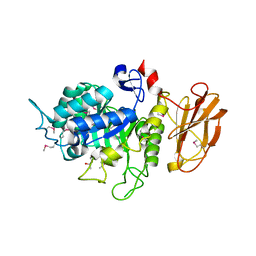
4QFU
 
 | |
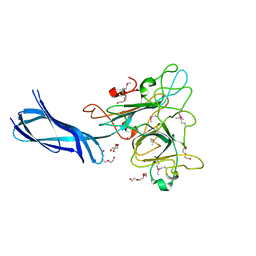
4Q1Z
 
 | |
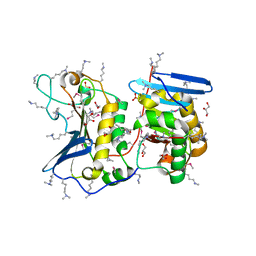
4QE0
 
 | |
4QHB
 
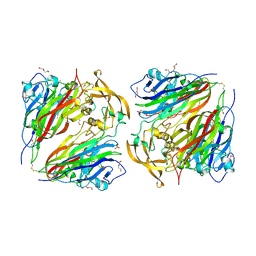 | |
4QNI
 
 | |
4QVU
 
 | |
4R7F
 
 | |
4QRL
 
 | |
4R1M
 
 | |
4R1L
 
 | |
4R7S
 
 | |
4RWV
 
 | | Crystal structure of PIP3 bound human nuclear receptor LRH-1 (Liver Receptor Homolog 1, NR5A2) in complex with a co-regulator DAX-1 (NR0B1) peptide at 1.86 A resolution | | Descriptor: | (2S)-3-{[(R)-{[(1S,2S,3R,4S,5S,6S)-2,6-dihydroxy-3,4,5-tris(phosphonooxy)cyclohexyl]oxy}(hydroxy)phosphoryl]oxy}propane -1,2-diyl dihexadecanoate, 1,2-ETHANEDIOL, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, ... | | Authors: | Joint Center for Structural Genomics (JCSG), Partnership for Stem Cell Biology, Partnership for Stem Cell Biology (STEMCELL) | | Deposit date: | 2014-12-05 | | Release date: | 2015-01-21 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (1.859 Å) | | Cite: | Crystal structure of a Homo sapiens hepatocytic transcription factor hB1F-2 (B1F2) in complex with nuclear receptor subfamily 0 group B member 1 (NR0B1, residues 140-154) from human at 1.86 A resolution
To be published
|
|
4RVO
 
 | |
4RU2
 
 | |
4RVN
 
 | |
4PWU
 
 | |
4Q5T
 
 | |
4Q69
 
 | |
4Q1V
 
 | |
4Q34
 
 | |
4QHZ
 | |
4QAN
 
 | |
4QGP
 
 | |
4QOA
 
 | |
4QPV
 
 | |
