3HEU
 
 | |
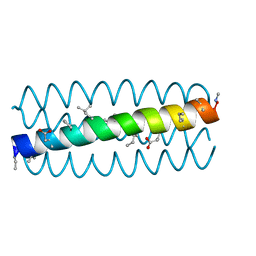
3HET
 | |
3HEX
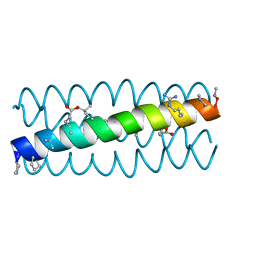 
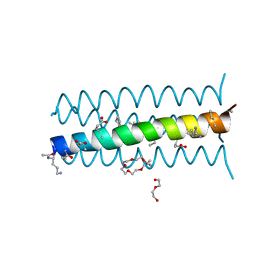
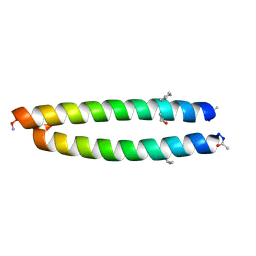
 | | Cyclic residues in alpha/beta-peptide helix bundles: GCN4-pLI side chain sequence on an (alpha-alpha-beta) backbone with cyclic beta-residues at positions 1, 4, 19 and 28 | | Descriptor: | alpha/beta-peptide based on the GCN4-pLI side chain sequence with an (alpha-alpha-beta) backbone and cyclic beta-residues at positions 1, 4, 19 and 28 | | Authors: | Horne, W.S, Price, J.L, Gellman, S.H. | | Deposit date: | 2009-05-10 | | Release date: | 2010-04-21 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structural consequences of beta-amino acid preorganization in a self-assembling alpha/beta-peptide: fundamental studies of foldameric helix bundles.
J.Am.Chem.Soc., 132, 2010
|
|
6E5K
 
 | | Heterogeneous-Backbone Mimics of a Designed Disulfide-Rich Protein: Aib turn, Aib helix, N-methyl hairpin | | Descriptor: | Designed peptide NC_HEE_D1: Aib turn, Aib helix, N-methyl hairpin mutant | | Authors: | Cabalteja, C.C, Mihalko, D.S, Horne, W.S. | | Deposit date: | 2018-07-20 | | Release date: | 2018-11-21 | | Last modified: | 2020-01-01 | | Method: | SOLUTION NMR | | Cite: | Heterogeneous-Backbone Foldamer Mimics of a Computationally Designed, Disulfide-Rich Miniprotein.
Chembiochem, 20, 2019
|
|
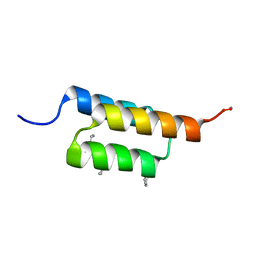
7URJ
 | |
4WH4
 
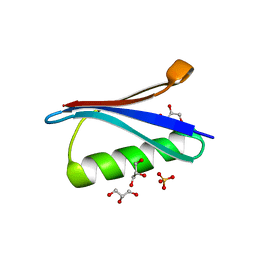 | | Protein GB1 Quadruple Mutant I6H/N8H/K28H/Q32H | | Descriptor: | GLYCEROL, Immunoglobulin G-binding protein G, SULFATE ION | | Authors: | Cunningham, T.C, Horne, W.S, Saxena, S. | | Deposit date: | 2014-09-19 | | Release date: | 2015-08-05 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The double-histidine Cu2+-binding motif: a highly rigid, site-specific spin probe for electron spin resonance distance measurements.
Angew.Chem.Int.Ed.Engl., 54, 2015
|
|
4TL1
 | | GCN4-p1 with mutation to 1-Aminocyclohexanecarboxylic acid at residue 10 | | Descriptor: | General control protein GCN4, ISOPROPYL ALCOHOL | | Authors: | Tavenor, N.A, Silva, K.I, Saxena, S, Horne, W.S. | | Deposit date: | 2014-05-28 | | Release date: | 2014-08-06 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Origins of Structural Flexibility in Protein-Based Supramolecular Polymers Revealed by DEER Spectroscopy.
J.Phys.Chem.B, 118, 2014
|
|
7RAP
 
 | |
6E5I
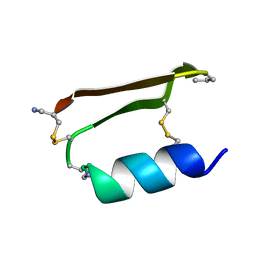 
 | |
6E5J
 
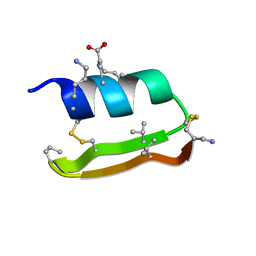 | | Heterogeneous-Backbone Mimics of a Designed Disulfide-Rich Protein: Aib turn, beta3 helix, N-methyl hairpin | | Descriptor: | Designed peptide NC_HEE_D1: Aib turn, beta3 helix, N-methyl hairpin mutant | | Authors: | Cabalteja, C.C, Mihalko, D.S, Horne, W.S. | | Deposit date: | 2018-07-20 | | Release date: | 2018-11-21 | | Last modified: | 2023-11-15 | | Method: | SOLUTION NMR | | Cite: | Heterogeneous-Backbone Foldamer Mimics of a Computationally Designed, Disulfide-Rich Miniprotein.
Chembiochem, 20, 2019
|
|
6E5H
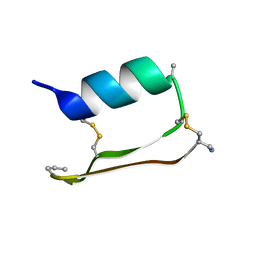 
 | |
4NJ0
 
 | | GCN4-p1 single Val9 to Ile mutant | | Descriptor: | General control protein GCN4 | | Authors: | Oshaben, K.M, Horne, W.S. | | Deposit date: | 2013-11-08 | | Release date: | 2014-08-20 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Tuning assembly size in Peptide-based supramolecular polymers by modulation of subunit association affinity.
Biomacromolecules, 15, 2014
|
|
4NJ1
 
 | | GCN4-p1 double Val9, 23 to Ile mutant | | Descriptor: | General control protein GCN4 | | Authors: | Oshaben, K.M, Horne, W.S. | | Deposit date: | 2013-11-08 | | Release date: | 2014-08-20 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Tuning assembly size in Peptide-based supramolecular polymers by modulation of subunit association affinity.
Biomacromolecules, 15, 2014
|
|
4NIZ
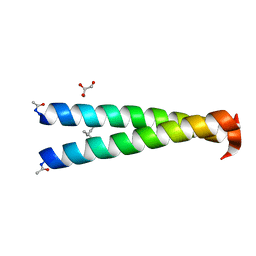 
 | | GCN4-p1 single Val9 to aminobutyric acid mutant | | Descriptor: | GLYCEROL, General control protein GCN4 | | Authors: | Oshaben, K.M, Horne, W.S. | | Deposit date: | 2013-11-08 | | Release date: | 2014-08-20 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Tuning assembly size in Peptide-based supramolecular polymers by modulation of subunit association affinity.
Biomacromolecules, 15, 2014
|
|
4NJ2
 
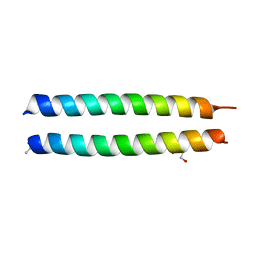 | | GCN4-p1 triple Val9, 23,30 to Ile mutant | | Descriptor: | GLYCEROL, General control protein GCN4 | | Authors: | Oshaben, K.M, Horne, W.S. | | Deposit date: | 2013-11-08 | | Release date: | 2014-08-20 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Tuning assembly size in Peptide-based supramolecular polymers by modulation of subunit association affinity.
Biomacromolecules, 15, 2014
|
|
9BB5
 | |
9BB2
 
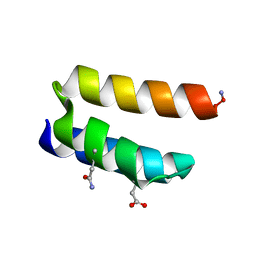 | |
9BB1
 | |
9BB4
 | |
9BB7
 
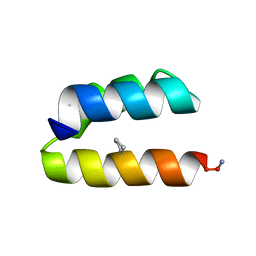 | |
9BB3
 
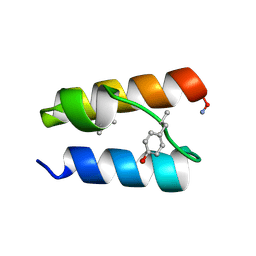 | |
9BB6
 | |
6OLN
 
 | |
3V3X
 
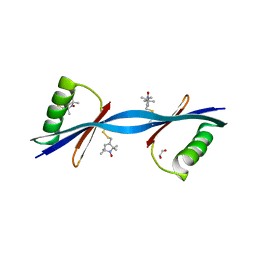 | | Nitroxide Spin Labels in Protein GB1: N8/K28 Double Mutant | | Descriptor: | ACETATE ION, GLYCEROL, Immunoglobulin G-binding protein G, ... | | Authors: | Cunningham, T.F, McGoff, M.S, Sengupta, I, Jaroniec, C.P, Horne, W.S, Saxena, S.K. | | Deposit date: | 2011-12-14 | | Release date: | 2012-08-29 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | High-resolution structure of a protein spin-label in a solvent-exposed beta-sheet and comparison with DEER spectroscopy.
Biochemistry, 51, 2012
|
|
4A1U
 
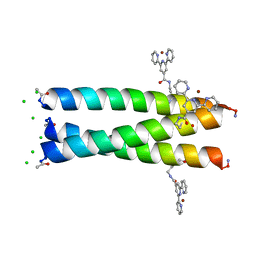
 | | Crystal structure of alpha-beta-foldamer 2c in complex with Bcl-xL | | Descriptor: | ALPHA-BETA-FOLDAMER 2C, BCL-2-LIKE PROTEIN 1, CHLORIDE ION, ... | | Authors: | Boersma, M.D, Haase, H.S, Kaufman, K.J, Horne, W.S, Lee, E.F, Clarke, O.B, Smith, B.J, Colman, P.M, Gellman, S.H, Fairlie, W.D. | | Deposit date: | 2011-09-20 | | Release date: | 2011-12-28 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.54 Å) | | Cite: | Evaluation of Diverse Alpha/Beta-Backbone Patterns for Functional Alpha-Helix Mimicry: Analogues of the Bim Bh3 Domain.
J.Am.Chem.Soc., 134, 2012
|
|
