8CTO
 
 | |
8CUN
 
 | |
8CWA
 
 | |
4YZB
 
 | | CDPK1 from Eimeria tenella in complex with inhibitor UW1521 | | Descriptor: | 4-(6-ethoxynaphthalen-2-yl)-6-(piperazin-1-ylmethyl)-2H-indazol-3-amine, CALCIUM ION, Calmodulin-like domain protein kinase | | Authors: | Merritt, E.A. | | Deposit date: | 2015-03-24 | | Release date: | 2016-04-06 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | CDPK is a druggable target in the apicomplexan parasite Eimeria
to be published
|
|
5UJ5
 
 | |
6ALI
 
 | |
6AMR
 
 | |
2A0M
 
 | |
4WI1
 | |
4M97
 
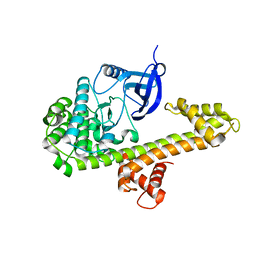 | | Calcium-Dependent Protein Kinase 1 from Neospora caninum | | Descriptor: | Calmodulin-like domain protein kinase isoenzyme gamma, related | | Authors: | Merritt, E.A. | | Deposit date: | 2013-08-14 | | Release date: | 2013-10-02 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Neospora caninum Calcium-Dependent Protein Kinase 1 Is an Effective Drug Target for Neosporosis Therapy.
Plos One, 9, 2014
|
|
5EJ2
 
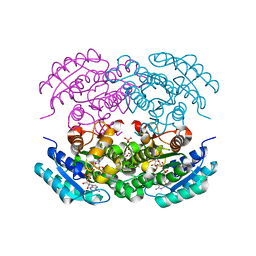 | |
4YSM
 
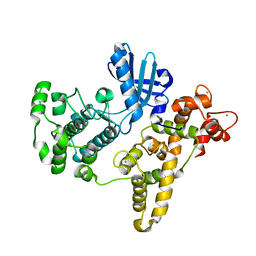 | |
4YUQ
 
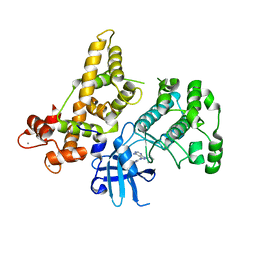 | | CDPK1 from Eimeria tenella in complex with inhibitor UW1354 | | Descriptor: | 1-cyclopentyl-3-(1H-pyrrolo[2,3-b]pyridin-5-yl)-1H-pyrazolo[3,4-d]pyrimidin-4-amine, CALCIUM ION, Calmodulin-like domain protein kinase | | Authors: | Merritt, E.A. | | Deposit date: | 2015-03-19 | | Release date: | 2016-04-06 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | CDPK is a druggable target in the apicomplexan parasite Eimeria
to be published
|
|
4MX9
 
 | | CDPK1 from Neospora caninum in complex with inhibitor UW1294 | | Descriptor: | 3-(6-ethoxynaphthalen-2-yl)-1-[(1-methylpiperidin-4-yl)methyl]-1H-pyrazolo[3,4-d]pyrimidin-4-amine, Calmodulin-like domain protein kinase isoenzyme gamma, related | | Authors: | Merritt, E.A. | | Deposit date: | 2013-09-26 | | Release date: | 2013-10-09 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Neospora caninum Calcium-Dependent Protein Kinase 1 Is an Effective Drug Target for Neosporosis Therapy.
Plos One, 9, 2014
|
|
4MXA
 
 | | CDPK1 from Neospora caninum in complex with inhibitor RM-1-132 | | Descriptor: | 3-(6-ethoxynaphthalen-2-yl)-1-(piperidin-4-ylmethyl)-1H-pyrazolo[3,4-d]pyrimidin-4-amine, Calmodulin-like domain protein kinase isoenzyme gamma, related | | Authors: | Merritt, E.A. | | Deposit date: | 2013-09-26 | | Release date: | 2013-10-09 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Neospora caninum Calcium-Dependent Protein Kinase 1 Is an Effective Drug Target for Neosporosis Therapy.
Plos One, 9, 2014
|
|
3URR
 
 | |
3V9O
 
 | |
3V8H
 
 | |
3VAV
 
 | |
3V7N
 
 | |
3V9P
 
 | |
3UW1
 
 | |
3V2I
 
 | |
3UW2
 
 | |
3UPT
 
 | |
