6HHB
 
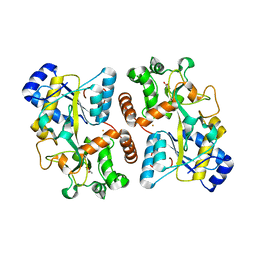 | | Structure of iron bound IbpS from Dickeya dadantii | | Descriptor: | ABC-type Fe3+ transport system, periplasmic component, ACETATE ION, ... | | Authors: | Gueguen-Chaignon, V, Condemine, G, Terradot, L. | | Deposit date: | 2018-08-27 | | Release date: | 2019-09-11 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.80000341 Å) | | Cite: | A secreted metal-binding protein protects necrotrophic phytopathogens from reactive oxygen species.
Nat Commun, 10, 2019
|
|
9B22
 
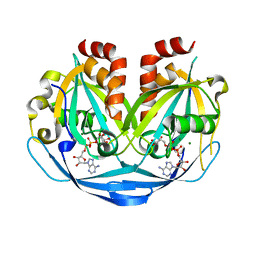 | |
9B1Z
 
 | |
9B20
 
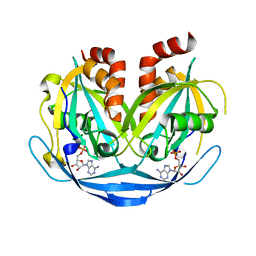 | |
9BN9
 
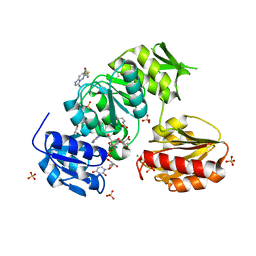 | |
9BN8
 
 | |
9BJL
 
 | |
9BKZ
 
 | |
7SD5
 
 | | Crystallographic structure of neutralizing antibody 10-40 in complex with SARS-CoV-2 spike receptor binding domain | | Descriptor: | 10-40 Heavy chain, 10-40 Light chain, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Reddem, E.R, Casner, R.G, Shapiro, L. | | Deposit date: | 2021-09-29 | | Release date: | 2022-04-27 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.53 Å) | | Cite: | An antibody class with a common CDRH3 motif broadly neutralizes sarbecoviruses.
Sci Transl Med, 14, 2022
|
|
7SI2
 
 | |
6FJL
 
 | | Structure of IbpS from Dickeya dadantii | | Descriptor: | ABC-type Fe3+ transport system, periplasmic component, ACETATE ION, ... | | Authors: | Gueguen-Chaignon, V, Condemine, G, Terradot, L. | | Deposit date: | 2018-01-22 | | Release date: | 2019-02-06 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.70000041 Å) | | Cite: | A secreted metal-binding protein protects necrotrophic phytopathogens from reactive oxygen species.
Nat Commun, 10, 2019
|
|
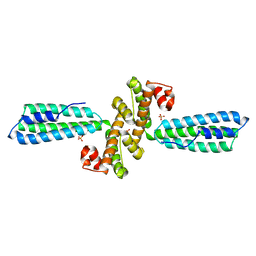
3KPH
 | |
3DKE
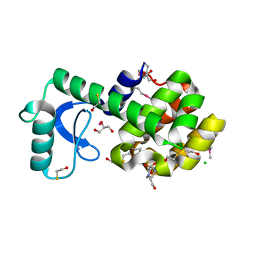 
 | | Polar and non-polar cavities in phage T4 lysozyme | | Descriptor: | 2-HYDROXYETHYL DISULFIDE, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, AZIDE ION, ... | | Authors: | Liu, L.J, Matthews, B.W. | | Deposit date: | 2008-06-24 | | Release date: | 2008-11-25 | | Last modified: | 2021-10-20 | | Method: | X-RAY DIFFRACTION (1.25 Å) | | Cite: | Use of experimental crystallographic phases to examine the hydration of polar and nonpolar cavities in T4 lysozyme
Proc.Natl.Acad.Sci.USA, 105, 2008
|
|
9BKY
 
 | |
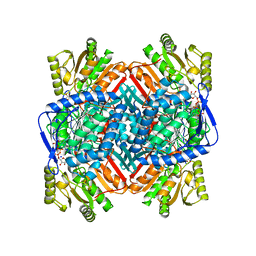
8UZM
 | |
8UZO
 | |
9AV6
 
 | |
8UZI
 
 | |
9AV7
 
 | |
8V4R
 
 | |
8UZ4
 | |
8UR4
 
 | |
8W0J
 
 | |
8W0H
 
 | |
8UZ8
 
 | |
