3UYL
 | | Spinosyn Rhamnosyltransferase SpnG complexed with thymidine diphosphate | | Descriptor: | MAGNESIUM ION, NDP-rhamnosyltransferase, THYMIDINE-5'-DIPHOSPHATE, ... | | Authors: | Isiorho, E.A, Liu, H.-W, Keatinge-Clay, A.T. | | Deposit date: | 2011-12-06 | | Release date: | 2012-02-15 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Structural Studies of the Spinosyn Rhamnosyltransferase, SpnG.
Biochemistry, 51, 2012
|
|
4J1W
 
 | |
4N6F
 
 | | Crystal structure of Amycolatopsis orientalis BexX complexed with G6P | | Descriptor: | CALCIUM ION, FRUCTOSE -6-PHOSPHATE, Putative thiosugar synthase | | Authors: | Zhang, X, Zhang, Y, Kinsland, C, Sasaki, E, Sun, H.G, Lu, M.J, Liu, T, Ou, A, Li, J, Chen, Y, Liu, H, Ealick, S.E. | | Deposit date: | 2013-10-11 | | Release date: | 2014-05-14 | | Last modified: | 2017-11-15 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Co-opting sulphur-carrier proteins from primary metabolic pathways for 2-thiosugar biosynthesis.
Nature, 509, 2014
|
|
4N6E
 
 | | Crystal structure of Amycolatopsis orientalis BexX/CysO complex | | Descriptor: | Putative thiosugar synthase, SULFATE ION, ThiS/MoaD family protein | | Authors: | Zhang, X, Zhang, Y, Kinsland, C, Sasaki, E, Sun, H.G, Lu, M.J, Liu, T, Ou, A, Li, J, Chen, Y, Liu, H, Ealick, S.E. | | Deposit date: | 2013-10-11 | | Release date: | 2014-05-14 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Co-opting sulphur-carrier proteins from primary metabolic pathways for 2-thiosugar biosynthesis.
Nature, 509, 2014
|
|
4LEI
 
 | |
5TK6
 
 | |
5UL2
 
 | |
5UL3
 
 | | Structure of Cobalamin-dependent S-adenosylmethionine radical enzyme OxsB with aqua-cobalamin bound | | Descriptor: | 1,2-ETHANEDIOL, 2,3-DIHYDROXY-1,4-DITHIOBUTANE, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, ... | | Authors: | Bridwell-Rabb, J, Drennan, C.L. | | Deposit date: | 2017-01-24 | | Release date: | 2017-04-19 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | A B12-dependent radical SAM enzyme involved in oxetanocin A biosynthesis.
Nature, 544, 2017
|
|
5TKA
 
 | |
5TK7
 
 | |
5TK8
 
 | |
5TK9
 
 | | Structure of the HD-domain phosphohydrolase OxsA with Oxetanocin-A bound | | Descriptor: | MAGNESIUM ION, OxsA protein, [(2S,3R,4R)-4-(6-amino-9H-purin-9-yl)oxetane-2,3-diyl]dimethanol | | Authors: | Bridwell-Rabb, J, Drennan, C.L. | | Deposit date: | 2016-10-06 | | Release date: | 2016-11-16 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.843 Å) | | Cite: | An HD domain phosphohydrolase active site tailored for oxetanocin-A biosynthesis.
Proc. Natl. Acad. Sci. U.S.A., 113, 2016
|
|
5UL4
 
 | |
2JVN
 
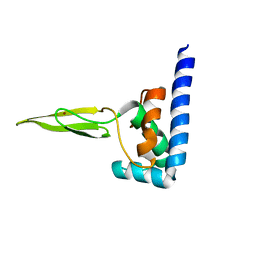 | | Domain C of human PARP-1 | | Descriptor: | Poly [ADP-ribose] polymerase 1, ZINC ION | | Authors: | Hoffman, D, Tao, Z, Liu, H. | | Deposit date: | 2007-09-24 | | Release date: | 2008-05-13 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Domain C of human poly(ADP-ribose) polymerase-1 is important for enzyme activity and contains a novel zinc-ribbon motif.
Biochemistry, 47, 2008
|
|
6NKO
 
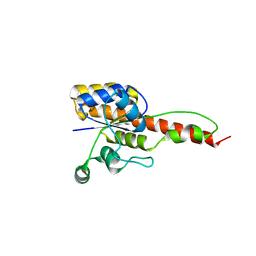 | | Crystal structure of ForH | | Descriptor: | ForH | | Authors: | Zheng, J, Irani, S, Zhang, Y. | | Deposit date: | 2019-01-07 | | Release date: | 2019-04-17 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.403 Å) | | Cite: | Identification of the Formycin A Biosynthetic Gene Cluster from Streptomyces kaniharaensis Illustrates the Interplay between Biological Pyrazolopyrimidine Formation and de Novo Purine Biosynthesis.
J. Am. Chem. Soc., 141, 2019
|
|
7SCP
 
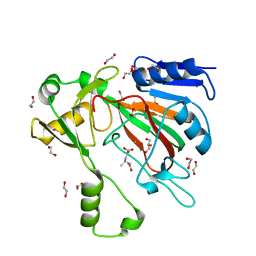 | | The crystal structure of ScoE in complex with intermediate | | Descriptor: | (3R)-3-(oxaloamino)butanoic acid, 1,2-ETHANEDIOL, FE (II) ION, ... | | Authors: | Cha, L, Chen, J, Zhou, J, Chang, W. | | Deposit date: | 2021-09-28 | | Release date: | 2022-03-23 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.99 Å) | | Cite: | Deciphering the Reaction Pathway of Mononuclear Iron Enzyme-Catalyzed N-C Triple Bond Formation in Isocyanide Lipopeptide and Polyketide Biosynthesis
Acs Catalysis, 12, 2022
|
|
7V6H
 
 | | Crystal Structure of the SpnL | | Descriptor: | Cyclopropane fatty-acyl-phospholipid synthase-like methyltransferase, S-ADENOSYL-L-HOMOCYSTEINE | | Authors: | Wu, H.-H, Ko, T.-P, Liu, H.-W, Tsai, M.-D. | | Deposit date: | 2021-08-20 | | Release date: | 2021-12-29 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (3.054 Å) | | Cite: | Evidence for an Enzyme-Catalyzed Rauhut-Currier Reaction during the Biosynthesis of Spinosyn A.
J.Am.Chem.Soc., 143, 2021
|
|
3SCH
 
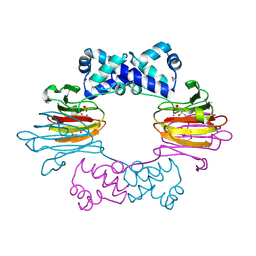 | | Co(II)-HppE with R-HPP | | Descriptor: | COBALT (II) ION, Epoxidase, [(2R)-2-hydroxypropyl]phosphonic acid | | Authors: | Drennan, C.L. | | Deposit date: | 2011-06-07 | | Release date: | 2011-07-06 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural basis of regiospecificity of a mononuclear iron enzyme in antibiotic fosfomycin biosynthesis.
J.Am.Chem.Soc., 133, 2011
|
|
3SCF
 
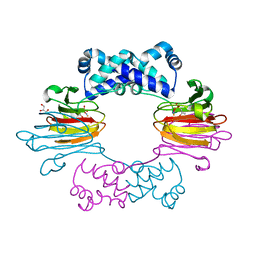 | | Fe(II)-HppE with S-HPP and NO | | Descriptor: | (S)-2-HYDROXYPROPYLPHOSPHONIC ACID, Epoxidase, FE (II) ION, ... | | Authors: | Drennan, C.L. | | Deposit date: | 2011-06-07 | | Release date: | 2011-07-06 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | Structural basis of regiospecificity of a mononuclear iron enzyme in antibiotic fosfomycin biosynthesis.
J.Am.Chem.Soc., 133, 2011
|
|
3SCG
 
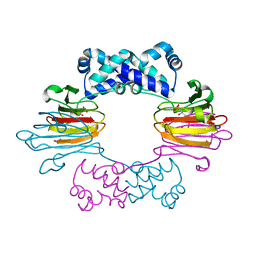 | | Fe(II)-HppE with R-HPP | | Descriptor: | Epoxidase, FE (II) ION, [(2R)-2-hydroxypropyl]phosphonic acid | | Authors: | Drennan, C.L. | | Deposit date: | 2011-06-07 | | Release date: | 2011-07-06 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structural basis of regiospecificity of a mononuclear iron enzyme in antibiotic fosfomycin biosynthesis.
J.Am.Chem.Soc., 133, 2011
|
|
8Q72
 
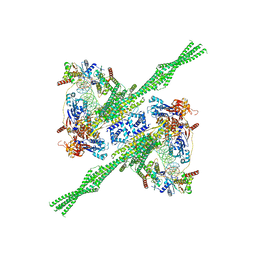 | | E. coli plasmid-borne JetABCD(E248A) core in a cleavage-competent state | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, Circular plasmid DNA (1840-MER), JetA, ... | | Authors: | Roisne-Hamelin, F, Li, Y, Gruber, S. | | Deposit date: | 2023-08-15 | | Release date: | 2024-01-31 | | Last modified: | 2024-03-20 | | Method: | ELECTRON MICROSCOPY (4.17 Å) | | Cite: | Structural basis for plasmid restriction by SMC JET nuclease.
Mol.Cell, 84, 2024
|
|
7VOC
 
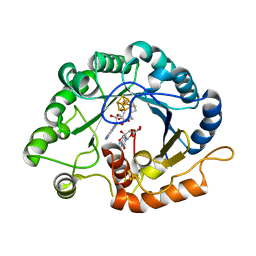 | | The crystal structure of a Radical SAM Enzyme BlsE involved in the Biosynthesis of Blasticidin S | | Descriptor: | (2~{S},3~{S},4~{S},5~{R},6~{R})-6-(4-azanyl-2-oxidanylidene-pyrimidin-1-yl)-3,4,5-tris(oxidanyl)oxane-2-carboxylic acid, Cytosylglucuronate decarboxylase, GLYCEROL, ... | | Authors: | Hou, X.L, Zhou, J.H. | | Deposit date: | 2021-10-13 | | Release date: | 2022-05-04 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.62005424 Å) | | Cite: | Radical S -Adenosyl Methionine Enzyme BlsE Catalyzes a Radical-Mediated 1,2-Diol Dehydration during the Biosynthesis of Blasticidin S.
J.Am.Chem.Soc., 144, 2022
|
|
7VOB
 
 | |
8HZY
 
 | |
8HZV
 
 | | The crystal structure of a Radical SAM Enzyme DesII | | Descriptor: | GLYCEROL, IRON/SULFUR CLUSTER, METHIONINE, ... | | Authors: | Hou, X.L, Zhou, J.H. | | Deposit date: | 2023-01-09 | | Release date: | 2024-07-10 | | Method: | X-RAY DIFFRACTION (2.33001685 Å) | | Cite: | Mechanistic Insights from the Crystal Structure and Computational Analysis of the Radical SAM Deaminase DesII.
Adv Sci, 2024
|
|
