1RDC
 
 | |
1KVA
 
 | | E. COLI RIBONUCLEASE HI D134A MUTANT | | 分子名称: | RIBONUCLEASE H | | 著者 | Kashiwagi, T, Jeanteur, D, Haruki, M, Katayanagi, K, Kanaya, S, Morikawa, K. | | 登録日 | 1996-10-04 | | 公開日 | 1997-03-12 | | 最終更新日 | 2021-11-03 | | 実験手法 | X-RAY DIFFRACTION (1.8 Å) | | 主引用文献 | Proposal for new catalytic roles for two invariant residues in Escherichia coli ribonuclease HI.
Protein Eng., 9, 1996
|
|
1KVC
 
 | | E. COLI RIBONUCLEASE HI D134N MUTANT | | 分子名称: | RIBONUCLEASE H | | 著者 | Kashiwagi, T, Jeanteur, D, Haruki, M, Katayanagi, K, Kanaya, S, Morikawa, K. | | 登録日 | 1996-10-04 | | 公開日 | 1997-03-12 | | 最終更新日 | 2024-02-14 | | 実験手法 | X-RAY DIFFRACTION (1.9 Å) | | 主引用文献 | Proposal for new catalytic roles for two invariant residues in Escherichia coli ribonuclease HI.
Protein Eng., 9, 1996
|
|
7E0G
 | |
5X0K
 
 | |
5XI8
 
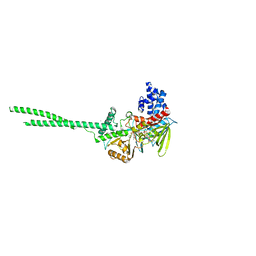
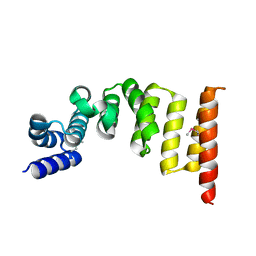 | | Structure and function of the TPR domain | | 分子名称: | Beta-barrel assembly-enhancing protease | | 著者 | Tanaka, Y, Tsukazaki, T. | | 登録日 | 2017-04-26 | | 公開日 | 2017-10-18 | | 最終更新日 | 2017-12-13 | | 実験手法 | X-RAY DIFFRACTION (1.7 Å) | | 主引用文献 | The TPR domain of BepA is required for productive interaction with substrate proteins and the beta-barrel assembly machinery complex.
Mol. Microbiol., 106, 2017
|
|
1RDD
 
 | |
2CWR
 | |
2CZN
 
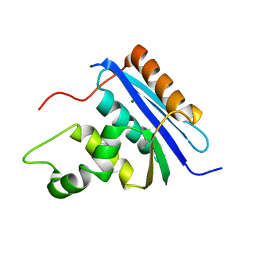
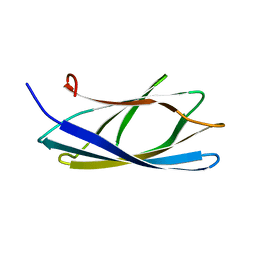
 | | Solution structure of the chitin-binding domain of hyperthermophilic chitinase from pyrococcus furiosus | | 分子名称: | chitinase | | 著者 | Uegaki, T, Ikegami, T, Nakamura, T, Hagihara, Y, Mine, S, Inoue, T, Matsumura, H, Ataka, M, Ishikawa, K. | | 登録日 | 2005-07-13 | | 公開日 | 2006-07-18 | | 最終更新日 | 2024-05-29 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Tertiary structure and carbohydrate recognition by the chitin-binding domain of a hyperthermophilic chitinase from Pyrococcus furiosus.
J.Mol.Biol., 381, 2008
|
|
5X0G
 
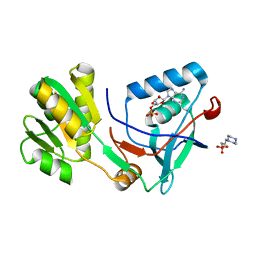 | | Free serine kinase (E30A mutant) in complex with ADP | | 分子名称: | 2-hydroxy-3-[4-(2-hydroxy-3-sulfopropyl)piperazin-1-yl]propane-1-sulfonic acid, ADENOSINE MONOPHOSPHATE, ADENOSINE-5'-DIPHOSPHATE, ... | | 著者 | Nagata, R, Fujihashi, M, Miki, K. | | 登録日 | 2017-01-20 | | 公開日 | 2017-04-12 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (1.9 Å) | | 主引用文献 | Structural Study on the Reaction Mechanism of a Free Serine Kinase Involved in Cysteine Biosynthesis
ACS Chem. Biol., 12, 2017
|
|
5X0J
 
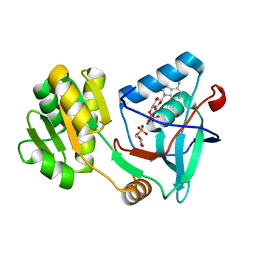 | | Free serine kinase (E30Q mutant) in complex with phosphoserine and AMP | | 分子名称: | ADENOSINE MONOPHOSPHATE, Free serine kinase, MAGNESIUM ION, ... | | 著者 | Nagata, R, Fujihashi, M, Miki, K. | | 登録日 | 2017-01-20 | | 公開日 | 2017-04-12 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (1.43 Å) | | 主引用文献 | Structural Study on the Reaction Mechanism of a Free Serine Kinase Involved in Cysteine Biosynthesis
ACS Chem. Biol., 12, 2017
|
|
5X0F
 
 | |
3LHM
 
 | |
1RDA
 | |
3VI7
 
 | |
1WSI
 
 | |
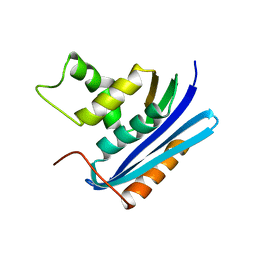
1WSH
 
 | |
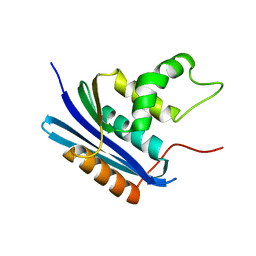
1WSJ
 
 | |
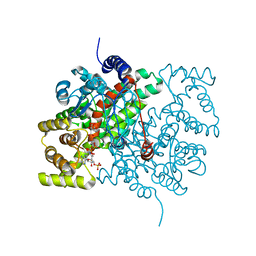
2CTS
 
 | |
7F3O
 
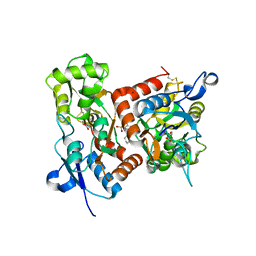 | | Crystal structure of the GluA2o LBD in complex with glutamate and TAK-653 | | 分子名称: | 7-(4-cyclohexyloxyphenyl)-9-methyl-4$l^{6}-thia-1$l^{4},5,8-triazabicyclo[4.4.0]deca-1(10),6,8-triene 4,4-dioxide, ACETATE ION, GLUTAMIC ACID, ... | | 著者 | Sogabe, S, Igaki, S, Hirokawa, A, Zama, Y, Lane, W, Snell, G. | | 登録日 | 2021-06-16 | | 公開日 | 2021-07-28 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (1.44 Å) | | 主引用文献 | Strictly regulated agonist-dependent activation of AMPA-R is the key characteristic of TAK-653 for robust synaptic responses and cognitive improvement.
Sci Rep, 11, 2021
|
|
3VI5
 
 | | Human hematopoietic prostaglandin D synthase inhibitor complex structures | | 分子名称: | 1-amino-9,10-dioxo-4-[(4-sulfamoylphenyl)amino]-9,10-dihydroanthracene-2-sulfonic acid, CALCIUM ION, GLUTATHIONE, ... | | 著者 | Kado, Y, Inoue, T. | | 登録日 | 2011-09-21 | | 公開日 | 2012-04-18 | | 最終更新日 | 2024-03-20 | | 実験手法 | X-RAY DIFFRACTION (2 Å) | | 主引用文献 | Human hematopoietic prostaglandin D synthase inhibitor complex structures
J.Biochem., 151, 2012
|
|
7F3A
 
 | | Arabidopsis thaliana GH1 beta-glucosidase AtBGlu42 | | 分子名称: | Beta-glucosidase 42, GLYCEROL | | 著者 | Horikoshi, S, Saburi, W, Yu, J, Yao, M. | | 登録日 | 2021-06-16 | | 公開日 | 2022-03-16 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (1.7 Å) | | 主引用文献 | Substrate specificity of glycoside hydrolase family 1 beta-glucosidase AtBGlu42 from Arabidopsis thaliana and its molecular mechanism.
Biosci.Biotechnol.Biochem., 86, 2022
|
|
2CSC
 
 | |
1HYT
 
 | |
2LHM
 
 | |
