6Y74
 
 | |
7UWU
 
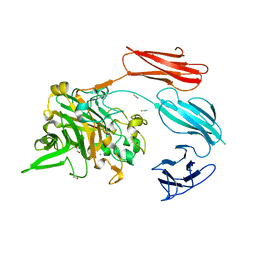 | | Starch adherence system protein 6 (Sas6) | | Descriptor: | 1,2-ETHANEDIOL, CALCIUM ION, Starch Adherence System protein 6 (Sas6), ... | | Authors: | Photenhauer, A.L, Koropatkin, N.M. | | Deposit date: | 2022-05-04 | | Release date: | 2023-06-14 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.19 Å) | | Cite: | The Ruminococcus bromii amylosome protein Sas6 binds single and double helical alpha-glucan structures in starch.
Nat.Struct.Mol.Biol., 31, 2024
|
|
7UWW
 
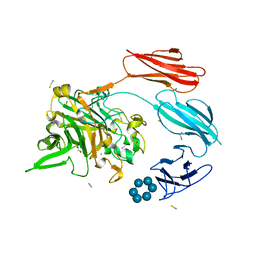 | | Sas6 with alpha-cyclodextrin | | Descriptor: | 1,2-ETHANEDIOL, CALCIUM ION, Cyclohexakis-(1-4)-(alpha-D-glucopyranose), ... | | Authors: | Photenhauer, A.L, Koropatkin, N.M. | | Deposit date: | 2022-05-04 | | Release date: | 2023-06-14 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.61 Å) | | Cite: | The Ruminococcus bromii amylosome protein Sas6 binds single and double helical alpha-glucan structures in starch.
Nat.Struct.Mol.Biol., 31, 2024
|
|
7UWV
 
 | |
6VSK
 
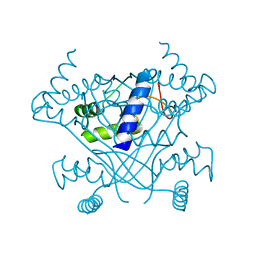 | | Crystal structure of the P-Rex1 DEP1 domain | | Descriptor: | Phosphatidylinositol 3,4,5-trisphosphate-dependent Rac exchanger 1 protein | | Authors: | Tesmer, J, Ravala, S.K. | | Deposit date: | 2020-02-11 | | Release date: | 2020-07-22 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (3.12 Å) | | Cite: | The first DEP domain of the RhoGEF P-Rex1 autoinhibits activity and contributes to membrane binding.
J.Biol.Chem., 295, 2020
|
|
8EDC
 
 | | Structure of C. elegans UNC-5 IG 1+2 Domains | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, Netrin receptor unc-5, SULFATE ION | | Authors: | Priest, J.M, Ozkan, E. | | Deposit date: | 2022-09-04 | | Release date: | 2023-01-11 | | Last modified: | 2024-08-14 | | Method: | X-RAY DIFFRACTION (2.89 Å) | | Cite: | Structural insights into the formation of repulsive netrin guidance complexes.
Sci Adv, 10, 2024
|
|
8EDI
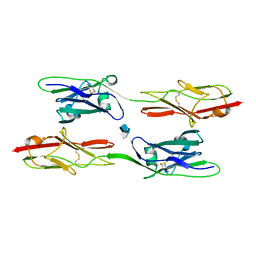 
 | | Structure of C. elegans UNC-5 IG 1+2 Domains bound to Heparin dp4 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, 4-deoxy-2-O-sulfo-alpha-L-threo-hex-4-enopyranuronic acid-(1-4)-2-deoxy-6-O-sulfo-2-(sulfoamino)-alpha-D-glucopyranose-(1-4)-2-O-sulfo-alpha-L-idopyranuronic acid-(1-4)-2-deoxy-6-O-sulfo-2-(sulfoamino)-alpha-D-glucopyranose, Netrin receptor unc-5 | | Authors: | Priest, J.M, Ozkan, E. | | Deposit date: | 2022-09-04 | | Release date: | 2023-01-11 | | Last modified: | 2024-08-14 | | Method: | X-RAY DIFFRACTION (2.11 Å) | | Cite: | Structural insights into the formation of repulsive netrin guidance complexes.
Sci Adv, 10, 2024
|
|
8EDK
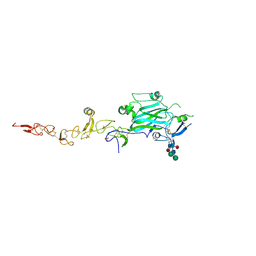 
 | | Structure of C. elegans UNC-6 LamN and EGF domains | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, Netrin unc-6, ... | | Authors: | Priest, J.M, Ozkan, E. | | Deposit date: | 2022-09-04 | | Release date: | 2023-01-11 | | Last modified: | 2024-08-14 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural insights into the formation of repulsive netrin guidance complexes.
Sci Adv, 10, 2024
|
|
7RPY
 
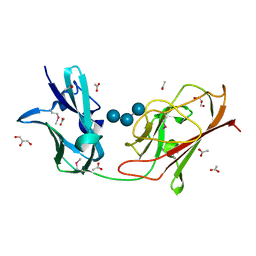 | | X25-2 domain of Sca5 from Ruminococcus bromii | | Descriptor: | ACETATE ION, Cohesin-containing protein, GLYCEROL, ... | | Authors: | Cerqueira, F, Koropatkin, N. | | Deposit date: | 2021-08-04 | | Release date: | 2022-04-13 | | Last modified: | 2022-05-25 | | Method: | X-RAY DIFFRACTION (1.67 Å) | | Cite: | Sas20 is a highly flexible starch-binding protein in the Ruminococcus bromii cell-surface amylosome.
J.Biol.Chem., 298, 2022
|
|
7RFT
 
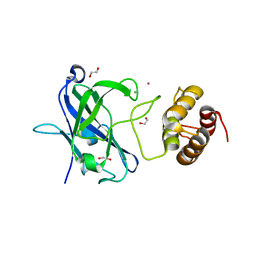 | |
7RAW
 
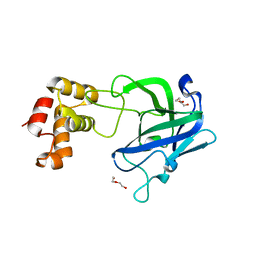 | |
