3F7M
 
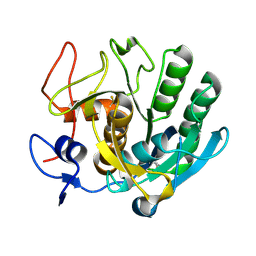 | | Crystal structure of apo Cuticle-Degrading Protease (ver112) from Verticillium psalliotae | | Descriptor: | Alkaline serine protease ver112 | | Authors: | Liang, L, Lou, Z, Ye, F, Meng, Z, Rao, Z, Zhang, K. | | Deposit date: | 2008-11-09 | | Release date: | 2009-11-17 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | The crystal structures of two cuticle-degrading proteases from nematophagous fungi and their contribution to infection against nematodes.
Faseb J., 24, 2010
|
|
3DE5
 
 | |
3DVR
 
 | |
3DWE
 
 | |
3DW3
 
 | |
3F49
 
 | |
1SIU
 
 | | KUMAMOLISIN-AS E78H MUTANT | | Descriptor: | CALCIUM ION, SULFATE ION, kumamolisin-As | | Authors: | Li, M, Wlodawer, A, Gustchina, A, Tsuruoka, N, Ashida, M, Minakata, H, Oyama, H, Oda, K, Nishino, T, Nakayama, T. | | Deposit date: | 2004-03-01 | | Release date: | 2004-03-30 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.31 Å) | | Cite: | Crystallographic and biochemical investigations of kumamolisin-As, a serine-carboxyl peptidase with collagenase activity
J.Biol.Chem., 279, 2004
|
|
1SBH
 
 | | SUBTILISIN BPN' 8397+1 (E.C. 3.4.21.14) MUTANT (M50F, N76D, G169A, Q206C, N218S, K256Y) | | Descriptor: | CALCIUM ION, SUBTILISIN 8397+1 | | Authors: | Kidd, R.D, Farber, G.K. | | Deposit date: | 1995-09-01 | | Release date: | 1995-12-07 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | A weak calcium binding site in subtilisin BPN' has a dramatic effect on protein stability.
J.Am.Chem.Soc., 118, 1996
|
|
1SBT
 
 | | ATOMIC COORDINATES FOR SUBTILISIN BPN (OR NOVO) | | Descriptor: | SUBTILISIN BPN' | | Authors: | Alden, R.A, Birktoft, J.J, Kraut, J, Robertus, J.D, Wright, C.S. | | Deposit date: | 1972-08-11 | | Release date: | 1977-01-06 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Atomic coordinates for subtilisin BPN' (or Novo).
Biochem.Biophys.Res.Commun., 45, 1971
|
|
1SBC
 
 | |
1SEL
 
 | |
1SBI
 
 | | SUBTILISIN BPN' 8397 (E.C. 3.4.21.14) MUTANT (M50F, N76D, G169A, Q206C, N218S) | | Descriptor: | CALCIUM ION, SUBTILISIN 8397 | | Authors: | Kidd, R.D, Yennawar, H.P, Farber, G.K. | | Deposit date: | 1995-09-01 | | Release date: | 1995-12-07 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | A weak calcium binding site in subtilisin BPN' has a dramatic effect on protein stability.
J.Am.Chem.Soc., 118, 1996
|
|
1SUB
 
 | | CALCIUM-INDEPENDENT SUBTILISIN BY DESIGN | | Descriptor: | ACETONE, CALCIUM ION, POTASSIUM ION, ... | | Authors: | Gallagher, T, Bryan, P, Gilliland, G.L. | | Deposit date: | 1992-06-10 | | Release date: | 1994-01-31 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Calcium-independent subtilisin by design.
Proteins, 16, 1993
|
|
1SUE
 
 | | SUBTILISIN BPN' FROM BACILLUS AMYLOLIQUEFACIENS, MUTANT | | Descriptor: | DIISOPROPYL PHOSPHONATE, SODIUM ION, SUBTILISIN BPN' | | Authors: | Gallagher, D.T, Bryan, P, Pan, Q, Gilliland, G.L. | | Deposit date: | 1998-02-17 | | Release date: | 1998-10-14 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Mechanism of ionic strength dependence of crystal growth rates in a subtilisin variant.
J.Cryst.Growth, 193, 1998
|
|
1ST2
 
 | |
1UBN
 
 | | SELENOSUBTILISIN BPN | | Descriptor: | CALCIUM ION, PROTEIN (SELENOSUBTILISIN BPN) | | Authors: | McRee, D.E, McTigue, M, Hilvert, D. | | Deposit date: | 1999-06-02 | | Release date: | 1999-06-08 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Electric fields in active sites: substrate switching from null to strong fields in thiol- and selenol-subtilisins.
Biochemistry, 38, 1999
|
|
1S2N
 
 | |
1SCA
 
 | | ENZYME CRYSTAL STRUCTURE IN A NEAT ORGANIC SOLVENT | | Descriptor: | CALCIUM ION, SODIUM ION, SUBTILISIN CARLSBERG | | Authors: | Fitzpatrick, P.A, Steinmetz, A.C.U, Ringe, D, Klibanov, A.M. | | Deposit date: | 1993-07-19 | | Release date: | 1994-01-31 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Enzyme crystal structure in a neat organic solvent.
Proc.Natl.Acad.Sci.USA, 90, 1993
|
|
1SUC
 
 | |
1T1I
 
 | | High Resolution Crystal Structure of Mutant W129A of Kumamolisin, a Sedolisin Type Proteinase (previously called Kumamolysin or KSCP) | | Descriptor: | CALCIUM ION, SULFATE ION, kumamolisin | | Authors: | Comellas-Bigler, M, Maskos, K, Huber, R, Oyama, H, Oda, K, Bode, W. | | Deposit date: | 2004-04-16 | | Release date: | 2004-08-03 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.28 Å) | | Cite: | 1.2 a crystal structure of the serine carboxyl proteinase pro-kumamolisin: structure of an intact pro-subtilase
Structure, 12, 2004
|
|
1SUA
 
 | | SUBTILISIN BPN' | | Descriptor: | SUBTILISIN BPN', TETRAPEPTIDE ALA-LEU-ALA-LEU | | Authors: | Almog, O, Gilliland, G.L. | | Deposit date: | 1997-01-14 | | Release date: | 1998-01-14 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structure of calcium-independent subtilisin BPN' with restored thermal stability folded without the prodomain.
Proteins, 31, 1998
|
|
1S01
 
 | |
1S02
 
 | | EFFECTS OF ENGINEERED SALT BRIDGES ON THE STABILITY OF SUBTILISIN BPN' | | Descriptor: | CALCIUM ION, SUBTILISIN BPN', SULFATE ION | | Authors: | Erwin, C.R, Barnett, B.L, Oliver, J.D, Sullivan, J.F. | | Deposit date: | 1991-02-20 | | Release date: | 1992-01-15 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Effects of engineered salt bridges on the stability of subtilisin BPN'.
Protein Eng., 4, 1990
|
|
1SCN
 
 | | INACTIVATION OF SUBTILISIN CARLSBERG BY N-(TERT-BUTOXYCARBONYL-ALANYL-PROLYL-PHENYLALANYL)-O-BENZOL HYDROXYLAMINE: FORMATION OF COVALENT ENZYME-INHIBITOR LINKAGE IN THE FORM OF A CARBAMATE DERIVATIVE | | Descriptor: | CALCIUM ION, N-(tert-butoxycarbonyl)-L-alanyl-N-[(1R)-1-(carboxyamino)-2-phenylethyl]-L-prolinamide, SODIUM ION, ... | | Authors: | Steinmetz, A.C.U, Demuth, H.-U, Ringe, D. | | Deposit date: | 1994-03-02 | | Release date: | 1994-08-31 | | Last modified: | 2017-11-29 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Inactivation of subtilisin Carlsberg by N-((tert-butoxycarbonyl)alanylprolylphenylalanyl)-O-benzolhydroxyl- amine: formation of a covalent enzyme-inhibitor linkage in the form of a carbamate derivative.
Biochemistry, 33, 1994
|
|
1ST3
 
 | |
