1T6W
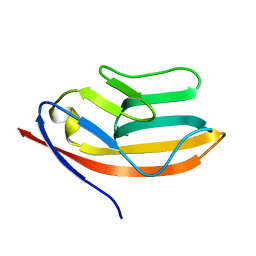 
 | | RATIONAL DESIGN OF A CALCIUM-BINDING ADHESION PROTEIN NMR, 20 STRUCTURES | | Descriptor: | CALCIUM ION, hypothetical protein XP_346638 | | Authors: | Yang, W, Wilkins, A.L, Ye, Y, Liu, Z.-R, Urbauer, J.L, Kearney, A, van der Merwe, P.A, Yang, J.J. | | Deposit date: | 2004-05-07 | | Release date: | 2005-02-15 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Design of a calcium-binding protein with desired structure in a cell adhesion molecule.
J.Am.Chem.Soc., 127, 2005
|
|
1T8J
 
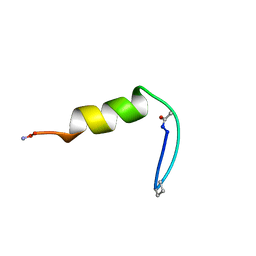 | | NMR Structure of BBA5, A Compact, Independently Folded BBA Motif | | Descriptor: | BBA5 | | Authors: | Struthers, M.D, Ottesen, J.J, Imperiali, B. | | Deposit date: | 2004-05-13 | | Release date: | 2004-05-25 | | Last modified: | 2024-11-13 | | Method: | SOLUTION NMR | | Cite: | Design and NMR Analyses of Compact, Independently Folded BBA Motifs
Fold.Des., 3, 1998
|
|
1T8V
 
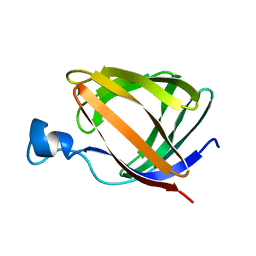 | |
1T9E
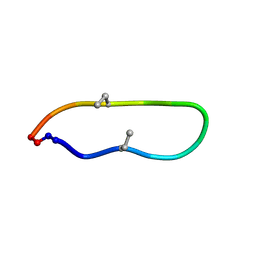 
 | |
1TAC
 | |
1TAP
 
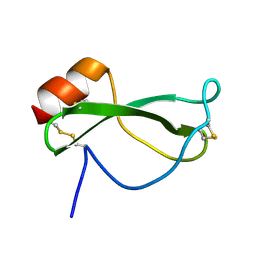 | | NMR SOLUTION STRUCTURE OF RECOMBINANT TICK ANTICOAGULANT PROTEIN (RTAP), A FACTOR XA INHIBITOR FROM THE TICK ORNITHODOROS MOUBATA | | Descriptor: | FACTOR XA INHIBITOR | | Authors: | Antuch, W, Guntert, P, Billeter, M, Wuthrich, K. | | Deposit date: | 1994-08-16 | | Release date: | 1994-11-30 | | Last modified: | 2024-11-20 | | Method: | SOLUTION NMR | | Cite: | NMR solution structure of the recombinant tick anticoagulant protein (rTAP), a factor Xa inhibitor from the tick Ornithodoros moubata.
FEBS Lett., 352, 1994
|
|
1TBA
 
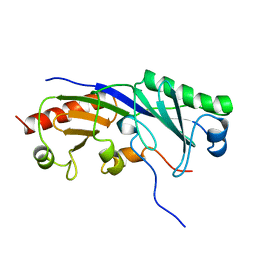 | | SOLUTION STRUCTURE OF A TBP-TAFII230 COMPLEX: PROTEIN MIMICRY OF THE MINOR GROOVE SURFACE OF THE TATA BOX UNWOUND BY TBP, NMR, 25 STRUCTURES | | Descriptor: | TRANSCRIPTION INITIATION FACTOR IID 230K CHAIN, TRANSCRIPTION INITIATION FACTOR TFIID | | Authors: | Liu, D, Ishima, R, Tong, K.I, Bagby, S, Kokubo, T, Muhandiram, D.R, Kay, L.E, Nakatani, Y, Ikura, M. | | Deposit date: | 1998-08-16 | | Release date: | 1999-08-16 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure of a TBP-TAF(II)230 complex: protein mimicry of the minor groove surface of the TATA box unwound by TBP.
Cell(Cambridge,Mass.), 94, 1998
|
|
1TBC
 | |
1TBD
 
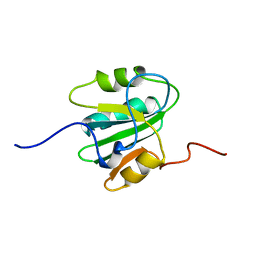 | | SOLUTION STRUCTURE OF THE ORIGIN DNA BINDING DOMAIN OF SV40 T-ANTIGEN, NMR, MINIMIZED AVERAGE STRUCTURE | | Descriptor: | SV40 T-ANTIGEN | | Authors: | Luo, X, Sanford, D.G, Bullock, P.A, Bachovchin, W.W. | | Deposit date: | 1996-11-04 | | Release date: | 1997-03-12 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the origin DNA-binding domain of SV40 T-antigen.
Nat.Struct.Biol., 3, 1996
|
|
1TBK
 
 | |
1TCG
 
 | |
1TCH
 
 | |
1TCP
 
 | |
1TDP
 
 | |
1TE4
 
 | | Solution structure of MTH187. Ontario Centre for Structural Proteomics target MTH0187_1_111; Northeast Structural Genomics Target TT740 | | Descriptor: | conserved protein MTH187 | | Authors: | Gignac, I, Julien, O, Yee, A, Arrowsmith, C.H, Gagne, S.M, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2004-05-24 | | Release date: | 2004-07-13 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | MTH187 from Methanobacterium thermoautotrophicum has three HEAT-like Repeats.
J.Biomol.Nmr, 35, 2006
|
|
1TE7
 
 | | Solution NMR Structure of Protein yqfB from Escherichia coli. Northeast Structural Genomics Consortium Target ET99 | | Descriptor: | Hypothetical UPF0267 protein yqfB | | Authors: | Atreya, H.S, Shen, Y, Yee, A, Arrowsmith, C, Szyperski, T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2004-05-24 | | Release date: | 2005-01-04 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | G-Matrix Fourier Transform NOESY-Based Protocol for High-Quality Protein Structure Determination
J.Am.Chem.Soc., 127, 2005
|
|
1TEY
 
 | | NMR structure of human histone chaperone, ASF1A | | Descriptor: | ASF1 anti-silencing function 1 homolog A | | Authors: | Mousson, F, Lautrette, A, Thuret, J.Y, Agez, M, Amigues, B, Courbeyrette, R, Neumann, J.M, Guerois, R, Mann, C, Ochsenbein, F. | | Deposit date: | 2004-05-26 | | Release date: | 2005-04-12 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structural basis for the interaction of Asf1 with histone H3 and its functional implications.
Proc.Natl.Acad.Sci.Usa, 102, 2005
|
|
1TF3
 
 | | TFIIIA FINGER 1-3 BOUND TO DNA, NMR, 22 STRUCTURES | | Descriptor: | 5S RNA GENE, TRANSCRIPTION FACTOR IIIA, ZINC ION | | Authors: | Foster, M.P, Wuttke, D.S, Radhakrishnan, I, Case, D.A, Gottesfeld, J.M, Wright, P.E. | | Deposit date: | 1997-07-01 | | Release date: | 1997-09-17 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Domain packing and dynamics in the DNA complex of the N-terminal zinc fingers of TFIIIA.
Nat.Struct.Biol., 4, 1997
|
|
1TFB
 
 | |
1TFN
 
 | |
1TFS
 
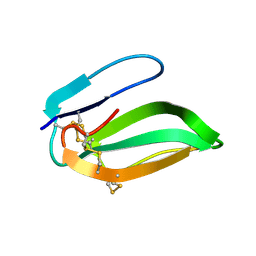 | | NMR AND RESTRAINED MOLECULAR DYNAMICS STUDY OF THE THREE-DIMENSIONAL SOLUTION STRUCTURE OF TOXIN FS2, A SPECIFIC BLOCKER OF THE L-TYPE CALCIUM CHANNEL, ISOLATED FROM BLACK MAMBA VENOM | | Descriptor: | TOXIN FS2 | | Authors: | Albrand, J.-P, Blackledge, M.J, Pascaud, F, Hollecker, M, Marion, D. | | Deposit date: | 1995-01-26 | | Release date: | 1995-03-31 | | Last modified: | 2024-10-23 | | Method: | SOLUTION NMR | | Cite: | NMR and restrained molecular dynamics study of the three-dimensional solution structure of toxin FS2, a specific blocker of the L-type calcium channel, isolated from black mamba venom.
Biochemistry, 34, 1995
|
|
1TH5
 | | Solution structure of C-terminal domain of NifU-like protein from Oryza sativa | | Descriptor: | NifU1 | | Authors: | Kumeta, H, Ogura, K, Asayama, M, Katoh, S, Katoh, E, Inagaki, F. | | Deposit date: | 2004-06-01 | | Release date: | 2005-09-27 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | The NMR structure of the domain II of a chloroplastic NifU-like protein OsNifU1A.
J.Biomol.Nmr, 38, 2007
|
|
1TI3
 
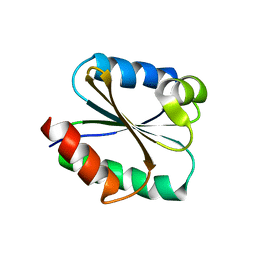 | | Solution structure of the Thioredoxin h1 from poplar, a CPPC active site variant | | Descriptor: | thioredoxin H | | Authors: | Coudevylle, N, Thureau, A, Hemmerlin, C, Gelhaye, E, Jacquot, J.P, Cung, M.T. | | Deposit date: | 2004-06-02 | | Release date: | 2004-12-14 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure of a natural CPPC active site variant, the reduced form of thioredoxin h1 from poplar.
Biochemistry, 44, 2005
|
|
1TIH
 
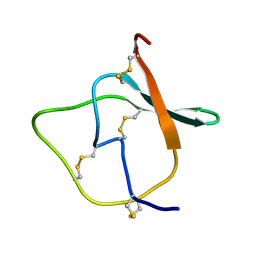 | |
1TIN
 
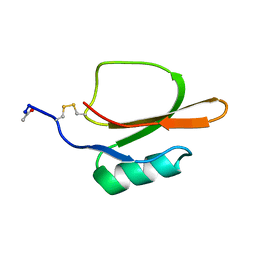 | |
