2VPV
 
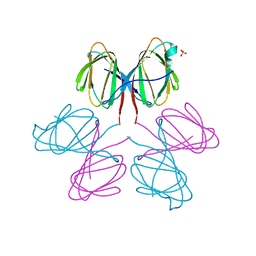 | | Dimerization Domain of Mif2p | | 分子名称: | PROTEIN MIF2, SULFATE ION | | 著者 | Cohen, R.L, Espelin, C.W, Sorger, P.K, Harrison, S.C, Simons, K.T. | | 登録日 | 2008-03-05 | | 公開日 | 2008-08-26 | | 最終更新日 | 2024-11-06 | | 実験手法 | X-RAY DIFFRACTION (2.7 Å) | | 主引用文献 | Structural and Functional Dissection of Mif2P, a Conserved DNA-Binding Kinetochore Protein.
Mol.Biol.Cell, 19, 2008
|
|
2YFV
 
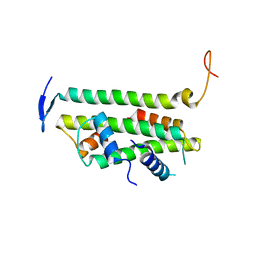 | |
5EN2
 
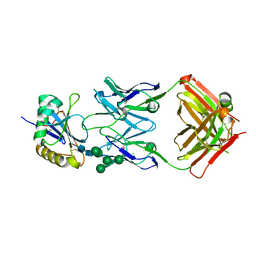 | | Molecular basis for antibody-mediated neutralization of New World hemorrhagic fever mammarenaviruses | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CHLORIDE ION, ... | | 著者 | Mahmutovic, S, Clark, L, Levis, S, Briggiler, A, Enria, D, Harrison, S.C, Abraham, J. | | 登録日 | 2015-11-09 | | 公開日 | 2015-12-30 | | 最終更新日 | 2024-11-20 | | 実験手法 | X-RAY DIFFRACTION (1.821 Å) | | 主引用文献 | Molecular Basis for Antibody-Mediated Neutralization of New World Hemorrhagic Fever Mammarenaviruses.
Cell Host Microbe, 18, 2015
|
|
4XI2
 
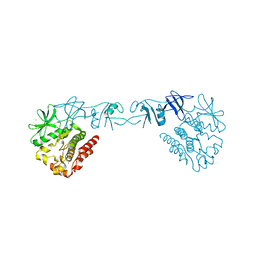 | |
4XDN
 | |
5IBU
 
 | | 6652 Fab (unbound) | | 分子名称: | 6652 Heavy Chain, 6652 Light Chain | | 著者 | Raymond, D.D, Harrison, S.C. | | 登録日 | 2016-02-22 | | 公開日 | 2016-11-09 | | 最終更新日 | 2024-11-13 | | 実験手法 | X-RAY DIFFRACTION (1.71 Å) | | 主引用文献 | Influenza immunization elicits antibodies specific for an egg-adapted vaccine strain.
Nat. Med., 22, 2016
|
|
5IBT
 
 | |
5IBL
 | |
3IYV
 | | Clathrin D6 coat as full-length Triskelions | | 分子名称: | Clathrin heavy chain, Clathrin light chain A | | 著者 | Johnson, G.T, Fotin, A, Cheng, Y, Sliz, P, Grigorieff, N, Harrison, S.C, Kirchhausen, T, Walz, T. | | 登録日 | 2010-06-17 | | 公開日 | 2010-07-21 | | 最終更新日 | 2024-02-21 | | 実験手法 | ELECTRON MICROSCOPY (7.9 Å) | | 主引用文献 | Molecular model for a complete clathrin lattice from electron cryomicroscopy.
Nature, 432, 2004
|
|
4YJZ
 
 | |
4YK4
 
 | |
8T0P
 
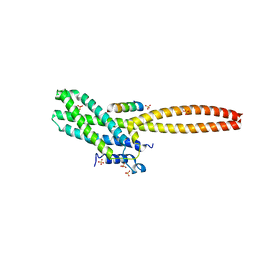 | | Structure of Cse4 bound to Ame1 and Okp1 | | 分子名称: | Histone H3-like centromeric protein CSE4, Inner kinetochore subunit AME1, Inner kinetochore subunit OKP1, ... | | 著者 | Deng, S, Harrison, S.C. | | 登録日 | 2023-06-01 | | 公開日 | 2023-09-27 | | 最終更新日 | 2023-12-27 | | 実験手法 | X-RAY DIFFRACTION (1.73 Å) | | 主引用文献 | Recognition of centromere-specific histone Cse4 by the inner kinetochore Okp1-Ame1 complex.
Embo Rep., 24, 2023
|
|
8UDG
 
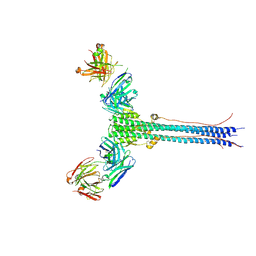 | | S1V2-72 Fab bound to EHA2 from influenza B/Malaysia/2506/2004 | | 分子名称: | Hemagglutinin, S1V2-72 heavy chain, S1V2-72 light chain | | 著者 | Finney, J, Kong, S, Walsh Jr, R.M, Harrison, S.C, Kelsoe, G. | | 登録日 | 2023-09-28 | | 公開日 | 2023-11-15 | | 最終更新日 | 2024-10-16 | | 実験手法 | ELECTRON MICROSCOPY (4.98 Å) | | 主引用文献 | Protective human antibodies against a conserved epitope in pre- and postfusion influenza hemagglutinin.
Proc.Natl.Acad.Sci.USA, 121, 2024
|
|
8UK3
 | | The rotavirus VP5*/VP8* conformational transition permeabilizes membranes to Ca2+ (class 6 reconstruction) | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, Outer capsid glycoprotein VP7, ... | | 著者 | De Sautu, M, Herrmann, T, Jenni, S, Harrison, S.C. | | 登録日 | 2023-10-12 | | 公開日 | 2024-03-27 | | 最終更新日 | 2024-10-23 | | 実験手法 | ELECTRON MICROSCOPY (8 Å) | | 主引用文献 | The rotavirus VP5*/VP8* conformational transition permeabilizes membranes to Ca2.
Plos Pathog., 20, 2024
|
|
8UK2
 | | The rotavirus VP5*/VP8* conformational transition permeabilizes membranes to Ca2+ (class 5 reconstruction) | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, Outer capsid glycoprotein VP7, ... | | 著者 | De Sautu, M, Herrmann, T, Jenni, S, Harrison, S.C. | | 登録日 | 2023-10-12 | | 公開日 | 2024-03-27 | | 最終更新日 | 2024-10-23 | | 実験手法 | ELECTRON MICROSCOPY (8 Å) | | 主引用文献 | The rotavirus VP5*/VP8* conformational transition permeabilizes membranes to Ca2.
Plos Pathog., 20, 2024
|
|
6XPQ
 
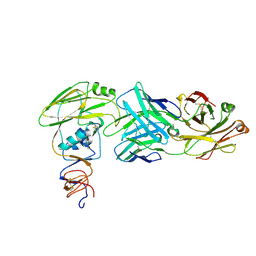 | |
6XPX
 
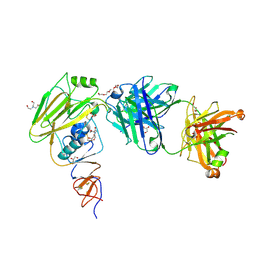 | |
6XPY
 
 | |
6XQ4
 
 | |
6XPR
 
 | | Human antibody D2 H1-1/H3-1 H3 in complex with the influenza hemagglutinin head domain of A/Texas/50/2012(H3N2) | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Hemagglutinin, ... | | 著者 | McCarthy, K.R, Harrison, S.C, Lee, J. | | 登録日 | 2020-07-08 | | 公開日 | 2021-05-19 | | 最終更新日 | 2024-11-06 | | 実験手法 | X-RAY DIFFRACTION (4.092 Å) | | 主引用文献 | A Prevalent Focused Human Antibody Response to the Influenza Virus Hemagglutinin Head Interface.
Mbio, 12, 2021
|
|
6XPO
 
 | | Influenza hemagglutinin A/Bangkok/01/1979(H3N2) | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Hemagglutinin, ... | | 著者 | McCarthy, K.R, Harrison, S.C. | | 登録日 | 2020-07-08 | | 公開日 | 2021-05-19 | | 最終更新日 | 2024-10-23 | | 実験手法 | X-RAY DIFFRACTION (3 Å) | | 主引用文献 | A Prevalent Focused Human Antibody Response to the Influenza Virus Hemagglutinin Head Interface.
Mbio, 12, 2021
|
|
6XPZ
 
 | |
6XQ0
 | |
6XQ2
 
 | |
6WXG
 | | Cryo-EM reconstruction of VP5*/VP8* assembly from rhesus rotavirus particles - Reversed conformation | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, Intermediate capsid protein VP6, ... | | 著者 | Herrmann, T, Harrison, S.C, Jenni, S. | | 登録日 | 2020-05-10 | | 公開日 | 2021-01-20 | | 最終更新日 | 2024-11-13 | | 実験手法 | ELECTRON MICROSCOPY (3.3 Å) | | 主引用文献 | Functional refolding of the penetration protein on a non-enveloped virus.
Nature, 590, 2021
|
|
