7K56
 
 | | Structure of VCP dodecamer purified from H1299 cells | | 分子名称: | Transitional endoplasmic reticulum ATPase | | 著者 | Yu, G, Bai, Y, Li, K, Jiang, W, Zhang, Z.Y. | | 登録日 | 2020-09-16 | | 公開日 | 2021-10-13 | | 最終更新日 | 2024-05-29 | | 実験手法 | ELECTRON MICROSCOPY (3.9 Å) | | 主引用文献 | Cryo-electron microscopy structures of VCP/p97 reveal a new mechanism of oligomerization regulation.
Iscience, 24, 2021
|
|
7K63
 
 | |
7K61
 
 | |
7K5Y
 | |
7K60
 
 | |
7K5X
 | |
8DK5
 
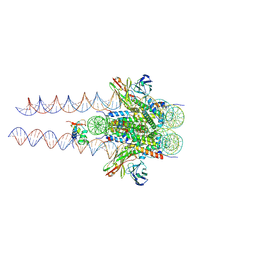
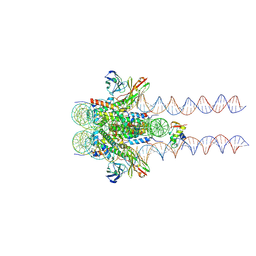
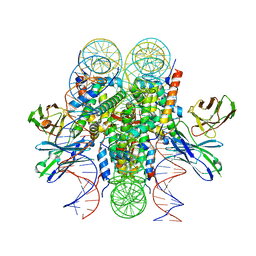 | | Structure of 187bp LIN28b nucleosome with site 0 mutation | | 分子名称: | DNA (187-MER), Histone H2A type 2-C, Histone H2B type 2-E, ... | | 著者 | Lian, T, Guan, R, Bai, Y. | | 登録日 | 2022-07-02 | | 公開日 | 2023-06-28 | | 実験手法 | ELECTRON MICROSCOPY (2.71 Å) | | 主引用文献 | Structural mechanism of LIN28B nucleosome targeting by OCT4.
Mol.Cell, 83, 2023
|
|
3T4B
 
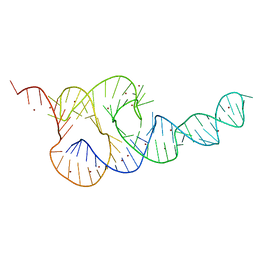 | | Crystal Structure of the HCV IRES pseudoknot domain | | 分子名称: | HCV IRES pseudoknot domain plus crystallization module, NICKEL (II) ION | | 著者 | Berry, K.E, Waghray, S, Mortimer, S.A, Bai, Y, Doudna, J.A. | | 登録日 | 2011-07-25 | | 公開日 | 2011-10-12 | | 最終更新日 | 2024-02-28 | | 実験手法 | X-RAY DIFFRACTION (3.55 Å) | | 主引用文献 | Crystal structure of the HCV IRES central domain reveals strategy for start-codon positioning.
Structure, 19, 2011
|
|
7D1O
 
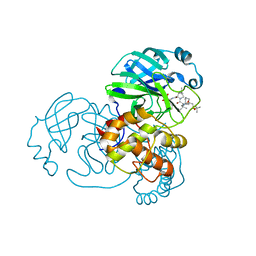 | | Crystal structure of SARS-Cov-2 main protease with narlaprevir | | 分子名称: | (1R,2S,5S)-3-[N-({1-[(tert-butylsulfonyl)methyl]cyclohexyl}carbamoyl)-3-methyl-L-valyl]-N-{(1S)-1-[(1R)-2-(cyclopropylamino)-1-hydroxy-2-oxoethyl]pentyl}-6,6-dimethyl-3-azabicyclo[3.1.0]hexane-2-carboxamide, 3C-like proteinase | | 著者 | Fu, L.F, Feng, Y, Qi, J.X. | | 登録日 | 2020-09-15 | | 公開日 | 2020-09-23 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (1.78 Å) | | 主引用文献 | Structural basis for the inhibition of the SARS-CoV-2 main protease by the anti-HCV drug narlaprevir.
Signal Transduct Target Ther, 6, 2021
|
|
5JDS
 
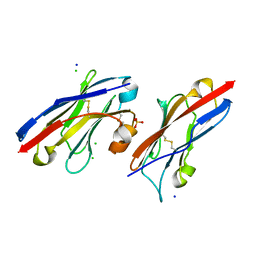 | |
5JDR
 
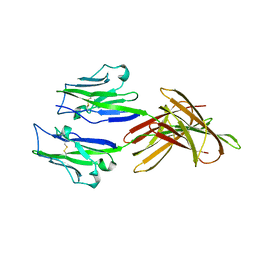 | | Structure of PD-L1 | | 分子名称: | Programmed cell death 1 ligand 1 | | 著者 | Zhou, A, Wei, H. | | 登録日 | 2016-04-17 | | 公開日 | 2017-04-12 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (2.7 Å) | | 主引用文献 | Structural basis of a novel PD-L1 nanobody for immune checkpoint blockade.
Cell Discov, 3, 2017
|
|
8HWK
 
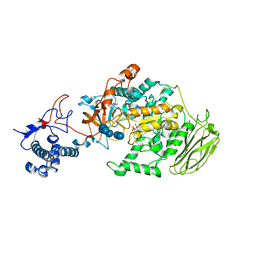 | | Limosilactobacillus reuteri N1 GtfB-maltohexaose | | 分子名称: | CITRIC ACID, DI(HYDROXYETHYL)ETHER, SODIUM ION, ... | | 著者 | Dong, J.J, Bai, Y.X. | | 登録日 | 2022-12-30 | | 公開日 | 2024-01-03 | | 最終更新日 | 2024-03-27 | | 実験手法 | X-RAY DIFFRACTION (2.9 Å) | | 主引用文献 | Insights into the Structure-Function Relationship of GH70 GtfB alpha-Glucanotransferases from the Crystal Structure and Molecular Dynamic Simulation of a Newly Characterized Limosilactobacillus reuteri N1 GtfB Enzyme.
J.Agric.Food Chem., 72, 2024
|
|
8HW3
 
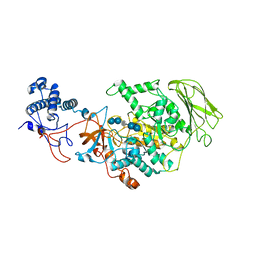 | | Limosilactobacillus reuteri N1 GtfB-acarbose | | 分子名称: | 4,6-dideoxy-4-{[(1S,4R,5S,6S)-4,5,6-trihydroxy-3-(hydroxymethyl)cyclohex-2-en-1-yl]amino}-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-beta-D-glucopyranose, GLYCEROL, SODIUM ION, ... | | 著者 | Dong, J.J, Bai, Y.X. | | 登録日 | 2022-12-28 | | 公開日 | 2024-01-03 | | 最終更新日 | 2024-03-27 | | 実験手法 | X-RAY DIFFRACTION (2.66 Å) | | 主引用文献 | Insights into the Structure-Function Relationship of GH70 GtfB alpha-Glucanotransferases from the Crystal Structure and Molecular Dynamic Simulation of a Newly Characterized Limosilactobacillus reuteri N1 GtfB Enzyme.
J.Agric.Food Chem., 72, 2024
|
|
8HU4
 
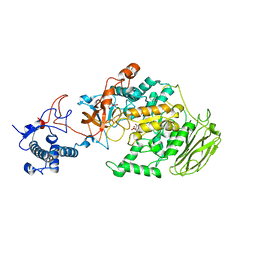 | | Limosilactobacillus reuteri N1 GtfB | | 分子名称: | CITRIC ACID, DI(HYDROXYETHYL)ETHER, SODIUM ION, ... | | 著者 | Dong, J.J, Bai, Y.X. | | 登録日 | 2022-12-22 | | 公開日 | 2023-12-27 | | 最終更新日 | 2024-03-27 | | 実験手法 | X-RAY DIFFRACTION (2.76 Å) | | 主引用文献 | Insights into the Structure-Function Relationship of GH70 GtfB alpha-Glucanotransferases from the Crystal Structure and Molecular Dynamic Simulation of a Newly Characterized Limosilactobacillus reuteri N1 GtfB Enzyme.
J.Agric.Food Chem., 72, 2024
|
|
6E0P
 
 | |
6DZT
 
 | |
6E0C
 
 | |
8WMN
 
 | |
8WMM
 
 | |
8WMH
 
 | |
8WR4
 
 | |
4WYU
 
 | |
4WYT
 
 | |
4X23
 
 | |
4QLC
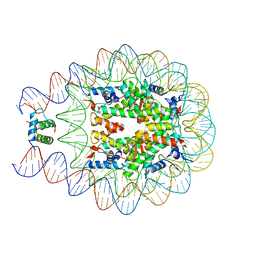 
 | | Crystal structure of chromatosome at 3.5 angstrom resolution | | 分子名称: | CITRIC ACID, DNA (167-mer), H5, ... | | 著者 | Jiang, J.S, Zhou, B.R, Xiao, T.S, Bai, Y.W. | | 登録日 | 2014-06-11 | | 公開日 | 2015-07-22 | | 最終更新日 | 2023-09-20 | | 実験手法 | X-RAY DIFFRACTION (3.503 Å) | | 主引用文献 | Structural Mechanisms of Nucleosome Recognition by Linker Histones.
Mol.Cell, 33 Suppl 1, 2015
|
|
