1X01
 
 | |
1X45
 
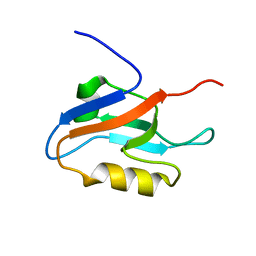 | | Solution structure of the first PDZ domain of amyloid beta A4 precursor protein-binding family A, member 1 | | Descriptor: | amyloid beta (A4) precursor protein-binding, family A, member 1 (X11) | | Authors: | Qin, X.R, Hayashi, F, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2005-05-13 | | Release date: | 2005-11-13 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the first PDZ domain of amyloid beta A4 precursor protein-binding family A, member 1
to be published
|
|
1X4M
 
 | | Solution structure of KH domain in Far upstream element binding protein 1 | | Descriptor: | Far upstream element binding protein 1 | | Authors: | He, F, Muto, Y, Inoue, M, Kigawa, T, Shirouzu, M, Terada, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2005-05-14 | | Release date: | 2005-11-14 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of KH domain in Far upstream element binding protein 1
To be Published
|
|
1X54
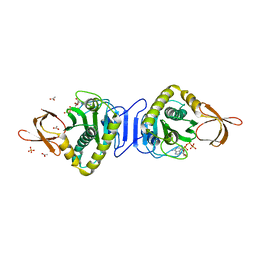
 | | Crystal structure of asparaginyl-tRNA synthetase from Pyrococcus horikoshii complexed with asparaginyl-adenylate | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 4-AMINO-1,4-DIOXOBUTAN-2-AMINIUM ADENOSINE-5'-MONOPHOSPHATE, Asparaginyl-tRNA synthetase, ... | | Authors: | Iwasaki, W, Sekine, S, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2005-05-15 | | Release date: | 2006-05-23 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Structural Basis of the Water-assisted Asparagine Recognition by Asparaginyl-tRNA Synthetase.
J.Mol.Biol., 360, 2006
|
|
1WEO
 
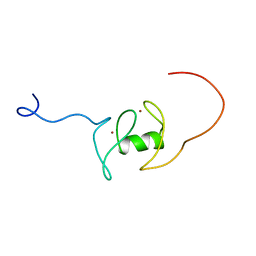 | | Solution structure of RING-finger in the catalytic subunit (IRX3) of cellulose synthase | | Descriptor: | ZINC ION, cellulose synthase, catalytic subunit (IRX3) | | Authors: | He, F, Muto, Y, Inoue, M, Kigawa, T, Shirouzu, M, Terada, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2004-05-25 | | Release date: | 2004-11-25 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of RING-finger in the catalytic subunit (IRX3) of cellulose synthase
To be Published
|
|
1X5I
 
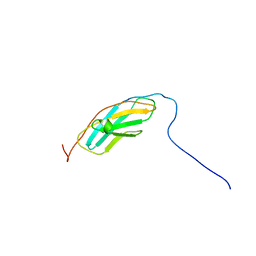 | | The solution structure of the fourth fibronectin type III domain of human Neogenin | | Descriptor: | Neogenin | | Authors: | Tochio, N, Koshiba, S, Inoue, M, Kigawa, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2005-05-15 | | Release date: | 2005-11-15 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | The solution structure of the fourth fibronectin type III domain of human Neogenin
To be Published
|
|
1X5Y
 
 | | Solution structure of the fibronectin type-III domain of mouse myosin-binding protein C, Fast-type homolog | | Descriptor: | myosin binding protein C, fast-type | | Authors: | Yoneyama, M, Tochio, N, Koshiba, S, Inoue, M, Kigawa, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2005-05-17 | | Release date: | 2005-11-17 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the fibronectin type-III domain of mouse myosin-binding protein C, Fast-type homolog
To be Published
|
|
1WG1
 
 | | Solution structure of RNA binding domain in BAB13405(homolog EXC-7) | | Descriptor: | KIAA1579 protein | | Authors: | Tsuda, K, Muto, Y, Nagata, T, Suzuki, S, Someya, T, Kigawa, T, Terada, T, Shirouzu, M, Inoue, M, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2004-05-27 | | Release date: | 2004-11-27 | | Last modified: | 2024-10-09 | | Method: | SOLUTION NMR | | Cite: | Solution structure of RNA binding domain in BAB13405(homolog EXC-7)
To be Published
|
|
2ZIN
 
 | | Crystal structure of the catalytic domain of pyrrolysyl-tRNA synthetase in complex with BocLys and an ATP analogue | | Descriptor: | 1,2-ETHANEDIOL, MAGNESIUM ION, N~6~-(tert-butoxycarbonyl)-L-lysine, ... | | Authors: | Yanagisawa, T, Ishii, R, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2008-02-19 | | Release date: | 2008-12-02 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | Multistep Engineering of Pyrrolysyl-tRNA Synthetase to Genetically Encode N(varepsilon)-(o-Azidobenzyloxycarbonyl) lysine for Site-Specific Protein Modification
Chem.Biol., 15, 2008
|
|
2ZKT
 
 | |
1WP4
 
 | |
1X3H
 
 | | Solution structure of the LIM domain of human Leupaxin | | Descriptor: | Leupaxin, ZINC ION | | Authors: | Yoneyama, M, Tochio, N, Koshiba, S, Inoue, M, Kigawa, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2005-05-09 | | Release date: | 2005-11-09 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the LIM domain of human Leupaxin
To be Published
|
|
1X4E
 
 | | Solution structure of RRM domain in RNA binding motif, single-stranded interacting protein 2 | | Descriptor: | RNA binding motif, single-stranded interacting protein 2 | | Authors: | He, F, Muto, Y, Inoue, M, Kigawa, T, Shirouzu, M, Terada, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2005-05-14 | | Release date: | 2005-11-14 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of RRM domain in RNA binding motif, single-stranded interacting protein 2
To be Published
|
|
1X4W
 
 | | Solution structure of the zf-AN1 domain from human hypothetical protein FLJ13222 | | Descriptor: | Hypothetical protein FLJ13222, ZINC ION | | Authors: | Tomizawa, T, Kigawa, T, Koshiba, S, Inoue, M, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2005-05-15 | | Release date: | 2005-11-15 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the zf-AN1 domain from human hypothetical protein FLJ13222
To be Published
|
|
1X5M
 
 | |
1WYP
 
 | | Solution structure of the CH domain of human Calponin 1 | | Descriptor: | Calponin 1 | | Authors: | Tomizawa, T, Kigawa, T, Koshiba, S, Inoue, M, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2005-02-15 | | Release date: | 2005-08-15 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the CH domain of human Calponin 1
To be Published
|
|
1WFI
 
 | |
1X0J
 
 | | Crystal structure analysis of the N-terminal bromodomain of human Brd2 | | Descriptor: | 2,3-DIHYDROXY-1,4-DITHIOBUTANE, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, Bromodomain-containing protein 2 | | Authors: | Nakamura, Y, Umehara, T, Shirouzu, M, Padmanabhan, B, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2005-03-23 | | Release date: | 2006-06-27 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural basis for acetylated histone H4 recognition by the human Brd2 bromodomain
To be Published
|
|
2ZTG
 
 | | Crystal structure of Archaeoglobus fulgidus alanyl-tRNA synthetase lacking the C-terminal dimerization domain in complex with Ala-SA | | Descriptor: | '5'-O-(N-(L-ALANYL)-SULFAMOYL)ADENOSINE, Alanyl-tRNA synthetase, ZINC ION | | Authors: | Naganuma, M, Sekine, S, Fukunaga, R, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2008-10-01 | | Release date: | 2009-06-30 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Unique protein architecture of alanyl-tRNA synthetase for aminoacylation, editing, and dimerization.
Proc.Natl.Acad.Sci.USA, 106, 2009
|
|
1WJ2
 
 | | Solution Structure of the C-terminal WRKY Domain of AtWRKY4 | | Descriptor: | Probable WRKY transcription factor 4, ZINC ION | | Authors: | Yamasaki, K, Inoue, M, Kigawa, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2004-05-28 | | Release date: | 2004-11-28 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of an Arabidopsis WRKY DNA binding domain.
Plant Cell, 17, 2005
|
|
1WM9
 
 | |
1WU8
 
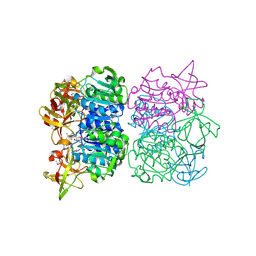 | |
1WUR
 
 | | Structure of GTP cyclohydrolase I Complexed with 8-oxo-dGTP | | Descriptor: | 8-OXO-2'-DEOXYGUANOSINE-5'-TRIPHOSPHATE, GTP cyclohydrolase I, ZINC ION | | Authors: | Tanaka, Y, Nakagawa, N, Masui, R, Yokoyama, S, Kuramitsu, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2004-12-08 | | Release date: | 2005-07-19 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.82 Å) | | Cite: | Novel reaction mechanism of GTP cyclohydrolase I. High-resolution X-ray crystallography of Thermus thermophilus HB8 enzyme complexed with a transition state analogue, the 8-oxoguanine derivative
J.Biochem.(Tokyo), 138, 2005
|
|
1WV8
 
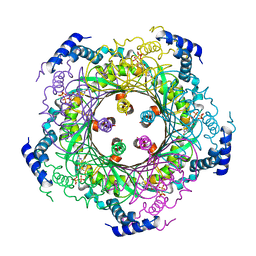
 | | Crystal structure of hypothetical protein TTHA1013 from an extremely thermophilic bacterium thermus thermophilus HB8 | | Descriptor: | hypothetical protein TTHA1013 | | Authors: | Mizohata, E, Hattori, M, Kuramitsu, S, Shirouzu, M, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2004-12-12 | | Release date: | 2005-06-12 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of the hypothetical protein TTHA1013 from Thermus thermophilus HB8
Proteins, 61, 2005
|
|
1X2M
 
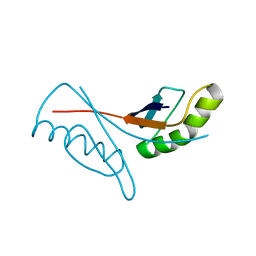
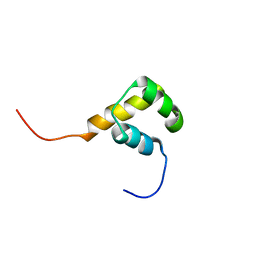 | | Solution structure of the homeobox domain of mouse LAG1 longevity assurance homolog 6 | | Descriptor: | LAG1 longevity assurance homolog 6 | | Authors: | Kamatari, Y.O, Tomizawa, T, Koshiba, S, Inoue, M, Kigawa, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2005-04-26 | | Release date: | 2005-10-26 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the homeobox domain of mouse LAG1 longevity assurance homolog 6
To be Published
|
|
