1NZA
 
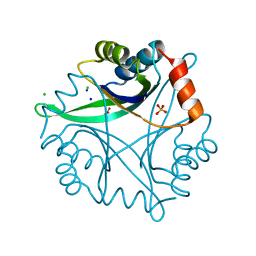 | | Divalent cation tolerance protein (Cut A1) from thermus thermophilus HB8 | | Descriptor: | CHLORIDE ION, Divalent cation tolerance protein, GLYCEROL, ... | | Authors: | Bagautdinov, B, Miyano, M, Tahirov, T.H, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2003-02-17 | | Release date: | 2003-03-11 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | The structures of the CutA1 proteins from Thermus thermophilus and Pyrococcus horikoshii: characterization of metal-binding sites and metal-induced assembly.
Acta Crystallogr.,Sect.F, 70, 2014
|
|
1J3N
 
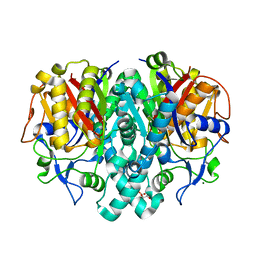 | | Crystal Structure of 3-oxoacyl-(acyl-carrier protein) Synthase II from Thermus thermophilus HB8 | | Descriptor: | 3-oxoacyl-(acyl-carrier protein) synthase II, CITRIC ACID, MAGNESIUM ION | | Authors: | Bagautdinov, B, Miyano, M, Tahirov, T.H, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2003-02-10 | | Release date: | 2003-03-11 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure of 3-oxoacyl-(acyl-carrier protein) synthase II from Thermus thermophilus HB8.
Acta Crystallogr.,Sect.F, 64, 2008
|
|
2ZGW
 
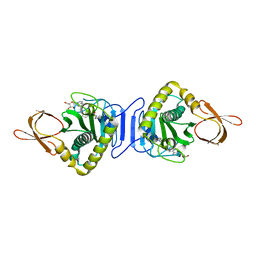 | | Crystal Structure Of Biotin Protein Ligase From Pyrococcus Horikoshii Complexed with Adenosine and Biotin, Mutations R48A and K111A | | Descriptor: | ADENOSINE, BIOTIN, biotin--[acetyl-CoA-carboxylase] ligase | | Authors: | Bagautdinov, B, Matsuura, Y, Bagautdinova, S, Kunishima, N, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2008-01-28 | | Release date: | 2008-02-05 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Protein biotinylation visualized by a complex structure of biotin protein ligase with a substrate
J.Biol.Chem., 283, 2008
|
|
4NYO
 
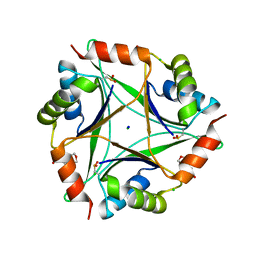 | | The 1.8 Angstrom Crystal Structure of the Periplasmic Divalent Cation Tolerance Protein Cuta from Pyrococcus Horikoshii OT3 | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, CHLORIDE ION, Divalent-cation tolerance protein CutA, ... | | Authors: | Bagautdinov, B, Tahirov, T.H, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2013-12-11 | | Release date: | 2014-01-01 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The structures of the CutA1 proteins from Thermus thermophilus and Pyrococcus horikoshii: characterization of metal-binding sites and metal-induced assembly
ACTA CRYSTALLOGR.,SECT.F, 70, 2014
|
|
4NYP
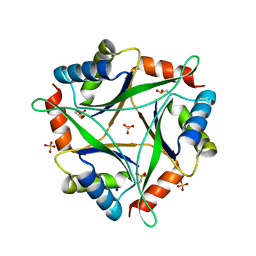 
 | |
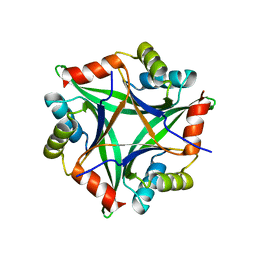
2ZFH
 
 | |
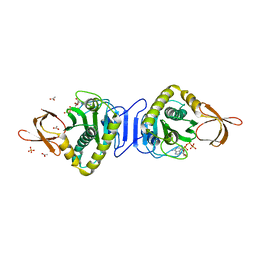
1X01
 
 | |
1UFV
 
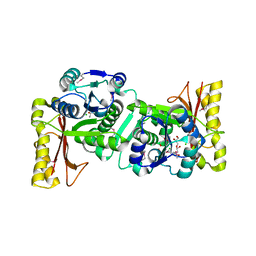 | | Crystal Structure Of Pantothenate Synthetase From Thermus Thermophilus HB8 | | Descriptor: | CHLORIDE ION, GLYCEROL, Pantoate-beta-alanine ligase | | Authors: | Bagautdinov, B, Kuramitsu, S, Yokoyama, S, Miyano, M, Tahirov, T.H, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2003-06-10 | | Release date: | 2003-06-24 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Crystal Structure Of Pantothenate Synthetase From Thermus Thermophilus HB8
To be Published
|
|
1UB0
 
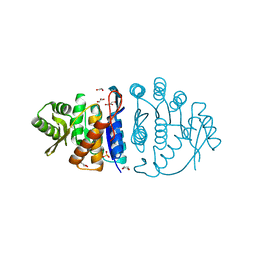 | | Crystal Structure Analysis of Phosphomethylpyrimidine Kinase (ThiD) from Thermus Thermophilus Hb8 | | Descriptor: | 1,4-DIETHYLENE DIOXIDE, GLYCEROL, Phosphomethylpyrimidine Kinase, ... | | Authors: | Bagautdinov, B, Kuramitsu, S, Yokoyama, S, Miyano, M, Tahirov, T.H, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2003-03-26 | | Release date: | 2003-04-08 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Crystal Structure Analysis of Phosphomethylpyrimidine Kinase (ThiD) from Thermus Thermophilus Hb8
To be Published
|
|
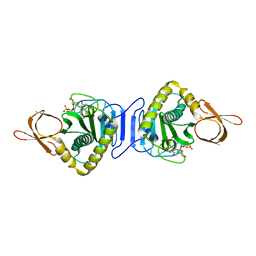
1WNL
 
 | |
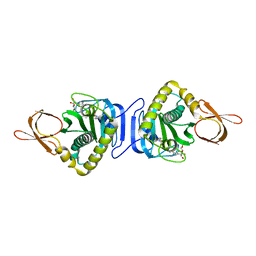
1WQW
 
 | |
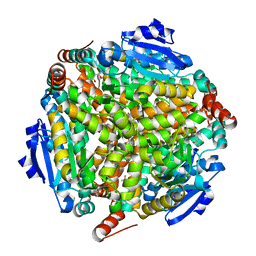
1WZ8
 
 | |
1WQ7
 
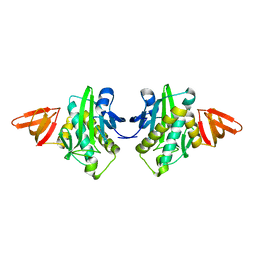 | |
1VC4
 
 | | Crystal Structure of Indole-3-Glycerol Phosphate Synthase (TrpC) from Thermus Thermophilus At 1.8 A Resolution | | Descriptor: | ACETIC ACID, GLYCEROL, Indole-3-Glycerol Phosphate Synthase, ... | | Authors: | Bagautdinov, B, Tahirov, T.H, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2004-03-04 | | Release date: | 2004-03-23 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structure of indole-3-glycerol phosphate synthase from Thermus thermophilus HB8: implications for thermal stability.
Acta Crystallogr.,Sect.D, 67, 2011
|
|
1WXD
 
 | |
1WPY
 
 | |
1V4V
 
 | |
1V8F
 
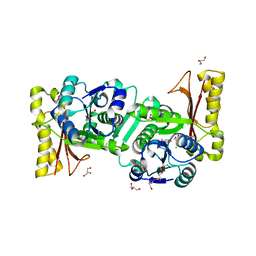 | |
1V6H
 
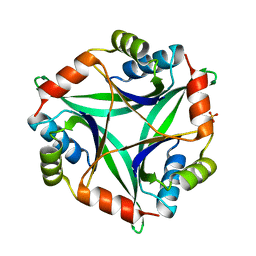 | |
2E41
 
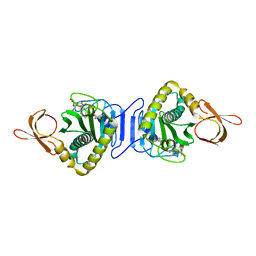 | | Crystal Structure Of Biotin Protein Ligase From Pyrococcus Horikoshii Complexed with the Reaction Product Analog Biotinol-5'-AMP, Mutations R48A and K111A | | Descriptor: | ((2R,3S,4R,5R)-5-(6-AMINO-9H-PURIN-9-YL)-3,4-DIHYDROXY-TETRAHYDROFURAN-2-YL)METHYL 5-((3AS,4S,6AR)-2-OXO-HEXAHYDRO-1H-THIENO[3,4-D]IMIDAZOL-4-YL)PENTYL HYDROGEN PHOSPHATE, biotin--[acetyl-CoA-carboxylase] ligase | | Authors: | Bagautdinov, B, Matsuura, Y, Bagautdinova, S, Kunishima, N, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2006-12-01 | | Release date: | 2007-06-05 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Protein biotinylation visualized by a complex structure of biotin protein ligase with a substrate
J.Biol.Chem., 283, 2008
|
|
2EJG
 
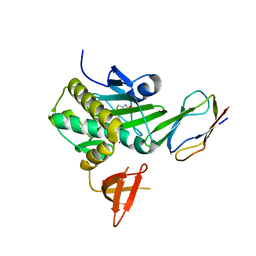 | | Crystal Structure Of The Biotin Protein Ligase (Mutation R48A) and Biotin Carboxyl Carrier Protein Complex From Pyrococcus Horikoshii OT3 | | Descriptor: | 149aa long hypothetical methylmalonyl-CoA decarboxylase gamma chain, 235aa long hypothetical biotin--[acetyl-CoA-carboxylase] ligase, ADENOSINE, ... | | Authors: | Bagautdinov, B, Matsuura, Y, Bagautdinova, S, Kunishima, N, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2007-03-16 | | Release date: | 2008-03-18 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.71 Å) | | Cite: | Protein biotinylation visualized by a complex structure of biotin protein ligase with a substrate
J.Biol.Chem., 283, 2008
|
|
2EJF
 
 | | Crystal Structure Of The Biotin Protein Ligase (Mutations R48A and K111A) and Biotin Carboxyl Carrier Protein Complex From Pyrococcus Horikoshii OT3 | | Descriptor: | 149aa long hypothetical methylmalonyl-CoA decarboxylase gamma chain, 235aa long hypothetical biotin--[acetyl-CoA-carboxylase] ligase, ADENOSINE, ... | | Authors: | Bagautdinov, B, Matsuura, Y, Bagautdinova, S, Kunishima, N, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2007-03-16 | | Release date: | 2008-03-18 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Protein biotinylation visualized by a complex structure of biotin protein ligase with a substrate
J.Biol.Chem., 283, 2008
|
|
2EJ9
 
 | |
2E66
 
 | | Crystal Structure Of CutA1 From Pyrococcus Horikoshii OT3, Mutation D60A | | Descriptor: | CHLORIDE ION, Divalent-cation tolerance protein cutA, SODIUM ION | | Authors: | Bagautdinov, B, Sawano, M, Bagautdinova, S, Yutani, K, Kunishima, N, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2006-12-25 | | Release date: | 2007-06-26 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural basis of the hyper-thermostability of CutA1
To be Published
|
|
2DXU
 
 | | Crystal Structure Of Biotin Protein Ligase From Pyrococcus Horikoshii Complexed with Biotinyl-5'-AMP, Mutation R48A | | Descriptor: | BIOTINYL-5-AMP, biotin--[acetyl-CoA-carboxylase] ligase | | Authors: | Bagautdinov, B, Taketa, M, Matsuura, Y, Kunishima, N, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2006-08-30 | | Release date: | 2007-03-01 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.28 Å) | | Cite: | Protein biotinylation visualized by a complex structure of biotin protein ligase with a substrate
J.Biol.Chem., 283, 2008
|
|
