5IT4
 
 | |
5JLR
 
 | | Crystal structure of Mycobacterium avium SerB2 with serine present at slightly different position near ACT domain | | Descriptor: | 1,2-ETHANEDIOL, 2-(2-METHOXYETHOXY)ETHANOL, MAGNESIUM ION, ... | | Authors: | Shree, S, Agrawal, A, Dubey, S, Ramachandran, R. | | Deposit date: | 2016-04-27 | | Release date: | 2017-05-03 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.261 Å) | | Cite: | Crystal structure of Mycobacterium avium SerB2 with serine present at slightly different position near ACT domain
To be published
|
|
5JJB
 
 | |
5JMA
 
 | | Crystal structure of Mycobacterium avium SerB2 in complex with serine at catalytic (PSP) domain | | Descriptor: | 1,2-ETHANEDIOL, MAGNESIUM ION, Phosphoserine phosphatase, ... | | Authors: | Shree, S, Dubey, S, Agrawal, A, Ramachandran, R. | | Deposit date: | 2016-04-28 | | Release date: | 2017-05-03 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | Crystal structure of Mycobacterium avium SerB2 in complex with serine at catalytic (PSP) domain
To be published
|
|
5IT0
 
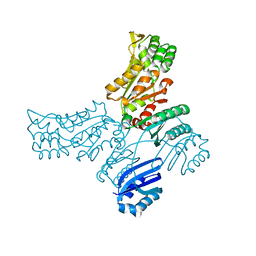 | |
5JLP
 
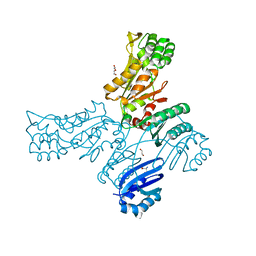 | |
8EPS
 
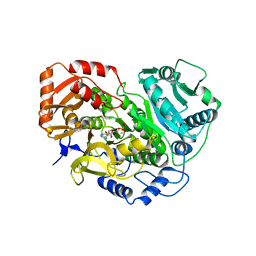 | |
8F82
 
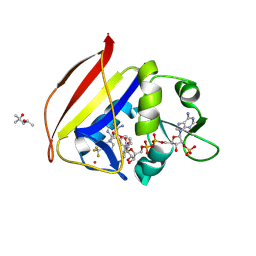 | |
8F83
 
 | |
8F85
 
 | |
8F81
 
 | |
8F84
 
 | |
8F80
 
 | |
8FI4
 
 | |
8FI6
 
 | |
8FI5
 
 | |
8G0R
 
 | |
8G0V
 
 | |
8G0T
 
 | |
8G0U
 
 | |
8G0S
 
 | |
3KJR
 
 | | Crystal structure of dihydrofolate reductase/thymidylate synthase from Babesia bovis determined using SlipChip based microfluidics | | Descriptor: | 2-[N-CYCLOHEXYLAMINO]ETHANE SULFONIC ACID, Dihydrofolate reductase/thymidylate synthase, GLYCEROL, ... | | Authors: | Li, L, Du, W, Edwards, T.E, Staker, B.L, Phan, I, Stacy, R, Ismagilov, R.F, Accelerated Technologies Center for Gene to 3D Structure (ATCG3D), Seattle Structural Genomics Center for Infectious Disease (SSGCID) | | Deposit date: | 2009-11-03 | | Release date: | 2009-11-17 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Multiparameter screening on SlipChip used for nanoliter protein crystallization combining free interface diffusion and microbatch methods.
J.Am.Chem.Soc., 132, 2010
|
|
3HZU
 
 | |
3P3A
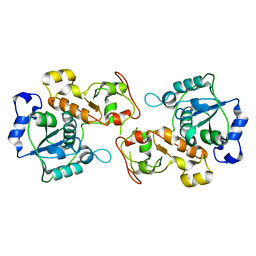 
 | |
3NFW
 
 | |
