3PJB
 
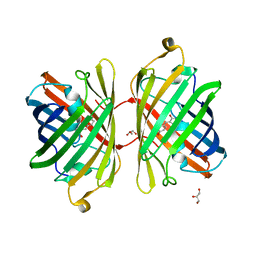 | |
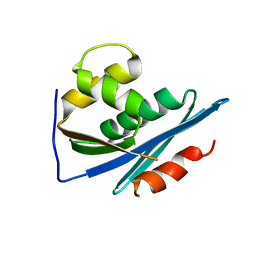
1RDH
 
 | |
3PJ5
 
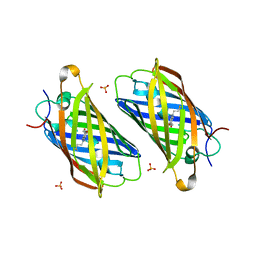 | |
5CNA
 
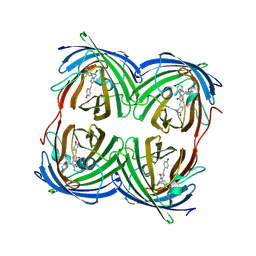
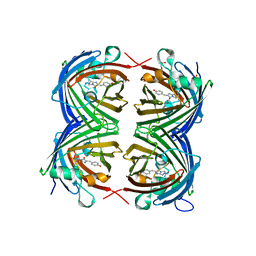
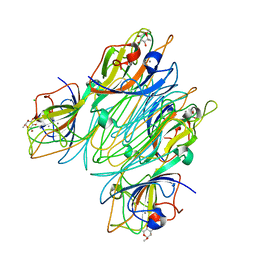 | | REFINED STRUCTURE OF CONCANAVALIN A COMPLEXED WITH ALPHA-METHYL-D-MANNOPYRANOSIDE AT 2.0 ANGSTROMS RESOLUTION AND COMPARISON WITH THE SACCHARIDE-FREE STRUCTURE | | Descriptor: | CALCIUM ION, CHLORIDE ION, CONCANAVALIN A, ... | | Authors: | Naismith, J.H, Emmerich, C, Habash, J, Harrop, S.J, Helliwell, J.R, Hunter, W.N, Raftery, J, Kalb(Gilboa), A.J, Yariv, J. | | Deposit date: | 1994-02-11 | | Release date: | 1994-05-31 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Refined structure of concanavalin A complexed with methyl alpha-D-mannopyranoside at 2.0 A resolution and comparison with the saccharide-free structure.
Acta Crystallogr.,Sect.D, 50, 1994
|
|
5EXB
 | |
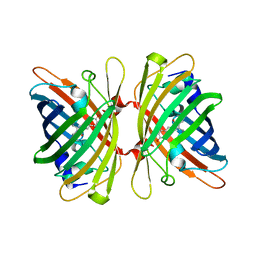
4HE4
 
 | |
5EXC
 
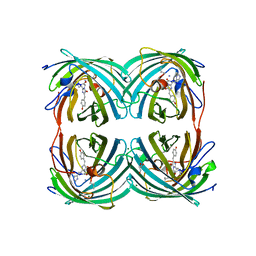 | |
6M9Z
 
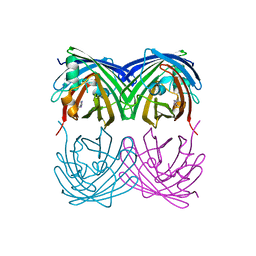 | |
6M9Y
 | |
6MAS
 
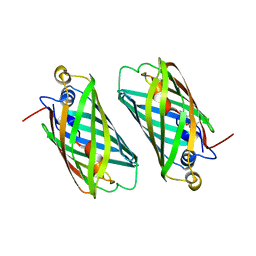
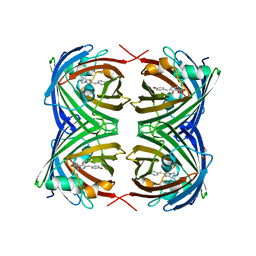 | | X-ray Structure of Branchiostoma floridae fluorescent protein lanFP10G | | Descriptor: | GLYCEROL, Uncharacterized protein | | Authors: | Muslinkina, L, Pletneva, N, Pletnev, V, Pletnev, S. | | Deposit date: | 2018-08-28 | | Release date: | 2019-03-13 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Structural Factors Enabling Successful GFP-Like Proteins with Alanine as the Third Chromophore-Forming Residue.
J. Mol. Biol., 431, 2019
|
|
6M9X
 
 | |
1HET
 
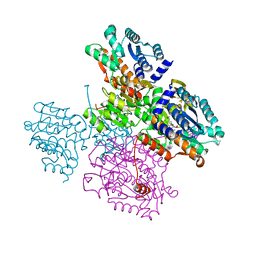
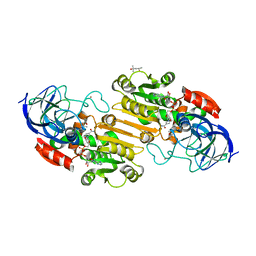 | | atomic X-ray structure of liver alcohol dehydrogenase containing a hydroxide adduct to NADH | | Descriptor: | (4R)-2-METHYLPENTANE-2,4-DIOL, ALCOHOL DEHYDROGENASE E CHAIN, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, ... | | Authors: | Meijers, R, Morris, R.J, Adolph, H.W, Merli, A, Lamzin, V.S, Cedergen-Zeppezauer, E.S. | | Deposit date: | 2000-11-25 | | Release date: | 2001-05-31 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.15 Å) | | Cite: | On the Enzymatic Activation of Nadh
J.Biol.Chem., 276, 2001
|
|
4JGE
 
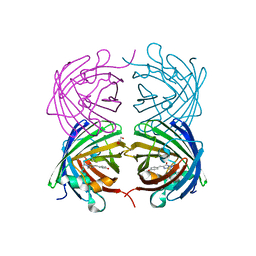 | |
4JEO
 
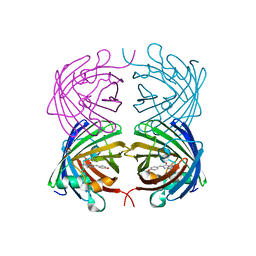 | |
4JF9
 
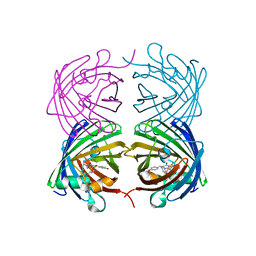 | |
4HVF
 
 | |
5TOV
 
 | |
5TOW
 
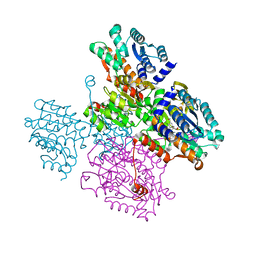 | | Crystal structure of the inactive form of S-adenosyl-L-homocysteine hydrolase from Thermotoga maritima in ternary complex with NADH and Adenosine | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 1,4-DIHYDRONICOTINAMIDE ADENINE DINUCLEOTIDE, ADENOSINE, ... | | Authors: | Czyrko, J, Brzezinski, K. | | Deposit date: | 2016-10-19 | | Release date: | 2017-07-05 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | S-adenosyl-L-homocysteine hydrolase from a hyperthermophile (Thermotoga maritima) is expressed in Escherichia coli in inactive form - Biochemical and structural studies.
Int. J. Biol. Macromol., 104, 2017
|
|
5T3F
 
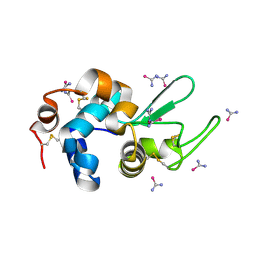 | |
5T3J
 
 | |
5T3I
 | |
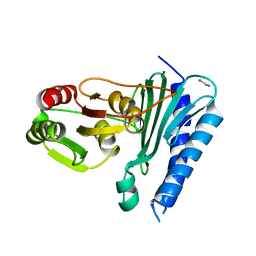
5T3H
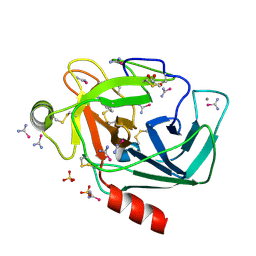 
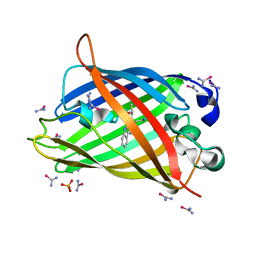
 | | bovine trypsin soaked with selenourea for 5 min | | Descriptor: | BENZAMIDINE, CALCIUM ION, Cationic trypsin, ... | | Authors: | Luo, Z, Dauter, Z. | | Deposit date: | 2016-08-25 | | Release date: | 2016-11-30 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Selenourea: a convenient phasing vehicle for macromolecular X-ray crystal structures.
Sci Rep, 6, 2016
|
|
5T3G
 
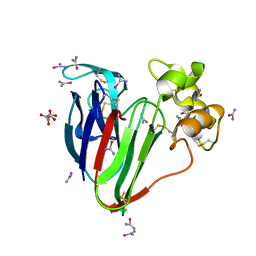 | | thaumatin soaked with selenourea for 10 min | | Descriptor: | L(+)-TARTARIC ACID, Thaumatin-1, selenourea | | Authors: | Luo, Z, Dauter, Z. | | Deposit date: | 2016-08-25 | | Release date: | 2016-11-30 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Selenourea: a convenient phasing vehicle for macromolecular X-ray crystal structures.
Sci Rep, 6, 2016
|
|
5T3L
 
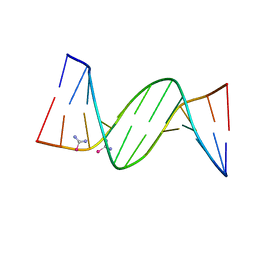 | | B-DNA (CGCGAATTCGCG)2 soaked with selenourea for 1 min | | Descriptor: | DNA (5'-D(*CP*GP*CP*GP*AP*AP*TP*TP*CP*GP*CP*G)-3'), selenourea | | Authors: | Luo, Z, Dauter, Z. | | Deposit date: | 2016-08-25 | | Release date: | 2016-11-30 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.58 Å) | | Cite: | Selenourea: a convenient phasing vehicle for macromolecular X-ray crystal structures.
Sci Rep, 6, 2016
|
|
1XYZ
 
 | |
