5Z9I
 
 | |
5UL7
 
 | |
8H67
 
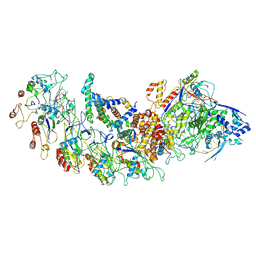 | | type I-B Cascade bound to a PAM-containing dsDNA target at 3.8 angstrom resolution. | | Descriptor: | CRISPR RNA, CRISPR associated protein Cas11b, CRISPR associated protein Cas5, ... | | Authors: | Xiao, Y, Lu, M, Yu, C, Zhang, Y. | | Deposit date: | 2022-10-15 | | Release date: | 2024-05-01 | | Last modified: | 2025-05-28 | | Method: | ELECTRON MICROSCOPY (3.8 Å) | | Cite: | Structure and genome editing of type I-B CRISPR-Cas.
Nat Commun, 15, 2024
|
|
8IP0
 
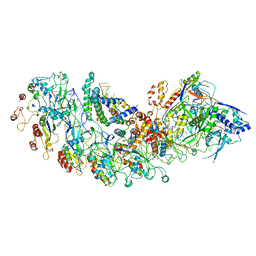 | | Cryo-EM structure of type I-B Cascade bound to a PAM-containing dsDNA target at 3.6 angstrom resolution | | Descriptor: | CRISPR associated protein Cas11b, DNA (41-MER), DNA (5'-D(P*AP*AP*AP*AP*AP*AP*AP*AP*AP*A)-3'), ... | | Authors: | Xiao, Y, Lu, M, Yu, C, Zhang, Y. | | Deposit date: | 2023-03-13 | | Release date: | 2024-03-20 | | Last modified: | 2024-05-29 | | Method: | ELECTRON MICROSCOPY (3.6 Å) | | Cite: | Structure and genome editing of type I-B CRISPR-Cas.
Nat Commun, 15, 2024
|
|
7CUT
 
 | | Crystal structure of the SARS-CoV-2 (COVID-19) main protease in complex with Z-VAD-FMK | | Descriptor: | 3C protein, DIMETHYL SULFOXIDE, GLYCEROL, ... | | Authors: | Lu, M, Yang, H.T, Wang, Z.Y, Zhao, Y, Xing, Y.F. | | Deposit date: | 2020-08-24 | | Release date: | 2021-06-23 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.82 Å) | | Cite: | Identification of proteasome and caspase inhibitors targeting SARS-CoV-2 M pro .
Signal Transduct Target Ther, 6, 2021
|
|
7CUU
 
 | | Crystal structure of the SARS-CoV-2 (COVID-19) main protease in complex with MG132 | | Descriptor: | 3C protein, DIMETHYL SULFOXIDE, GLYCEROL, ... | | Authors: | Lu, M, Yang, H.T, Wang, Z.Y, Zhao, Y, Xing, Y.F. | | Deposit date: | 2020-08-24 | | Release date: | 2021-06-23 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.68 Å) | | Cite: | Identification of proteasome and caspase inhibitors targeting SARS-CoV-2 M pro .
Signal Transduct Target Ther, 6, 2021
|
|
7EAX
 | | Crystal complex of p53-V272M and antimony ion | | Descriptor: | ANTIMONY (III) ION, Cellular tumor antigen p53, ZINC ION | | Authors: | Lu, M, Tang, Y. | | Deposit date: | 2021-03-08 | | Release date: | 2022-02-16 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Repurposing antiparasitic antimonials to noncovalently rescue temperature-sensitive p53 mutations.
Cell Rep, 39, 2022
|
|
7V97
 
 | | Arsenic-bound p53 DNA-binding domain mutant V272M | | Descriptor: | ARSENIC, Cellular tumor antigen p53, ZINC ION | | Authors: | Lu, M, Xing, Y.F, Wang, Z.Y, Ni, Y, Song, H.X. | | Deposit date: | 2021-08-24 | | Release date: | 2022-08-31 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.02 Å) | | Cite: | Diverse rescue potencies of p53 mutations to ATO are predetermined by intrinsic mutational properties.
Sci Transl Med, 15, 2023
|
|
9JI7
 
 | | NADP-dependent oxidoreductase complexed with NADP and substrate 2 | | Descriptor: | (1Z,3E,5E,7S,8R,9S,10S,11R,13R,15S,16Z,18E,24S)-11-ethyl-2,7-dihydroxy-10-methyl-21,25-diazatetracyclo[22.2.1.08,15.09,13]heptacosa-1,3,5,16,18-pentaene-20,26,27-trione, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, NADP-dependent oxidoreductase | | Authors: | Li, Y, Zhu, D, Xie, X, Li, F, Lu, M. | | Deposit date: | 2024-09-11 | | Release date: | 2025-06-18 | | Method: | X-RAY DIFFRACTION (2.53 Å) | | Cite: | Structural Basis for Medium-Chain Dehydrogenase/Reductase-Catalyzed Reductive Cyclization in Polycyclic Tetramate Macrolactam Biosynthesis.
J.Am.Chem.Soc., 147, 2025
|
|
9JML
 
 | | NADP-dependent oxidoreductase complexed with NADP and substrate 1 | | Descriptor: | (1Z,3Z,5Z,7S,8R,9S,10S,11R,13R,15R,16Z,18Z,24S)-11-ethyl-2,7-dihydroxy-10-methyl-21,25-diazatetracyclo[22.2.1.08,15.09,13]heptacosa-1,3,5,16,18-pentaene-20,26,27-trione, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, NADP-dependent oxidoreductase | | Authors: | Li, Y, Zhu, D, Xie, X, Li, F, Lu, M. | | Deposit date: | 2024-09-20 | | Release date: | 2025-06-18 | | Method: | X-RAY DIFFRACTION (2.02 Å) | | Cite: | Structural Basis for Medium-Chain Dehydrogenase/Reductase-Catalyzed Reductive Cyclization in Polycyclic Tetramate Macrolactam Biosynthesis.
J.Am.Chem.Soc., 147, 2025
|
|
9JP8
 
 | | NADP-dependent oxidoreductase | | Descriptor: | NADP-dependent oxidoreductase, SULFATE ION, UNKNOWN LIGAND | | Authors: | Li, Y, Zhu, D, Xie, X, Li, F, Lu, M. | | Deposit date: | 2024-09-25 | | Release date: | 2025-06-18 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural Basis for Medium-Chain Dehydrogenase/Reductase-Catalyzed Reductive Cyclization in Polycyclic Tetramate Macrolactam Biosynthesis.
J.Am.Chem.Soc., 147, 2025
|
|
9JLY
 
 | | NADP-dependent oxidoreductase complexed with NADP | | Descriptor: | NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, NADP-dependent oxidoreductase | | Authors: | Li, Y, Zhu, D, Xie, X, Li, F, Lu, M. | | Deposit date: | 2024-09-19 | | Release date: | 2025-06-18 | | Method: | X-RAY DIFFRACTION (3.11 Å) | | Cite: | Structural Basis for Medium-Chain Dehydrogenase/Reductase-Catalyzed Reductive Cyclization in Polycyclic Tetramate Macrolactam Biosynthesis.
J.Am.Chem.Soc., 147, 2025
|
|
9JJ4
 
 | | NADP-dependent oxidoreductase complexed with NADP and substrate 1a | | Descriptor: | (1Z,3E,5S,7S,8R,9S,10S,11R,13R,15R,16S,18Z,25S)-11-ethyl-2,7-dihydroxy-10-methyl-21,26-diazapentacyclo[23.2.1.09,13.08,15.05,16]octacosa-1(2),3,18-triene-20,27,28-trione, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, NADP-dependent oxidoreductase | | Authors: | Li, Y, Zhu, D, Xie, X, Li, F, Lu, M. | | Deposit date: | 2024-09-12 | | Release date: | 2025-06-18 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structural Basis for Medium-Chain Dehydrogenase/Reductase-Catalyzed Reductive Cyclization in Polycyclic Tetramate Macrolactam Biosynthesis.
J.Am.Chem.Soc., 147, 2025
|
|
9JLK
 
 | | NADP-dependent oxidoreductase | | Descriptor: | NADP-dependent oxidoreductase | | Authors: | Li, Y, Zhu, D, Xie, X, Li, F, Lu, M. | | Deposit date: | 2024-09-19 | | Release date: | 2025-06-18 | | Method: | X-RAY DIFFRACTION (2.33 Å) | | Cite: | Structural Basis for Medium-Chain Dehydrogenase/Reductase-Catalyzed Reductive Cyclization in Polycyclic Tetramate Macrolactam Biosynthesis.
J.Am.Chem.Soc., 147, 2025
|
|
9JM9
 
 | | NADP-dependent oxidoreductase complexed with NADP and substrate 3 | | Descriptor: | (1Z,3E,5E,7R,8S,10R,11R,12S,15S,16E,18E,25S)-11-ethyl-2-hydroxy-10-methyl-21,26-diazatetracyclo[23.2.1.07,15.08,12]octacosa-1,3,5,13,16,18-hexaene-20,27,28-trione, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, NADP-dependent oxidoreductase | | Authors: | Li, Y, Zhu, D, Xie, X, Li, F, Lu, M. | | Deposit date: | 2024-09-20 | | Release date: | 2025-06-18 | | Method: | X-RAY DIFFRACTION (3.38 Å) | | Cite: | Structural Basis for Medium-Chain Dehydrogenase/Reductase-Catalyzed Reductive Cyclization in Polycyclic Tetramate Macrolactam Biosynthesis.
J.Am.Chem.Soc., 147, 2025
|
|
3KQI
 
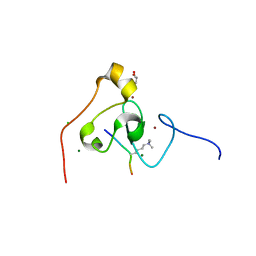 | | crystal structure of PHF2 PHD domain complexed with H3K4Me3 peptide | | Descriptor: | CHLORIDE ION, GLYCEROL, H3K4Me3 peptide, ... | | Authors: | Wen, H, Li, J.Z, Song, T, Lu, M, Lee, M. | | Deposit date: | 2009-11-17 | | Release date: | 2010-02-02 | | Last modified: | 2025-03-26 | | Method: | X-RAY DIFFRACTION (1.78 Å) | | Cite: | Recognition of histone H3K4 trimethylation by the plant homeodomain of PHF2 modulates histone demethylation.
J.Biol.Chem., 285, 2010
|
|
1CE0
 
 | |
1JCD
 | | Crystal Structure of a Novel Alanine-Zipper Trimer at 1.3 A Resolution, I6A,L9A,V13A,L16A,V20A,L23A,V27A,M30A,V34A,L48A,M51A mutations | | Descriptor: | MAJOR OUTER MEMBRANE LIPOPROTEIN | | Authors: | Liu, J, Lu, M. | | Deposit date: | 2001-06-08 | | Release date: | 2003-06-17 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | An Alanine-Zipper Structure
Determined by Long Range
Intermolecular Interactions
J.Biol.Chem., 277, 2002
|
|
1K34
 
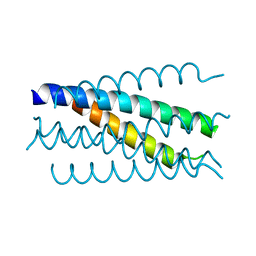 | | Crystal structure analysis of gp41 core mutant | | Descriptor: | Transmembrane glycoprotein GP41 | | Authors: | Shu, W, Lu, M. | | Deposit date: | 2001-10-01 | | Release date: | 2001-10-10 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Interhelical interactions in the gp41 core: implications for activation of HIV-1 membrane fusion.
Biochemistry, 41, 2002
|
|
1JQ0
 | | Mutation that destabilize the gp41 core: determinants for stabilizing the SIV/CPmac envelope glycoprotein complex. Mutant structure. | | Descriptor: | gp41 envelope protein | | Authors: | Liu, J, Wang, S, LaBranche, C.C, Hoxie, J.A, Lu, M. | | Deposit date: | 2001-08-03 | | Release date: | 2002-04-24 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Mutations that destabilize the gp41 core are determinants for stabilizing the simian immunodeficiency virus-CPmac envelope glycoprotein complex.
J.Biol.Chem., 277, 2002
|
|
5ZQI
 
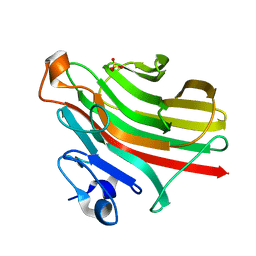 | | Alginate lyase AlgAT5 from Polysaccharide Lyase family 7 | | Descriptor: | Alginate lyase AlgAT5, SULFATE ION | | Authors: | Su, H, Dong, S, Feng, Y.G, Ji, S.Q, Lu, M, Li, F.L. | | Deposit date: | 2018-04-19 | | Release date: | 2019-04-24 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Alginate lyase AlgAT5 from Polysaccharide Lyase family 7
To Be Published
|
|
1JPX
 
 | | Mutation that destabilize the gp41 core: determinants for stabilizing the SIV/CPmac envelope glycoprotein complex. Wild type. | | Descriptor: | gp41 envelope protein | | Authors: | Liu, J, Wang, S, LaBranche, C.C, Hoxie, J.A, Lu, M. | | Deposit date: | 2001-08-03 | | Release date: | 2002-04-24 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Mutations that destabilize the gp41 core are determinants for stabilizing the simian immunodeficiency virus-CPmac envelope glycoprotein complex.
J.Biol.Chem., 277, 2002
|
|
1JCC
 | | Crystal Structure of a Novel Alanine-Zipper Trimer at 1.7 A Resolution, V13A,L16A,V20A,L23A,V27A,M30A,V34A mutations | | Descriptor: | MAJOR OUTER MEMBRANE LIPOPROTEIN, ZINC ION | | Authors: | Liu, J, Dai, J, Lu, M. | | Deposit date: | 2001-06-08 | | Release date: | 2003-06-17 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Zinc-Mediated Helix Capping in A Triple-Helical Protein
Biochemistry, 42, 2003
|
|
1K33
 
 | | Crystal structure analysis of the gp41 core mutant | | Descriptor: | Transmembrane glycoprotein GP41 | | Authors: | Shu, W, Lu, M. | | Deposit date: | 2001-10-01 | | Release date: | 2001-10-10 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Interhelical interactions in the gp41 core: implications for activation of HIV-1 membrane fusion.
Biochemistry, 41, 2002
|
|
1SZT
 
 | |
