7TOY
 
 | |
7TOZ
 
 | |
7TP0
 
 | |
7TP2
 
 | |
7TP8
 
 | |
7TPA
 
 | |
7TP1
 
 | |
7TPC
 
 | |
7TPE
 
 | |
7TPH
 
 | |
7TPF
 
 | |
7TLA
 
 | |
7TPL
 
 | |
7TP9
 
 | |
7TLD
 
 | |
7TLB
 
 | |
7TP7
 
 | |
7TLC
 
 | |
7THE
 
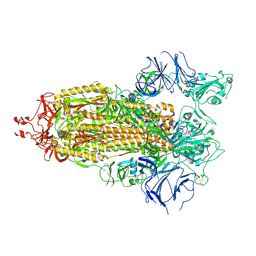
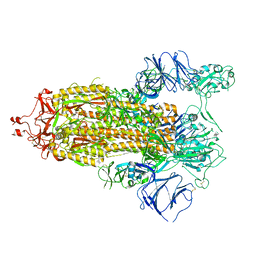
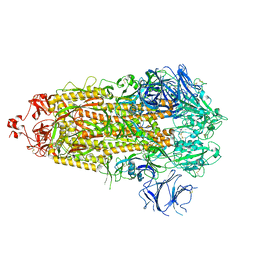
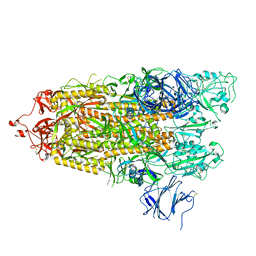
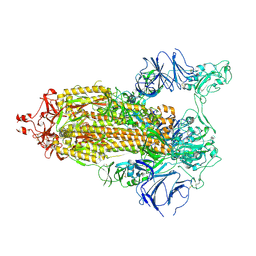
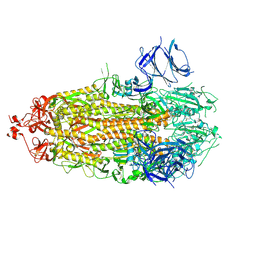
 | | Structure of RBD directed antibody DH1042 in complex with SARS-CoV-2 spike: Local refinement of RBD-Fab interface | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, DH1042 Fab Heavy Chain, DH1042 Fab Light Chain, ... | | Authors: | May, A.J, Manne, K, Acharya, P. | | Deposit date: | 2022-01-10 | | Release date: | 2022-02-16 | | Last modified: | 2022-08-03 | | Method: | ELECTRON MICROSCOPY (3.87 Å) | | Cite: | Structural diversity of the SARS-CoV-2 Omicron spike.
Mol.Cell, 82, 2022
|
|
7TOW
 
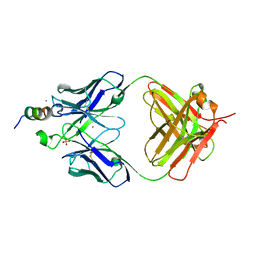 | |
3PRE
 
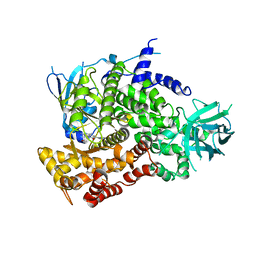 | | Quinazolines with intra-molecular hydrogen bonding scaffold (iMHBS) as PI3K/mTOR dual inhibitors. | | Descriptor: | 2-amino-8-(trans-4-methoxycyclohexyl)-4-methyl-6-(1H-pyrazol-3-yl)pyrido[2,3-d]pyrimidin-7(8H)-one, Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit gamma isoform | | Authors: | Knighton, D.R, Greasley, S.E, Rodgers, C.M.-L. | | Deposit date: | 2010-11-29 | | Release date: | 2011-02-09 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.91 Å) | | Cite: | Quinazolines with intra-molecular hydrogen bonding scaffold (iMHBS) as PI3K/mTOR dual inhibitors.
Bioorg.Med.Chem.Lett., 21, 2011
|
|
3PRZ
 
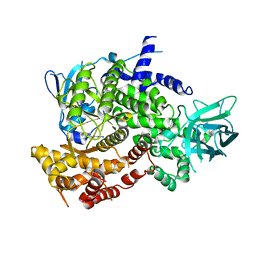 | |
3PS6
 
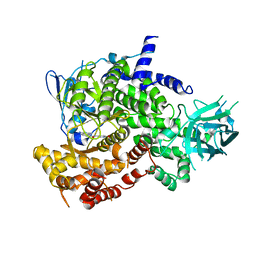 | |
6J69
 
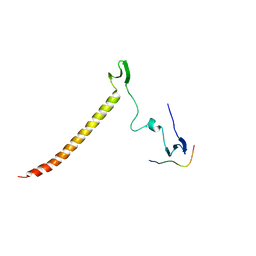 | | Structure of KIBRA and Dendrin Complex | | Descriptor: | Peptide from Dendrin, Protein KIBRA | | Authors: | Lin, Z, Yang, Z, Ji, Z, Zhang, M. | | Deposit date: | 2019-01-14 | | Release date: | 2019-03-20 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.753 Å) | | Cite: | Kibra Modulates Learning and Memory via Binding to Dendrin.
Cell Rep, 26, 2019
|
|
6JFP
 
 | | Crystal structure of the beta-glucosidase Bgl15 | | Descriptor: | beta-D-glucopyranose, beta-glucosidase 15 | | Authors: | Xie, W, Chen, R. | | Deposit date: | 2019-02-11 | | Release date: | 2020-02-12 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Engineering of beta-Glucosidase Bgl15 with Simultaneously Enhanced Glucose Tolerance and Thermostability To Improve Its Performance in High-Solid Cellulose Hydrolysis.
J.Agric.Food Chem., 68, 2020
|
|
