[English] 日本語
 Yorodumi
Yorodumi- PDB-3dw5: Crystal Structure of the Sarcin/Ricin Domain from E. COLI 23S rRN... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3dw5 | ||||||
|---|---|---|---|---|---|---|---|
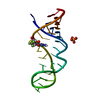
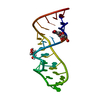
| Title | Crystal Structure of the Sarcin/Ricin Domain from E. COLI 23S rRNA, U2656-OCH3 modified | ||||||
 Components Components | Sarcin/Ricin Domain from E. Coli 23 S rRNA | ||||||
 Keywords Keywords | RNA / Sarcin Ricin Loop | ||||||
| Function / homology | RNA / RNA (> 10) Function and homology information Function and homology information | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / Resolution: 0.96 Å SYNCHROTRON / Resolution: 0.96 Å | ||||||
 Authors Authors | Olieric, V. / Rieder, U. / Lang, K. / Serganov, A. / Schulze-Briese, C. / Micura, R. / Dumas, P. / Ennifar, E. | ||||||
 Citation Citation |  Journal: Rna / Year: 2009 Journal: Rna / Year: 2009Title: A fast selenium derivatization strategy for crystallization and phasing of RNA structures. Authors: Olieric, V. / Rieder, U. / Lang, K. / Serganov, A. / Schulze-Briese, C. / Micura, R. / Dumas, P. / Ennifar, E. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3dw5.cif.gz 3dw5.cif.gz | 44.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3dw5.ent.gz pdb3dw5.ent.gz | 32.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3dw5.json.gz 3dw5.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  3dw5_validation.pdf.gz 3dw5_validation.pdf.gz | 393.8 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  3dw5_full_validation.pdf.gz 3dw5_full_validation.pdf.gz | 394 KB | Display | |
| Data in XML |  3dw5_validation.xml.gz 3dw5_validation.xml.gz | 5.3 KB | Display | |
| Data in CIF |  3dw5_validation.cif.gz 3dw5_validation.cif.gz | 7.2 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/dw/3dw5 https://data.pdbj.org/pub/pdb/validation_reports/dw/3dw5 ftp://data.pdbj.org/pub/pdb/validation_reports/dw/3dw5 ftp://data.pdbj.org/pub/pdb/validation_reports/dw/3dw5 | HTTPS FTP |
-Related structure data
| Related structure data |  3dvzC  3dw4C  3dw6C  3dw7C  1q9aS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: RNA chain | Mass: 8758.282 Da / Num. of mol.: 1 / Source method: obtained synthetically |
|---|---|
| #2: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.9 Å3/Da / Density % sol: 35.25 % | ||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 293 K / Method: vapor diffusion, hanging drop / pH: 7 Details: 3.2 M (NH4)2SO4, 50 mM K-MOPS pH 7.0, 10 mM MgCl2, 10 mM MnCl2, vapor diffusion, hanging drop, temperature 293K | ||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions |
|
-Data collection
| Diffraction | Mean temperature: 90 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SLS SLS  / Beamline: X06SA / Wavelength: 0.85 Å / Beamline: X06SA / Wavelength: 0.85 Å |
| Detector | Type: PSI PILATUS 6M / Detector: PIXEL / Date: Apr 22, 2008 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.85 Å / Relative weight: 1 |
| Reflection | Highest resolution: 0.96 Å / Num. obs: 39656 / % possible obs: 99.8 % / Redundancy: 11.9 % / Biso Wilson estimate: 12.625 Å2 / Rsym value: 0.032 / Net I/σ(I): 34.5 |
| Reflection shell | Resolution: 0.96→0.97 Å / Mean I/σ(I) obs: 3.1 / Num. measured obs: 9571 / Num. unique obs: 1123 / Rsym value: 0.634 / % possible all: 92.9 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Starting model: 1q9a Resolution: 0.96→27.557 Å / Occupancy max: 1 / Occupancy min: 0.25 / SU ML: 0.09 / σ(F): 2 / Stereochemistry target values: ML
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL / Bsol: 39.753 Å2 / ksol: 0.436 e/Å3 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 35.27 Å2 / Biso mean: 14.204 Å2 / Biso min: 7.01 Å2
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 0.96→27.557 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Refine-ID: X-RAY DIFFRACTION / Total num. of bins used: 14
|
 Movie
Movie Controller
Controller








 PDBj
PDBj






























