3F63
 
 | |
3F6D
 
 | | Crystal Structure of a Genetically Modified Delta Class GST (adGSTD4-4) from Anopheles dirus, F123A, in Complex with S-Hexyl Glutathione | | Descriptor: | Glutathione transferase GST1-4, S-HEXYLGLUTATHIONE | | Authors: | Wongsantichon, J, Robinson, R.C, Ketterman, A.J. | | Deposit date: | 2008-11-05 | | Release date: | 2009-10-27 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural contributions of delta class glutathione transferase active-site residues to catalysis
Biochem.J., 428, 2010
|
|
3GH6
 
 | |
4JOU
 
 | |
3G7J
 
 | | Crystal Structure of a Genetically Modified Delta Class GST (adGSTD4-4) from Anopheles dirus, Y119E, in Complex with S-Hexyl Glutathione | | Descriptor: | Glutathione transferase GST1-4, S-HEXYLGLUTATHIONE | | Authors: | Wongsantichon, J, Robinson, R.C, Ketterman, A.J. | | Deposit date: | 2009-02-10 | | Release date: | 2010-02-16 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural contributions of delta class glutathione transferase active-site residues to catalysis
Biochem.J., 428, 2010
|
|
3G7I
 
 | |
5A62
 
 | | Hydrolytic potential of the ammonia-oxidizing Thaumarchaeon Nitrososphaera gargenis - crystal structure and activity profiles of carboxylesterases linked to their metabolic function | | Descriptor: | ACETATE ION, PUTATIVE ALPHA/BETA HYDROLASE FOLD PROTEIN | | Authors: | Chow, J, Kaljunen, H, Nittinger, E, Spieck, E, Rarey, M, Mueller-Dieckmann, J, Streit, W.R. | | Deposit date: | 2015-06-24 | | Release date: | 2016-07-06 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Hydrolytic Potential of the Ammonia-Oxidizing Thaumarchaeon Nitrososphaera Gargenis - Crystal Structure and Activity Profiles of Carboxylesterases Linked to Their Metabolic Function
To be Published
|
|
4YH2
 
 | |
6KA7
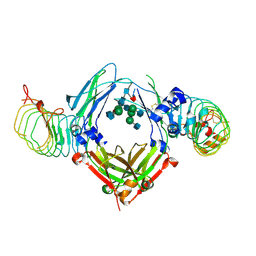 
 | | The complex structure of Human IgG Fc and its binding Repebody | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-2)-alpha-D-mannopyranose-(1-3)-[2-acetamido-2-deoxy-beta-D-glucopyranose-(1-2)-alpha-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, Immunoglobulin gamma-1 heavy chain, repebody | | Authors: | Choi, J, Kim, H. | | Deposit date: | 2019-06-21 | | Release date: | 2020-06-24 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | The complex structure of Human IgG Fc and its binding Repebody
To Be Published
|
|
4FBM
 
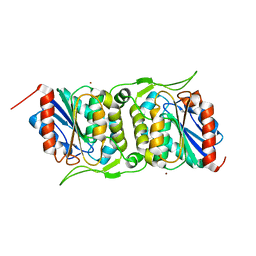 | | LipS and LipT, two metagenome-derived lipolytic enzymes increase the diversity of known lipase and esterase families | | Descriptor: | BROMIDE ION, LipS lipolytic enzyme | | Authors: | Chow, J, Krauss, U, Dall Antonia, Y, Fersini, F, Schmeisser, C, Schmidt, M, Menyes, I, Bornscheuer, U, Lauinger, B, Bongen, P, Pietruszka, J, Eckstein, M, Thum, O, Liese, A, Mueller-Dieckmann, J, Jaeger, K.-E, Kovavic, F, Streit, W.R, Structural Proteomics in Europe (SPINE) | | Deposit date: | 2012-05-23 | | Release date: | 2012-10-10 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | The Metagenome-Derived Enzymes LipS and LipT Increase the Diversity of Known Lipases.
Plos One, 7, 2012
|
|
4FBL
 
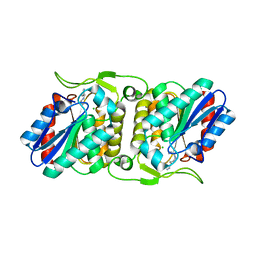 | | LipS and LipT, two metagenome-derived lipolytic enzymes increase the diversity of known lipase and esterase families | | Descriptor: | CHLORIDE ION, LipS lipolytic enzyme, SPERMIDINE | | Authors: | Chow, J, Krauss, U, Dall Antonia, Y, Fersini, F, Schmeisser, C, Schmidt, M, Menyes, I, Bornscheuer, U, Lauinger, B, Bongen, P, Pietruszka, J, Eckstein, M, Thum, O, Liese, A, Mueller-Dieckmann, J, Jaeger, K.-E, Kovacic, F, Streit, W.R, Structural Proteomics in Europe (SPINE) | | Deposit date: | 2012-05-23 | | Release date: | 2012-10-10 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.99 Å) | | Cite: | The Metagenome-Derived Enzymes LipS and LipT Increase the Diversity of Known Lipases.
Plos One, 7, 2012
|
|
3S4P
 
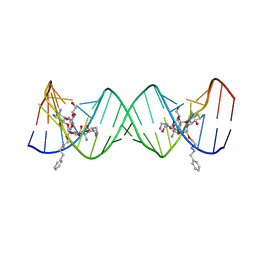 | | Crystal structure of the bacterial ribosomal decoding site complexed with an amphiphilic paromomycin O2''-ether analogue | | Descriptor: | (1R,2R,3S,4R,6S)-4,6-diamino-2-{[3-O-(2,6-diamino-2,6-dideoxy-beta-L-idopyranosyl)-2-O-{2-[(2-phenylethyl)amino]ethyl}-beta-D-ribofuranosyl]oxy}-3-hydroxycyclohexyl 2-amino-2-deoxy-alpha-D-glucopyranoside, RNA (5'-R(P*GP*CP*GP*UP*CP*AP*CP*AP*CP*CP*GP*GP*UP*GP*AP*AP*GP*UP*CP*GP*C)-3') | | Authors: | Szychowski, J, Kondo, J, Zahr, O, Auclair, K, Westhof, E, Hanessian, S, Keillor, J.W. | | Deposit date: | 2011-05-20 | | Release date: | 2011-09-21 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.56 Å) | | Cite: | Inhibition of aminoglycoside-deactivating enzymes APH(3')-IIIa and AAC(6')-Ii by amphiphilic paromomycin O2''-ether analogues
Chemmedchem, 6, 2011
|
|
8TUM
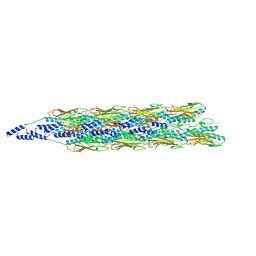 
 | |
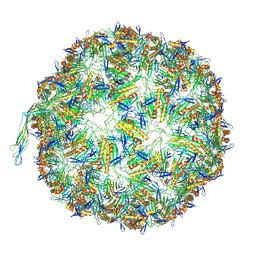
8TUX
 
 | |
8TUW
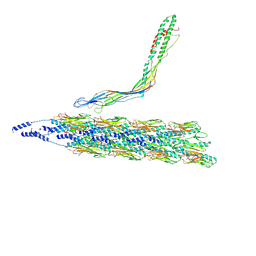 
 | |
3MAK
 
 | |
1U3I
 
 | | Crystal structure of glutathione S-tranferase from Schistosoma mansoni | | Descriptor: | GLUTATHIONE, Glutathione S-transferase 28 kDa | | Authors: | Chomilier, J, Vaney, M.C, Labesse, G, Trottein, F, Capron, A, Mormon, J.-P. | | Deposit date: | 2004-07-22 | | Release date: | 2005-07-26 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.89 Å) | | Cite: | Crystal structure of Schistosoma mansoni glutathione S-transferase
To be Published
|
|
4OZ5
 
 | | Bacillus subtilis HmoB | | Descriptor: | Bacillus subtilis HmoB, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Choe, J, Park, S. | | Deposit date: | 2014-02-13 | | Release date: | 2015-02-18 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.71 Å) | | Cite: | Bacillus subtilis HmoB
To Be Published
|
|
1N1E
 
 | |
1N1G
 
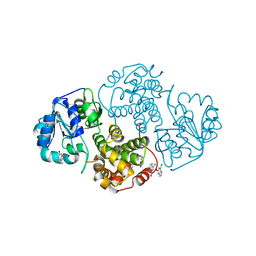 | |
1M66
 
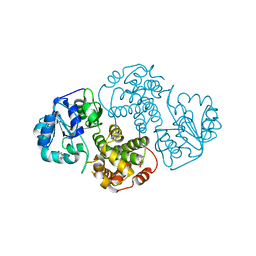 | | Crystal Structure of Leishmania mexicana GPDH Complexed with Inhibitor 2-bromo-6-chloro-purine | | Descriptor: | 2-BROMO-6-CHLORO-PURINE, Glycerol-3-phosphate dehydrogenase, PALMITIC ACID | | Authors: | Choe, J, Suresh, S, Wisedchaisri, G, Kennedy, K.J, Gelb, M.H, Hol, W.G.J. | | Deposit date: | 2002-07-12 | | Release date: | 2002-12-11 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Anomalous differences of light elements in determining precise binding modes of ligands to
glycerol-3-phosphate dehydrogenase
Chem.Biol., 9, 2002
|
|
1M67
 
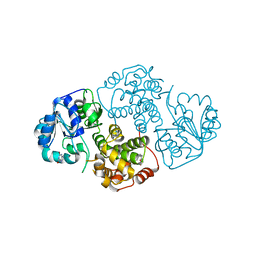 | | Crystal Structure of Leishmania mexicana GPDH Complexed with Inhibitor 2-bromo-6-hydroxy-purine | | Descriptor: | 2-BROMO-6-HYDROXY-PURINE, Glycerol-3-phosphate dehydrogenase, PALMITIC ACID | | Authors: | Choe, J, Suresh, S, Wisedchaisri, G, Kennedy, K.J, Gelb, M.H, Hol, W.G.J. | | Deposit date: | 2002-07-12 | | Release date: | 2002-12-11 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Anomalous differences of light elements in determining precise binding modes of ligands to
glycerol-3-phosphate dehydrogenase
Chem.Biol., 9, 2002
|
|
3RD3
 
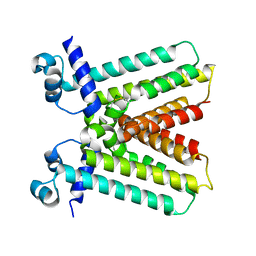 | |
7SS6
 
 | | Structure of Klebsiella LpxH in complex with JH-LPH-45 | | Descriptor: | 1,2-ETHANEDIOL, 5-{4-[3-chloro-5-(trifluoromethyl)phenyl]piperazine-1-sulfonyl}-N-[5-(hydroxyamino)-5-oxopentyl]-2,3-dihydro-1H-indole-1-carboxamide, MANGANESE (II) ION, ... | | Authors: | Cho, J, Cochrane, C.S, Zhou, P. | | Deposit date: | 2021-11-09 | | Release date: | 2023-04-12 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.74 Å) | | Cite: | Development of LpxH Inhibitors Chelating the Active Site Dimanganese Metal Cluster of LpxH.
Chemmedchem, 18, 2023
|
|
7SS7
 
 | | Crystal structure of Klebsiella LpxH in complex with JH-LPH-50 | | Descriptor: | 1,2-ETHANEDIOL, 6-{[(4-{4-[3-chloro-5-(trifluoromethyl)phenyl]piperazine-1-sulfonyl}phenyl)carbamoyl]amino}-N-hydroxyhexanamide, MANGANESE (II) ION, ... | | Authors: | Cho, J, Cochrane, C.S, Zhou, P. | | Deposit date: | 2021-11-09 | | Release date: | 2023-04-12 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.73 Å) | | Cite: | Development of LpxH Inhibitors Chelating the Active Site Dimanganese Metal Cluster of LpxH.
Chemmedchem, 18, 2023
|
|
