4N2O
 
 | | Structure of a novel autonomous cohesin protein from Ruminococcus flavefaciens | | Descriptor: | Autonomous cohesin, CHLORIDE ION | | Authors: | Frolow, F, Voronov-Goldman, M, Levy-Assaraf, M, Lamed, R, Bayer, E, Shimon, L. | | Deposit date: | 2013-10-05 | | Release date: | 2013-12-18 | | Last modified: | 2019-07-17 | | Method: | X-RAY DIFFRACTION (2.442 Å) | | Cite: | Structural characterization of a novel autonomous cohesin from Ruminococcus flavefaciens.
Acta Crystallogr F Struct Biol Commun, 70, 2014
|
|
4EYZ
 
 | | Crystal structure of an uncommon cellulosome-related protein module from Ruminococcus flavefaciens that resembles papain-like cysteine peptidases | | Descriptor: | 1,2-ETHANEDIOL, Cellulosome-related protein module from Ruminococcus flavefaciens that resembles papain-like cysteine peptidases | | Authors: | Frolow, F, Voronov-Goldman, M, Bayer, E, Lamed, R. | | Deposit date: | 2012-05-02 | | Release date: | 2013-03-20 | | Last modified: | 2017-11-15 | | Method: | X-RAY DIFFRACTION (1.383 Å) | | Cite: | Crystal Structure of an Uncommon Cellulosome-Related Protein Module from Ruminococcus flavefaciens That Resembles Papain-Like Cysteine Peptidases.
Plos One, 8, 2013
|
|
4IU2
 
 | | Cohesin-dockerin -X domain complex from Ruminococcus flavefacience | | Descriptor: | CALCIUM ION, CHLORIDE ION, Cell-wall anchoring protein, ... | | Authors: | Salama-Alber, O, Bayer, E, Frolow, F. | | Deposit date: | 2013-01-19 | | Release date: | 2013-04-24 | | Last modified: | 2013-07-03 | | Method: | X-RAY DIFFRACTION (2.001 Å) | | Cite: | Atypical Cohesin-Dockerin Complex Responsible for Cell Surface Attachment of Cellulosomal Components: BINDING FIDELITY, PROMISCUITY, AND STRUCTURAL BUTTRESSES.
J.Biol.Chem., 288, 2013
|
|
4IU3
 
 | | Cohesin-dockerin -X domain complex from Ruminococcus flavefacience | | Descriptor: | CALCIUM ION, Cell-wall anchoring protein, Cellulose-binding protein, ... | | Authors: | Salama-Alber, O, Bayer, E, Frolow, F. | | Deposit date: | 2013-01-19 | | Release date: | 2013-04-24 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.97 Å) | | Cite: | Atypical Cohesin-Dockerin Complex Responsible for Cell Surface Attachment of Cellulosomal Components: BINDING FIDELITY, PROMISCUITY, AND STRUCTURAL BUTTRESSES.
J.Biol.Chem., 288, 2013
|
|
3GHP
 
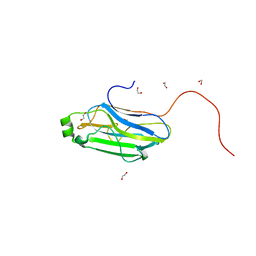 | |
3L8Q
 
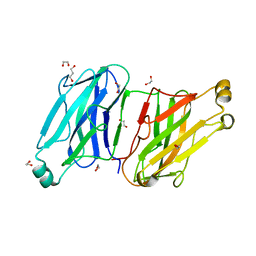 | | Structure analysis of the type II cohesin dyad from the adaptor ScaA scaffoldin of Acetivibrio cellulolyticus | | Descriptor: | 1,2-ETHANEDIOL, 1,3-PROPANDIOL, Cellulosomal scaffoldin adaptor protein B, ... | | Authors: | Noach, I, Frolow, F, Bayer, E.A. | | Deposit date: | 2010-01-03 | | Release date: | 2010-05-05 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.57 Å) | | Cite: | Modular Arrangement of a Cellulosomal Scaffoldin Subunit Revealed from the Crystal Structure of a Cohesin Dyad
J.Mol.Biol., 399, 2010
|
|
1QZN
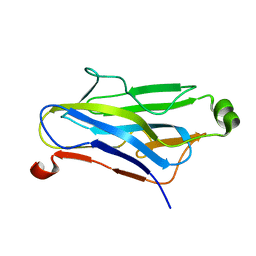 
 | | Crystal Structure Analysis of a type II cohesin domain from the cellulosome of Acetivibrio cellulolyticus | | Descriptor: | cellulosomal scaffoldin adaptor protein B | | Authors: | Frolow, F, Noach, I, Rosenheck, S, Lamed, R, Qi, X, Shimon, L.J.W, Bayer, E.A. | | Deposit date: | 2003-09-17 | | Release date: | 2004-09-21 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of a type-II cohesin module from the Bacteroides cellulosolvens cellulosome reveals novel and distinctive secondary structural elements.
J.Mol.Biol., 348, 2005
|
|
2ZF9
 
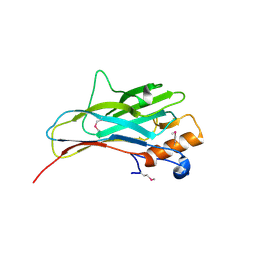 | |
3FNK
 
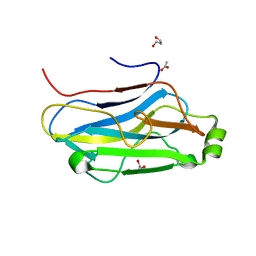 | | Crystal structure of the second type II cohesin module from the cellulosomal adaptor ScaA scaffoldin of Acetivibrio cellulolyticus | | Descriptor: | 1,2-ETHANEDIOL, 1,3-PROPANDIOL, 1,4-BUTANEDIOL, ... | | Authors: | Noach, I, Frolow, F, Bayer, E.A. | | Deposit date: | 2008-12-25 | | Release date: | 2009-06-23 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.99 Å) | | Cite: | Intermodular Linker Flexibility Revealed from Crystal Structures of Adjacent Cellulosomal Cohesins of Acetivibrio cellulolyticus
J.Mol.Biol., 391, 2009
|
|
5VSG
 
 | | Fibrils of the super helical repeat peptide, SHR-FF, grown at elevated temperature | | Descriptor: | Super Helical Repeat Peptide SHR-FF | | Authors: | Mondal, S, Sawaya, M.R, Eisenberg, D.S, Gazit, E. | | Deposit date: | 2017-05-11 | | Release date: | 2018-06-27 | | Last modified: | 2020-01-01 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | Transition of Metastable Cross-alpha Crystals into Cross-beta Fibrils by beta-Turn Flipping.
J.Am.Chem.Soc., 141, 2019
|
|
3D0A
 
 | |
3D07
 
 | |
3D09
 
 | |
3ZUC
 | | Structure of CBM3b of major scaffoldin subunit ScaA from Acetivibrio cellulolyticus determined from the crystals grown in the presence of Nickel | | Descriptor: | 1,2-ETHANEDIOL, CALCIUM ION, CELLULOSOMAL SCAFFOLDIN, ... | | Authors: | Yaniv, O, Halfon, Y, Lamed, R, Frolow, F. | | Deposit date: | 2011-07-18 | | Release date: | 2012-01-11 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.001 Å) | | Cite: | Structure of Cbm3B of the Major Scaffoldin Subunit Scaa from Acetivibrio Cellulolyticus
Acta Crystallogr.,Sect.F, 68, 2012
|
|
3ZQW
 | | Structure of CBM3b of major scaffoldin subunit ScaA from Acetivibrio cellulolyticus | | Descriptor: | 1,2-ETHANEDIOL, CALCIUM ION, CELLULOSOMAL SCAFFOLDIN, ... | | Authors: | Yaniv, O, Halfon, Y, Lamed, R, Frolow, F. | | Deposit date: | 2011-06-12 | | Release date: | 2012-01-11 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.07 Å) | | Cite: | Structure of Cbm3B of the Major Scaffoldin Subunit Scaa from Acetivibrio Cellulolyticus
Acta Crystallogr.,Sect.F, 68, 2012
|
|
3ZU8
 | | STRUCTURE OF CBM3B OF MAJOR SCAFFOLDIN SUBUNIT SCAA FROM ACETIVIBRIO CELLULOLYTICUS DETERMINED ON THE NIKEL ABSORPTION EDGE | | Descriptor: | 1,2-ETHANEDIOL, CALCIUM ION, CELLULOSOMAL SCAFFOLDIN, ... | | Authors: | Yaniv, O, Halfon, Y, Lamed, R, Frolow, F. | | Deposit date: | 2011-07-17 | | Release date: | 2012-01-11 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.801 Å) | | Cite: | Structure of Cbm3B of the Major Scaffoldin Subunit Scaa from Acetivibrio Cellulolyticus
Acta Crystallogr.,Sect.F, 68, 2012
|
|
4B96
 | | Family 3b carbohydrate-binding module from the biomass sensoring system of Clostridium clariflavum | | Descriptor: | CALCIUM ION, CELLULOSE BINDING DOMAIN-CONTAINING PROTEIN, CHLORIDE ION | | Authors: | Yaniv, O, Reddy, Y.H.K, Yoffe, H, Shimon, L.J.W, Bayer, E.A, Lamed, R, Frolow, F. | | Deposit date: | 2012-09-02 | | Release date: | 2013-09-18 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.911 Å) | | Cite: | Structure of Cbm3B from the Biomass Sensoring System of Clostridium Clarifalvum
To be Published
|
|
