2JZF
 
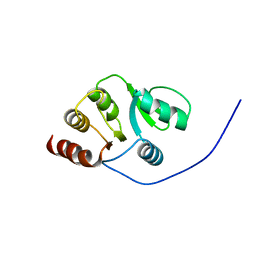 | | NMR Conformer closest to the mean coordinates of the domain 513-651 of the SARS-CoV nonstructural protein nsp3 | | Descriptor: | Replicase polyprotein 1ab | | Authors: | Chatterjee, A, Johnson, M.A, Serrano, P, Pedrini, B, Joseph, J, Saikatendu, K, Neuman, B, Stevens, R.C, Wilson, I.A, Buchmeier, M.J, Kuhn, P, Wuthrich, K, Joint Center for Structural Genomics (JCSG) | | Deposit date: | 2008-01-04 | | Release date: | 2008-02-05 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Nuclear magnetic resonance structure shows that the severe acute respiratory syndrome coronavirus-unique domain contains a macrodomain fold.
J.Virol., 83, 2009
|
|
4KIK
 
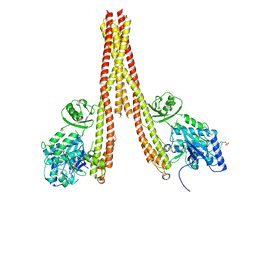 | | Human IkB kinase beta | | Descriptor: | Inhibitor of nuclear factor kappa-B kinase subunit beta, K-252A | | Authors: | Liu, S, Mosyak, L. | | Deposit date: | 2013-05-02 | | Release date: | 2013-06-26 | | Last modified: | 2013-08-28 | | Method: | X-RAY DIFFRACTION (2.83 Å) | | Cite: | Crystal Structure of a Human I kappa B Kinase beta Asymmetric Dimer.
J.Biol.Chem., 288, 2013
|
|
1PZ5
 
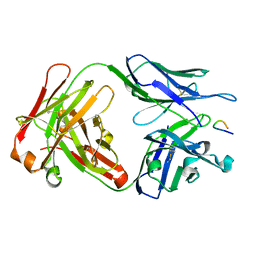 | | Structural basis of peptide-carbohydrate mimicry in an antibody combining site | | Descriptor: | Heavy chain of Fab (SYA/J6), Light chain of Fab (SYA/J6), Octapeptide (MDWNMHAA) | | Authors: | Vyas, N.K, Vyas, M.N, Chervenak, M.C, Bundle, D.R, Pinto, B.M, Quiocho, F.A. | | Deposit date: | 2003-07-09 | | Release date: | 2003-11-18 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural basis of peptide-carbohydrate mimicry in an antibody combining site.
Proc.Natl.Acad.Sci.USA, 100, 2003
|
|
