2LYZ
 
 | |
3LYZ
 
 | |
6LYZ
 | |
5LYT
 
 | |
5LYZ
 
 | |
6LYT
 
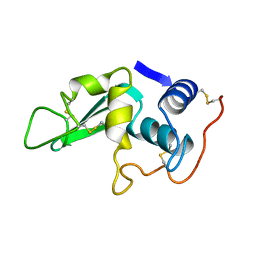 | |
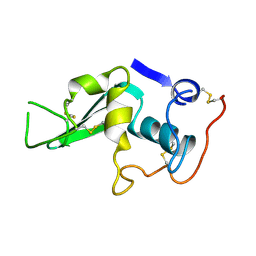
1LYZ
 | |
1C6T
 | |
1C6P
 | |
1C6Q
 | |
1EPY
 
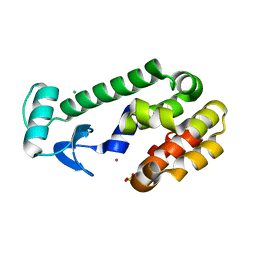 | | T4 LYSOZYME MUTANT, T21H/C54T/C97A/Q141H/T142H | | Descriptor: | CHLORIDE ION, COBALT (II) ION, LYSOZYME, ... | | Authors: | Wray, J.W, Baase, W.A, Ostheimer, G.J, Zhang, X.-J, Matthews, B.W. | | Deposit date: | 2000-03-29 | | Release date: | 2000-04-12 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Use of a non-rigid region in T4 lysozyme to design an adaptable metal-binding site.
Protein Eng., 13, 2000
|
|
3HTC
 
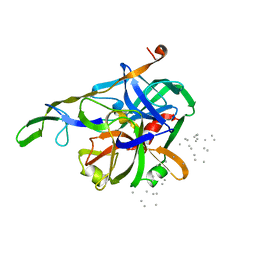 | | THE STRUCTURE OF A COMPLEX OF RECOMBINANT HIRUDIN AND HUMAN ALPHA-THROMBIN | | Descriptor: | ALPHA-THROMBIN (LARGE SUBUNIT), ALPHA-THROMBIN (SMALL SUBUNIT), HIRUDIN VARIANT 2 | | Authors: | Tulinsky, A, Rydel, T.J, Ravichandran, K.G, Huber, R, Bode, W. | | Deposit date: | 1993-06-11 | | Release date: | 1994-01-31 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The structure of a complex of recombinant hirudin and human alpha-thrombin.
Science, 249, 1990
|
|
