8TO6
 
 | | Escherichia coli RNA polymerase unwinding intermediate (I1d) at the lambda PR promoter | | 分子名称: | (3R,5S,7R,8R,9S,10S,12S,13R,14S,17R)-10,13-dimethyl-17-[(2R)-pentan-2-yl]-2,3,4,5,6,7,8,9,11,12,14,15,16,17-tetradecahydro-1H-cyclopenta[a]phenanthrene-3,7,12-triol, DNA-directed RNA polymerase subunit alpha, DNA-directed RNA polymerase subunit beta, ... | | 著者 | Darst, S.A, Saecker, R.M, Mueller, A.U. | | 登録日 | 2023-08-02 | | 公開日 | 2024-07-03 | | 最終更新日 | 2024-07-17 | | 実験手法 | ELECTRON MICROSCOPY (2.9 Å) | | 主引用文献 | Early intermediates in bacterial RNA polymerase promoter melting visualized by time-resolved cryo-electron microscopy.
Nat.Struct.Mol.Biol., 2024
|
|
8TOM
 
 | |
8TO8
 
 | | Escherichia coli RNA polymerase unwinding intermediate (I1b) at the lambda PR promoter | | 分子名称: | (3R,5S,7R,8R,9S,10S,12S,13R,14S,17R)-10,13-dimethyl-17-[(2R)-pentan-2-yl]-2,3,4,5,6,7,8,9,11,12,14,15,16,17-tetradecahydro-1H-cyclopenta[a]phenanthrene-3,7,12-triol, DNA-directed RNA polymerase subunit alpha, DNA-directed RNA polymerase subunit beta, ... | | 著者 | Darst, S.A, Saecker, R.M, Mueller, A.U. | | 登録日 | 2023-08-03 | | 公開日 | 2024-07-03 | | 最終更新日 | 2024-07-17 | | 実験手法 | ELECTRON MICROSCOPY (2.9 Å) | | 主引用文献 | Early intermediates in bacterial RNA polymerase promoter melting visualized by time-resolved cryo-electron microscopy.
Nat.Struct.Mol.Biol., 2024
|
|
8TO1
 
 | | Escherichia coli RNA polymerase unwinding intermediate (I1a) at the lambda PR promoter | | 分子名称: | (3R,5S,7R,8R,9S,10S,12S,13R,14S,17R)-10,13-dimethyl-17-[(2R)-pentan-2-yl]-2,3,4,5,6,7,8,9,11,12,14,15,16,17-tetradecahydro-1H-cyclopenta[a]phenanthrene-3,7,12-triol, DNA-directed RNA polymerase subunit alpha, DNA-directed RNA polymerase subunit beta, ... | | 著者 | Darst, S.A, Saecker, R.M, Mueller, A.U. | | 登録日 | 2023-08-02 | | 公開日 | 2024-07-03 | | 最終更新日 | 2024-07-17 | | 実験手法 | ELECTRON MICROSCOPY (2.8 Å) | | 主引用文献 | Early intermediates in bacterial RNA polymerase promoter melting visualized by time-resolved cryo-electron microscopy.
Nat.Struct.Mol.Biol., 2024
|
|
6KVH
 
 | | The mutant crystal structure of endo-polygalacturonase (T284A) from Talaromyces leycettanus JCM 12802 | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, alpha-D-mannopyranose, endo-polygalacturonase | | 著者 | Tu, T, Wang, Z, Luo, H, Yao, B. | | 登録日 | 2019-09-04 | | 公開日 | 2020-09-09 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (1.2 Å) | | 主引用文献 | Structural Insights into the Mechanisms Underlying the Kinetic Stability of GH28 Endo-Polygalacturonase.
J.Agric.Food Chem., 69, 2021
|
|
3MQ2
 
 | | Crystal Structure of 16S rRNA Methyltranferase KamB | | 分子名称: | 16S rRNA methyltransferase, S-ADENOSYL-L-HOMOCYSTEINE | | 著者 | Macmaster, R.A. | | 登録日 | 2010-04-27 | | 公開日 | 2010-12-08 | | 最終更新日 | 2024-02-21 | | 実験手法 | X-RAY DIFFRACTION (1.69 Å) | | 主引用文献 | Structural insights into the function of aminoglycoside-resistance A1408 16S rRNA methyltransferases from antibiotic-producing and human pathogenic bacteria.
Nucleic Acids Res., 38, 2010
|
|
6S8Z
 
 | |
4G55
 
 | | Clathrin terminal domain complexed with pitstop 2 | | 分子名称: | 1,2-ETHANEDIOL, ACETATE ION, Clathrin heavy chain 1, ... | | 著者 | Bulut, H, Von Kleist, L, Saenger, W, Haucke, V. | | 登録日 | 2012-07-17 | | 公開日 | 2012-08-01 | | 最終更新日 | 2024-02-28 | | 実験手法 | X-RAY DIFFRACTION (1.69 Å) | | 主引用文献 | Role of the clathrin terminal domain in regulating coated pit dynamics revealed by small molecule inhibition.
Cell(Cambridge,Mass.), 146, 2011
|
|
7WVR
 
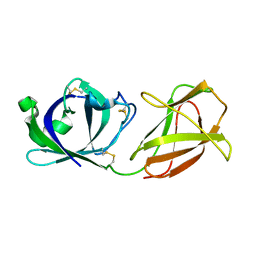 | |
7JVO
 
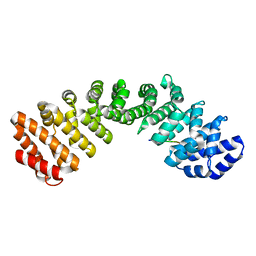 | |
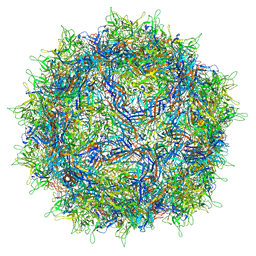
7NA6
 
 | |
7Z9P
 
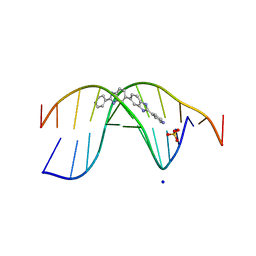 | | The novel DNA binding mechanism of ridinilazole, a precision Clostridiodes difficile antibiotic | | 分子名称: | DNA (5'-D(P*GP*CP*GP*AP*AP*TP*TP*CP*GP*CP*G*(RID))-3'), Ridinilazole, SODIUM ION, ... | | 著者 | Mason, S, Leonard, P.M. | | 登録日 | 2022-03-21 | | 公開日 | 2023-03-29 | | 最終更新日 | 2024-02-07 | | 実験手法 | X-RAY DIFFRACTION (2.2 Å) | | 主引用文献 | The Novel DNA Binding Mechanism of Ridinilazole, a Precision Clostridiodes difficile Antibiotic.
Antimicrob.Agents Chemother., 67, 2023
|
|
7SQ1
 
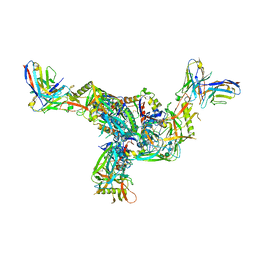 | | BG505.MD39TS Env trimer in complex with Fab from antibody C05 | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, C05 Fab Light chain, ... | | 著者 | Moore, A, Du, J, Xu, Z, Walker, S, Kulp, D.W, Pallesen, J. | | 登録日 | 2021-11-04 | | 公開日 | 2022-06-22 | | 最終更新日 | 2022-12-07 | | 実験手法 | ELECTRON MICROSCOPY (3.8 Å) | | 主引用文献 | Induction of tier-2 neutralizing antibodies in mice with a DNA-encoded HIV envelope native like trimer.
Nat Commun, 13, 2022
|
|
2RTL
 
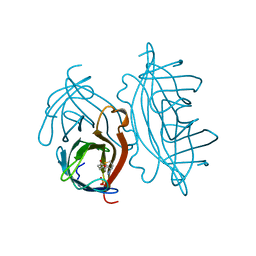 | |
2RTI
 
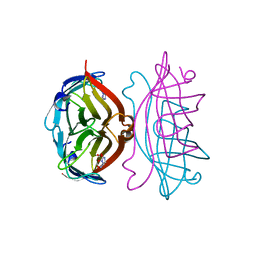 | |
2RTD
 
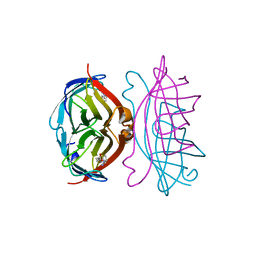 | |
2RTR
 
 | |
2RTO
 
 | |
2RTG
 
 | |
2RTQ
 
 | |
2RTC
 
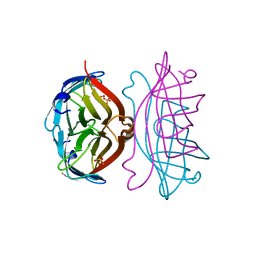 | | APOSTREPTAVIDIN, PH 3.60, SPACE GROUP I222 | | 分子名称: | STREPTAVIDIN, SULFATE ION | | 著者 | Katz, B.A. | | 登録日 | 1997-09-11 | | 公開日 | 1998-10-14 | | 最終更新日 | 2024-02-21 | | 実験手法 | X-RAY DIFFRACTION (1.5 Å) | | 主引用文献 | Binding of biotin to streptavidin stabilizes intersubunit salt bridges between Asp61 and His87 at low pH.
J.Mol.Biol., 274, 1997
|
|
2RTN
 
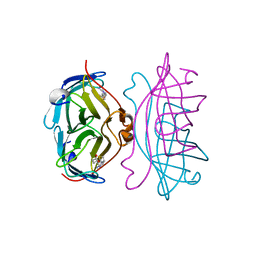 | |
2RTA
 
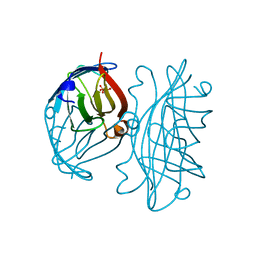 | | APOSTREPTAVIDIN, PH 2.97, SPACE GROUP I4122 | | 分子名称: | STREPTAVIDIN, SULFATE ION | | 著者 | Katz, B.A. | | 登録日 | 1997-09-11 | | 公開日 | 1998-10-14 | | 最終更新日 | 2024-02-21 | | 実験手法 | X-RAY DIFFRACTION (1.39 Å) | | 主引用文献 | Binding of biotin to streptavidin stabilizes intersubunit salt bridges between Asp61 and His87 at low pH.
J.Mol.Biol., 274, 1997
|
|
3QSY
 
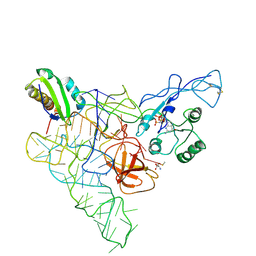 | | Recognition of the methionylated initiator tRNA by the translation initiation factor 2 in Archaea | | 分子名称: | METHIONINE, PHOSPHOAMINOPHOSPHONIC ACID-GUANYLATE ESTER, Translation initiation factor 2 subunit alpha, ... | | 著者 | Nikonov, O.S, Stolboushkina, E.A, Zelinskaya, N.V, Nikulin, A.D, Garber, M.B, Nikonov, S.V. | | 登録日 | 2011-02-22 | | 公開日 | 2012-03-21 | | 最終更新日 | 2023-11-01 | | 実験手法 | X-RAY DIFFRACTION (3.2 Å) | | 主引用文献 | Crystal structure of the archaeal translation initiation factor 2 in complex with a GTP analogue and Met-tRNAf(Met.)
J.Mol.Biol., 425, 2013
|
|
5N8Y
 
 | | KaiCBA circadian clock backbone model based on a Cryo-EM density | | 分子名称: | Circadian clock protein KaiA, Circadian clock protein KaiB, Circadian clock protein kinase KaiC | | 著者 | Schuller, J.M, Snijder, J, Loessl, P, Heck, A.J.R, Foerster, F. | | 登録日 | 2017-02-24 | | 公開日 | 2017-03-29 | | 最終更新日 | 2024-05-15 | | 実験手法 | ELECTRON MICROSCOPY (4.7 Å) | | 主引用文献 | Structures of the cyanobacterial circadian oscillator frozen in a fully assembled state.
Science, 355, 2017
|
|
