cell nuclear antigen, Proliferating cell nuclear antigen | | Authors: | Freudenthal, B.D, Gakhar, L, Ramaswamy, S, Washington, M.T. | | Deposit date: | 2009-12-10 | | Release date: | 2010-03-23 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structure of monoubiquitinated PCNA and implications for translesion synthesis and DNA polymerase exchange.
Nat.Struct.Mol.Biol., 17, 2010
|
|
3L10
 
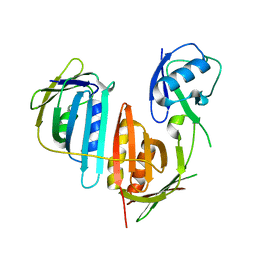 | | Structure of split monoubiquitinated PCNA with ubiquitin in position one | | Descriptor: | Monoubiquitinated Proliferating cell nuclear antigen, Proliferating cell nuclear antigen | | Authors: | Freudenthal, B.D, Gakhar, L, Ramaswamy, S, Washington, M.T. | | Deposit date: | 2009-12-10 | | Release date: | 2010-03-23 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structure of monoubiquitinated PCNA and implications for translesion synthesis and DNA polymerase exchange.
Nat.Struct.Mol.Biol., 17, 2010
|
|
5U9K
 
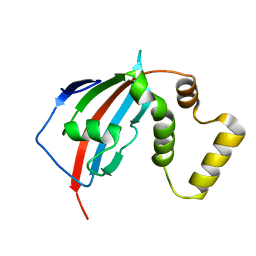 | |
3GPM
 
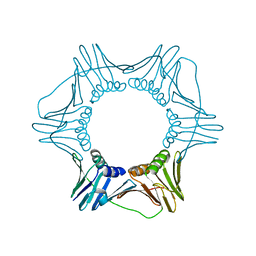 | | Structure of the trimeric form of the E113G PCNA mutant protein | | Descriptor: | Proliferating cell nuclear antigen | | Authors: | Freudenthal, B.D, Gakhar, L, Ramaswamy, S, Washington, M.T. | | Deposit date: | 2009-03-23 | | Release date: | 2009-06-16 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (3.8 Å) | | Cite: | A charged residue at the subunit interface of PCNA promotes trimer formation by destabilizing alternate subunit interactions.
Acta Crystallogr.,Sect.D, 65, 2009
|
|
5U9A
 
 | |
5U9J
 
 | | Crystal structure of the FKBP domain of human aryl hydrocarbon receptor-interacting protein-like 1 (AIPL1) complexed with geranyl geranyl pyrophoshate | | Descriptor: | Aryl hydrocarbon receptor-interacting protein-like 1 (AIPL1), GERAN-8-YL GERAN, ISOPROPYL ALCOHOL, ... | | Authors: | Yadav, R.P, Gakhar, L, Liping, Y, Artemyev, N.O. | | Deposit date: | 2016-12-16 | | Release date: | 2017-07-26 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Unique structural features of the AIPL1-FKBP domain that support prenyl lipid binding and underlie protein malfunction in blindness.
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
5U9I
 
 | | Crystal structure of the FKBP domain of human aryl hydrocarbon receptor-interacting protein-like 1 (AIPL1) complexed with S-farnesyl-L-cysteine methyl ester | | Descriptor: | Aryl hydrocarbon receptor-interacting protein-like 1 (AIPL1), FARNESYL | | Authors: | Yadav, R.P, Gakhar, L, Liping, Y, Artemyev, N.O. | | Deposit date: | 2016-12-16 | | Release date: | 2017-07-26 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Unique structural features of the AIPL1-FKBP domain that support prenyl lipid binding and underlie protein malfunction in blindness.
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
5V35
 
 | | Crystal structure of V71F mutant of the FKBP domain of human aryl hydrocarbon receptor-interacting protein-like 1 (AIPL1) complexed with S-farnesyl-L-cysteine methyl ester | | Descriptor: | Aryl hydrocarbon receptor-interacting protein-like 1 (AIPL1), FARNESYL | | Authors: | Yadav, R.P, Gakhar, L, Liping, Y, Artemyev, N.O. | | Deposit date: | 2017-03-06 | | Release date: | 2017-07-26 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Unique structural features of the AIPL1-FKBP domain that support prenyl lipid binding and underlie protein malfunction in blindness.
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
