8IJU
 
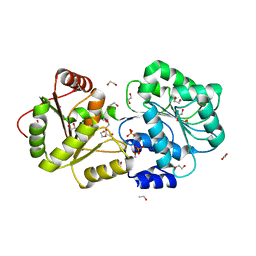 | | ATP-dependent RNA helicase DDX39A (URH49delta41) | | Descriptor: | 1,2-ETHANEDIOL, ATP-dependent RNA helicase DDX39A, PHOSPHATE ION, ... | | Authors: | Mikami, B, Fujita, K, Masuda, S, Kojima, M. | | Deposit date: | 2023-02-28 | | Release date: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.82 Å) | | Cite: | Structural differences between the closely related RNA helicases, UAP56 and URH49, fashion distinct functional apo-complexes.
Nat Commun, 15, 2024
|
|
1BOL
 
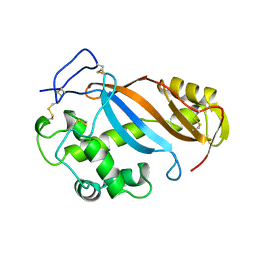 | |
3A75
 
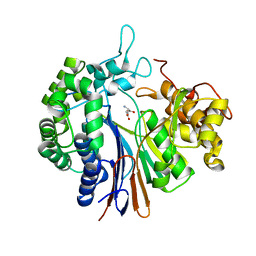 | |
4JU2
 
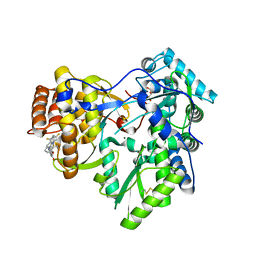 | | Crystal structure of hcv ns5b polymerase in complex with compound 12 | | Descriptor: | 6-[3-(pyridin-2-yl)-2-(trifluoromethyl)phenoxy]-1-(2,4,6-trifluorobenzyl)quinazolin-4(1H)-one, GLYCEROL, Genome polyprotein, ... | | Authors: | Coulombe, R. | | Deposit date: | 2013-03-24 | | Release date: | 2013-07-03 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Molecular Dynamics Simulations and Structure-Based Rational Design Lead to Allosteric HCV NS5B Polymerase Thumb Pocket 2 Inhibitor with Picomolar Cellular Replicon Potency.
J.Med.Chem., 57, 2014
|
|
4JTY
 
 | | Crystal structure of HCV NS5B polymerase with COMPOUND 2 | | Descriptor: | 3-{[4-oxo-1-(2,4,6-trifluorobenzyl)-1,4-dihydroquinazolin-6-yl]oxy}-2-(trifluoromethyl)benzamide, GLYCEROL, Genome polyprotein, ... | | Authors: | Coulombe, R. | | Deposit date: | 2013-03-24 | | Release date: | 2013-07-03 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Molecular Dynamics Simulations and Structure-Based Rational Design Lead to Allosteric HCV NS5B Polymerase Thumb Pocket 2 Inhibitor with Picomolar Cellular Replicon Potency.
J.Med.Chem., 57, 2014
|
|
4JTW
 
 | | Crystal structure of HCV NS5B polymerase in complex with coupound 1 | | Descriptor: | 1-(2,4,6-trifluorobenzyl)-6-[2-(trifluoromethyl)phenoxy]quinazolin-4(1H)-one, GLYCEROL, Genome polyprotein, ... | | Authors: | Coulombe, R. | | Deposit date: | 2013-03-24 | | Release date: | 2013-07-10 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Molecular Dynamics Simulations and Structure-Based Rational Design Lead to Allosteric HCV NS5B Polymerase Thumb Pocket 2 Inhibitor with Picomolar Cellular Replicon Potency.
J.Med.Chem., 57, 2014
|
|
4JU1
 
 | | Crystal structure of hcv ns5b polymerase in complex with compound 6 | | Descriptor: | 6-[3-([1,3]oxazolo[4,5-b]pyridin-2-yl)-2-(trifluoromethyl)phenoxy]-1-(2,4,6-trifluorobenzyl)quinazolin-4(1H)-one, GLYCEROL, Genome polyprotein, ... | | Authors: | Coulombe, R. | | Deposit date: | 2013-03-24 | | Release date: | 2013-07-03 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Molecular Dynamics Simulations and Structure-Based Rational Design Lead to Allosteric HCV NS5B Polymerase Thumb Pocket 2 Inhibitor with Picomolar Cellular Replicon Potency.
J.Med.Chem., 57, 2014
|
|
4JTZ
 
 | | Crystal structure of hcv ns5b polymerase in complex with compound 4 | | Descriptor: | 3-{[4-oxo-1-(2,4,6-trifluorobenzyl)-1,4-dihydroquinazolin-6-yl]oxy}-N-(pyridin-3-yl)-2-(trifluoromethyl)benzamide, GLYCEROL, Genome polyprotein, ... | | Authors: | Coulombe, R. | | Deposit date: | 2013-03-24 | | Release date: | 2013-07-03 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Molecular Dynamics Simulations and Structure-Based Rational Design Lead to Allosteric HCV NS5B Polymerase Thumb Pocket 2 Inhibitor with Picomolar Cellular Replicon Potency.
J.Med.Chem., 57, 2014
|
|
4JJU
 
 | | Crystal structure of HCV NS5B polymerase in complex with COMPOUND 29 | | Descriptor: | 1-(2,4-difluorobenzyl)-6-{[3-(trifluoromethyl)pyridin-2-yl]oxy}quinazolin-4(1H)-one, GLYCEROL, Genome polyprotein, ... | | Authors: | Coulombe, R. | | Deposit date: | 2013-03-08 | | Release date: | 2013-06-12 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.91 Å) | | Cite: | Structure-based design of novel HCV NS5B thumb pocket 2 allosteric inhibitors with submicromolar gt1 replicon potency: Discovery of a quinazolinone chemotype.
Bioorg.Med.Chem.Lett., 23, 2013
|
|
4JJS
 
 | | Crystal structure of HCV NS5B polymerase in complex with COMPOUND 2 | | Descriptor: | 2-{[(trans-4-methylcyclohexyl)carbonyl](propan-2-yl)amino}-5-[2-(trifluoromethyl)phenoxy]benzoic acid, Genome polyprotein, MAGNESIUM ION | | Authors: | Coulombe, R. | | Deposit date: | 2013-03-08 | | Release date: | 2013-06-12 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structure-based design of novel HCV NS5B thumb pocket 2 allosteric inhibitors with submicromolar gt1 replicon potency: Discovery of a quinazolinone chemotype.
Bioorg.Med.Chem.Lett., 23, 2013
|
|
6TL2
 
 | |
3MWV
 
 | | Crystal structure of HCV NS5B polymerase | | Descriptor: | Genome polyprotein | | Authors: | Coulombe, R. | | Deposit date: | 2010-05-06 | | Release date: | 2010-10-27 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Importance of ligand bioactive conformation in the discovery of potent indole-diamide inhibitors of the hepatitis C virus NS5B.
J.Am.Chem.Soc., 132, 2010
|
|
3MWW
 
 | | Crystal structure of HCV NS5B polymerase | | Descriptor: | 1-[2-(4-carboxypiperidin-1-yl)-2-oxoethyl]-3-cyclohexyl-2-furan-3-yl-1H-indole-6-carboxylic acid, Genome polyprotein, SULFATE ION | | Authors: | Coulombe, R. | | Deposit date: | 2010-05-06 | | Release date: | 2010-10-27 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Importance of ligand bioactive conformation in the discovery of potent indole-diamide inhibitors of the hepatitis C virus NS5B.
J.Am.Chem.Soc., 132, 2010
|
|
